Development of an Automated Decision Support System for Colon Examination Using Random Texture Descriptor and Decision Tree Induction
Petra Perner*
Institute of Computer Vision and Applied Computer Sciences, Germany
Submission: August 11, 2017; Published: August 29, 2017
*Corresponding author: Petra Perner, Institute of Computer Vision and Applied Computer Sciences, IBaI, Leipzig, Germany,Email: pperner@ibai-institut.de
How to cite this article: Petra P. Development of an Automated Decision Support System for Colon Examination Using Random Texture Descriptor and Decision Tree Induction. Adv Res Gastroentero Hepatol 2017; 6(5): 555698. DOI 10.19080/ARGH.2017.06.555698
Abstract
The aim of our research was to develop an automated decision support system for colon examination that allows us automatically diagnosing medical images in normal tissue images and images showing a polyp. For the system development we used a data set of images that came from an endoscope video system used for colon examination. The data set contains 283 normal tissue images and 61 polyp images of 33x33 image size. The images were cut-out semi-automatically by an expert from larger images. The 283 normal images consist of dark regions and reflection. One must decide if the image shows a polyp or not. This is a two-class problem. The unequal number of the data in the two classes makes our problem to an unbalanced data set problem. The polyps in the images were identified and selected by a "well-trained” medical expert. Our experiments have shown that such kind of images can be well described by a texture descriptor. We used our novel Random set texture descriptor that has shown superior performance on different medical image applications. We review the theory of our texture descriptor and then we apply it to our medical data set. We used a decision-tree induction method to learn the classification rules based on our tool "Decision Master®. The results show that decision tree induction and image analysis based on our novel texture descriptor is an excellent method to mine medical images for the decision rules even when the data set is unbalanced, but not only that makes our Random-set based texture descriptor favourable. It also gives a flexible way to describe the appearance of the medical objects in symbolic terms, the computation time is less, and it can be set up as software module that can be flexible used in different systems. Based on the learnt decision rules and our texture descriptor we can built a semi-automatic decision support system for colon examination where the experts chooses the image region for examination with the help of an image analysis tool, then the chosen sub-image is given to the texture descriptor module, and finally the texture features are given to our learnt decision tree for making the final diagnosis.
Keywords: Image analysis; Endoscope images; Colon examination; Polyp images; Decision tree induction, Random set texture descriptor; Unbalanced data set problem; Decision support system
Introduction
The aim of our research was to develop an automated decision support system for colon examination that allows us automatically diagnosing medical images in normal tissue images and images showing a polyp. We used a data set of medical images of the size 33x33 pixels that came from an endoscope video system used for colon examination. The images showing abnormal and normal tissues have been cut out by an expert from larger images with the help of an image analysing tool.
Texture seems to be a powerful tool to describe the appearances of medical objects into normal tissue and polyp's. Therefore, very flexible and powerful texture descriptors are of importance that allow to recognize the texture and to understand what makes up the texture. Texture seems to become an important role to describe the appearance of different medical and biological objects in images. Patterns on cells in cell images, on fungi images or polyp images can be described by texture.
Different texture descriptors have been developed over the past [1]. The most used texture descriptor is the well- known texture descriptor based on the co-occurrence matrix [2]. Although it works well on different applications we prefer to use our texture descriptor based on Random sets [3] since this descriptor gives us more freedom in describing different textures. In this paper we describe our method to learn decision rules for colon examination. We present our material and the application of the texture descriptors and the decision-tree induction-method in Section 2. The used data set of normal tissue image and polyp images is derived from colon examination. We calculated the texture features based on our novel Random-set method for each image of the data set and learn a decision tree classifier. The theory of the texture descriptor is given in Section 7
Material of the Application and the Data-Mining Tool Decision Master®
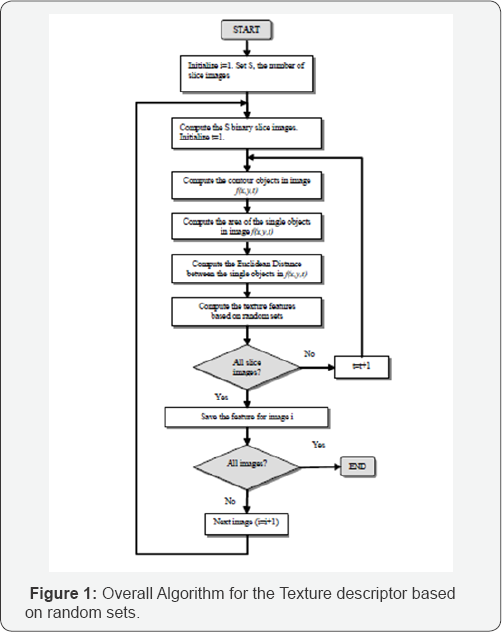
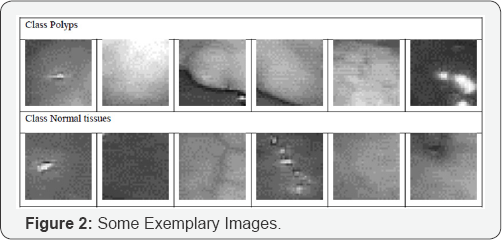
We studied the performance of our feature extraction and decision tree induction method based on a data set of 344 images. These images come from an endoscope video system used for The overall algorithm is shown in Figure 1 colon examination [4]. The data set contains 283 normal tissue images and 61 polyp images (Figure 2) in the form of sub-images of a size 33x33 that are derived from 37 original colonoscopy images. The sub-images have been cut out from the larger images by an expert with the help of an image analysis tool. The polyps in the 37 original colonoscopy images were identified and selected by a "well-trained” medical expert. The 283 normal images consist of dark regions, reflections etc. of the 37 original colonoscopy images.
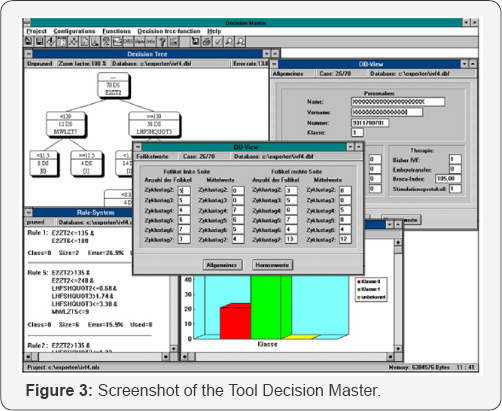
This presents a two-class problem; one must decide if the image shows a polyp or not. The texture descriptions were calculated from these images. The resulting data set was used to train a decision tree based on the C4.5 algorithm [3]. Crossvalidation was used to estimate the error rate (Figure 1). The tool Decision Master® [5] is a data mining tool based on decision tree induction. It contains binary and n-ary decision tree induction methods such as standard algorithm like C4.5, ID3 and n-ary decision-tree induction methods developed by the Institute of Computer Vision and applied Computer Sciences IBaI. It is a commercial tool now and sold worldwide. It allows to compare the learnt models based on standard algorithms and special developed algorithms. N-ary trees get usually more compact than binary trees. The explanation capability of the trees is then better than for binary trees. However, it cannot be said from scratch which decision tree induction method is best based on the error rate for the desired data set. Therefore, it should be easy for the user to check out several decision tree induction methods. The tool Decision Master® allows that in an excellent manner. It has functions for dealing with missing values, erroneous values, and outliers such as the box-plot method and others. N-fold cross-validation or test-and-train can evaluate the learnt model. Several error rates are calculated such as the overall error rate, the class-specific error rate, and the classification quality. The tool has a nice user interface so that a non-computer expert can easy handle it. The other option is to integrate the software as OEM component in larger systems for example in E-commerce suites for on-line user profiling or for learning other information from the trace of the online-user [3]. Figure 3 Screenshot of the data-mining tool Decision Master®
Texture Descriptor based on Random Sets
Boolean sets were invented by Matheron [6]. An indepth description of the theory can be found in Stoyan et al. [7]. The Boolean model allows to model and simulate a huge variety of textures e.g. for crystals, leaves, etc. The texture model X is obtained by taking various realizations of compact random sets, implanting them in Poisson points in Rn, and taking the supremum. The functional moment Q(B) of X, after Booleanization, is calculated as:
where is the set of the compact random set of Rn, the density of the process and
is an average measure that characterizes the geometric properties of the remaining set of objects after dilation. Relation (25) is the fundamental formula of the model. It completely characterizes the texture model. Q(B) does not depend on the location of B , i.e., it is stationary. One can also provide that it is ergodic so that we can peak the measure for a specific portion of the space without referring to the particular portion of the space. Formula 25 show us that the texture model depends on two parameters: the density of the process and a measure that characterizes the objects. In the one-dimensional space, it is the average length of the lines and in the two-dimensional space
is the average measure of the area and the perimeter of the objects under the assumption of convex shapes.
We consider the two-dimensional case and develop a proper texture descriptor. Suppose now that we have a texture image with 8 bit gray levels. Then we can consider the texture image as the superposition of various Boolean models, each of them having a different gray level value on the scale from 0 to 255 for the objects within the bit plane.
To reduce the dimensionality of the resulting feature vector, the gray levels ranging from 0 to 2 55 are now quantized into S intervals t. Each image f(x,y) is classified according to the gray level into t classes, with t= 0,1,2,..,S . For each class a binary image is calculated containing the value “1” for pixels with a gray level value falling into the gray level interval of class t and value “0” for all other pixels. The resulting bit plane f(x,y,t) can now be considered as a realization of the Boolean model. The quantization of the gray level into S intervals was done at equal distances. In the following, we call the image f(x,y,t) a class image. In the class image we can see a lot of different objects. These objects get labeled with the contour-following method [8]. Afterwards, features from the bit-plane and from these objects are calculated. Since it does not make sense to consider the features of every single object due to the curse of dimensionality, we calculate the mean and standard deviation for each feature that characterizes the objects such as the area and the contour. In addition to that, we calculate the number of objects and the areal density in the class image.
The list of features and their calculation are shown in Table 1. The first one is the areal density of the class image t which is the number of pixels in the class image, labeled by “1”, divided by the area of the image. If all pixels of an image are labeled by “1”, then the density is one. If no pixel in an image is labeled, then the density is zero (Table 1).
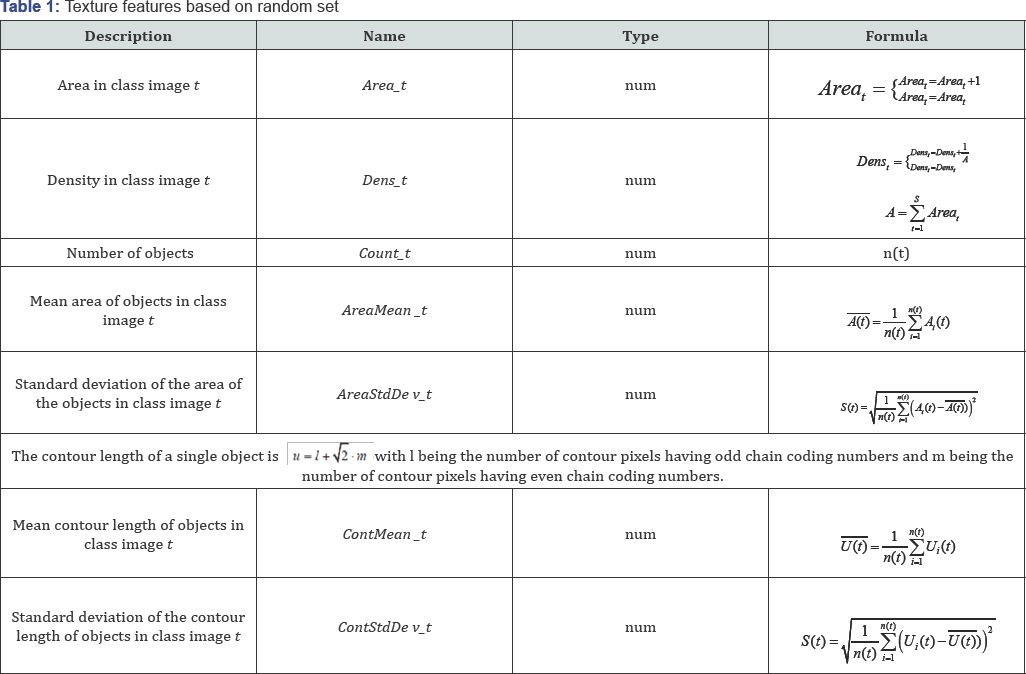
From the objects in the class image t, the area, a simple shape factor, and the length of the contour are calculated. Per the model, not a single feature of each object is taken for classification due to the curse of dimensionality, but the mean and the standard deviation of each feature are calculated over all the objects in the class image t. We also calculate the frequency of the object size in each class image t. Depending on the number of slices S we get a feature set of 42(S=6), 84(S=12), 112(S=16) features.
Result
For the texture descriptor based on random sets the choice of S is important. On the one hand, we need a sufficiently large S to separate the classes. On the other hand, with increasing S also the number of features increases and we run into the curse-of- dimensionality problem.
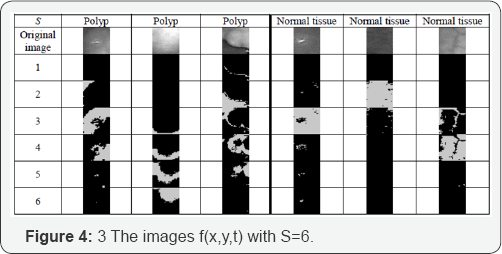
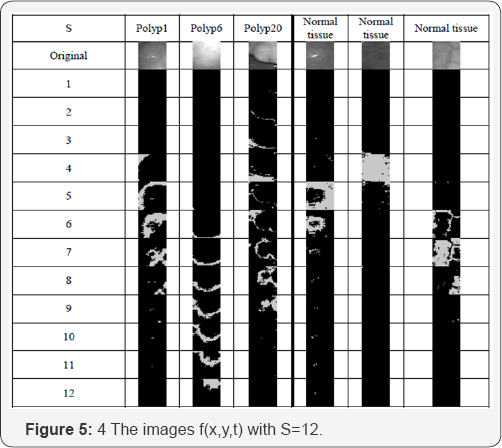
Figure 4 shows the class images for some polyp images and some normal tissue images for S=6. Figure 5 shows the class images for some polyp images and some tissue images for S=12. Figure 5 shows that most pixels of normal tissue images are located in only a few lower 1-3 class images. In contrast to this, in the polyp images the pixels are distributed more across the class images.
For our tests, we used S=6, S=12 and S=16. We have not yet developed a good procedure to estimate the number of S. The determination of the right number of S is still heuristic but in most of our applications S=12 turned out to be a good choice [3]. If we use S larger than twelve we will end up with only dots in the class images that do not give us discrimination power anymore.
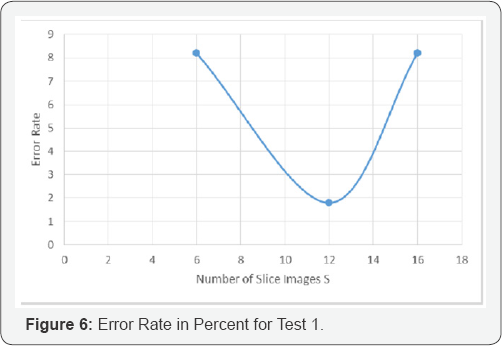
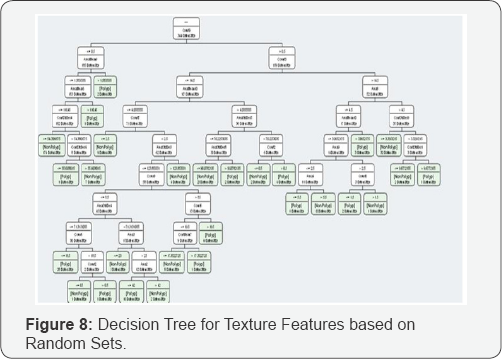
In the first test (test_1), we used 30 polyp images and 30 normal tissue images as a data base. The results are shown in Figure 7. In the second tests (test_2), we used all 344 images as a data base. The results are shown in Figure 8.
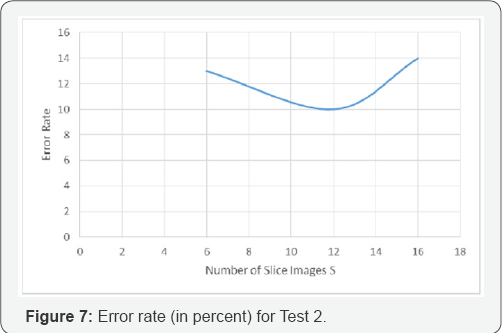
In both tests the texture descriptor based on random sets with S=12 is the best texture descriptor. The test shows that the choice of S=6 is too small and the choice of S=16 is already too large. This observation might demonstrate the effect we described above that the majority of objects in the images are only dots and that we end up in each slice image with only a few objects. The curse of dimensionality might also be a problem but the decision tree induction algorithm only selects the features that are of importance. Thus, the decision tree induction algorithm can be seen as a feature selector if it is also not optimum.
The texture descriptor based on random sets for S=12 has an error rate of 1.67% for the data set with 60 images (Figure 5) with equally distributed number of polyps and normal tissue.
The texture descriptor based on random sets for S=12 has an error rate of 9.88% for the data set with 334 images (Figure 6) with 283 normal tissues and 61 polyps. The result was expected since decision tree induction algorithm heavily rely on the distribution of the classes in the data set. We only report the results here to show this effect.
The resulting decision trees are shown in Figure 7 for the texture features based on random sets.
The feature selection method during decision tree induction selects only 22 features from 84 features for the texture descriptor based on random sets. The runtime of the program for the calculation of the texture descriptor based on random sets is 313.75 seconds. That is very fast compared to other texture descriptors.
Discussion
We have studied the behavior of our novel random set texture descriptor and decision tree induction. The random-set texture-descriptor is a statistical-based texture descriptors the same way the co-occurrence matrix based texture descriptor is. Decision trees are sensitive to unbalanced class distribution. Therefore, the error rate in the second experiment rises since the ratio of the two classes is 1/5 in the data set. Nonetheless, the tendency of the error rate of the three descriptors with its different choice of S is the same.
A further advantage of the texture descriptor based on random sets is the reduced time required for computing the features. In addition, we can understand the semantics behind the numerical texture description. The texture features based on random sets have a semantic meaning and give an expert an understanding about texture.
The choice of the number of slices S emerges to S=12 in the described experiment as well as in all the applications we have done until now. The number S=12 provides a feature set of 84 features. It might be that this is a compromise between a rich description of texture and the large feature set problem (curse- dimensionality). Besides that, our observations showed that the objects in the slice images converges to single points in case of S=16. If this happens then there is no information in the shape or contour anymore.
For the texture descriptor based on Random sets, semantic labels depending on the application can describe the texture and therefore the meaning of the texture is understandable by a human. In case of Figure 4 we can say for poly_1-image, it has objects in the higher slices. In case of poly_20-image, it is a homogenous texture since objects are distributed over all slices, and in case of normal tissue, the objects are in the middle of the slices. The semantic label helps the human to understand the texture and to talk about the texture in a common way. The medical texture objects such as for the polyp images and for cell images [4] are often not large objects. That limits the statistics we can use. We stayed on the first-order statistics. Higher-order statistics make no sense for small objects since the number of objects gets low and no sufficient statistic can be calculated.
The run-time of the random-set texture descriptor is low. This is a big advantage of the random-set texture descriptor over the co-occurrence texture descriptor. It helps to speed up the calculation of the image processing methods. The random- set texture descriptor can be given out a standard module for texture calculation. The input to the module is only the object points and the output of the module are the calculated features for the slices.
The decision tree induction method performs nicely on texture classification. The decision tree induction method is also a feature selector. Therefore, the method can be seen as a learning method for the classification model as well as a feature selector. The texture descriptor based on random sets may provide a richer description of texture. Features from almost all slices are included in the decision.
The data set contains 283 normal tissue images and 61 polyp images. This is a two-class problem. The unequal number of the data in the two classes makes our problem to an unbalanced data set problem. The polyps in the images were identified and selected by a “well-trained” medical expert. The 283 normal images consist of dark regions and reflection. For the full unequally distributed data set, we achieved an error rate of 9.88% based on cross-validation. We achieved an error rate of 1.67% by cross-validation when we created a data set with equally distributed data in each class. To sample the data into two equally distributed data sets is our strategy to deal with the unbalanced data set problem for decision tree induction.
The Architecture of an Online Learning System and an Automatic Decision
Support system
The architecture of a system that allows to learn decision rules and that can be used for automatic decision support is shown in Figure 9. For the learning mode, the video is given to the image and video analysis system. Each frame is shown to the expert and examined by the expert. The expert uses a window of 33x33 pixels and cuts out a sub-image with objects of interest from the displayed image. He labels the images as polyp-image or normal tissue image. This information is put into an image data base that is kept for further investigation purposes. The image is given to the feature-extractor and the texture features are calculated based on our novel Random set texture features. The features are stored together with the image and the respective class label in the image data base. Then a set of data with its features and the class label are given to the decision tree induction tool and the decision tree is learnt. It should be made sure that it is an equally distributed data set. The learnt tree is evaluated and the error rate is inspected by the expert. If he is satisfied with the error rate then the learnt decision tree is given to the classification unit. The classification unit stores the tree and uses the tree for routine inspection purposes of the colon images. The preprocessing of the colon video is the same as for the learning unit. From each video are preprocessed the frames, the cut out of the object of interests are made by the expert, the features are calculated for the 33x33 sub-image, and the features are given to the decision tree. The decision tree classifies the sub-image based on its features and reports the decided class label on the display.
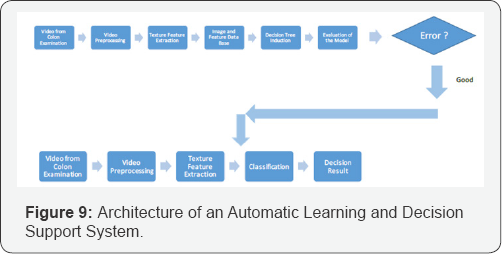
The shown architecture is two divided. It consists of a learning unit and a classification unit for decision support. It also contains a data base in which the images, the calculated texture features for the image, and the class label are stored. This data base is used for learning and can be constantly filled by new ARGHiving images and their respective features and class label. As soon as more data are available the experiment can be repeated in order to get a better performance of the model if wanted [6-11].
Conclusion
We proposed a method for learning decision rules for colon examination. We preprocessed the video into its frames and showed them to the expert on screen. The expert cuts out the region of interests by a window of 33x33 pixels and labels the data into normal tissue or polyp image. Later on the classifier should be automatically able to classify the images into normal and polyp images. This makes the task to a two-class problem. After a large enough data base has been set up we calculate for each image in the data base the features for the texture descriptor based on Random sets. The data set with its texture features and the labels is given to the decision tree induction algorithm. The algorithms learns the decision tree and the error rate based on cross-validation. If the expert is satisfied by the error rate then the decision tree is used for classification.
We have found that the texture descriptor based on Random sets is a superior choice based on the error rate, the tree properties and the runtime. The run-time of the Random-set texture descriptor is quick. This is a big advantage of the Random- set texture descriptor over other texture descriptors since the large computation time of the feature extraction algorithm is still a problem. The Random-set texture descriptor can form a software module that can be used for different applications and for different sizes of objects.
In addition, the texture descriptor based on Random sets has semantic meanings. An expert can understand the properties of the texture when looking at the slices produced during the calculation of the texture features. Therefore, the different appearances in the slices can be labeled by semantic terms that would give us explanation capability of the different textures.
The unbalanced data set problem as it often appears for medical data sets is handled in our study by sampling two equally distributed data sets together for the two-class problem. By doing that we can achieve a good accuracy for the classification.
The architecture of an automatic decision support system can be divided into a learning unit and a classification unit. The preprocessing of the video and the feature extraction is for both units the same. In addition the architecture has a data base where the images, their respective features, and the class label can be stored and used for other purposes.
Acknowledgment
This work has been sponsored under the grant title "Study of the Cognitive Aspects of Human Vision” Cog Vision under the grant number IS 2012-4.
Biography
Petra Perner (IAPR Fellow) is the director of the Institute of Computer Vision and Applied Computer Sciences IBaI. She received her Diploma degree in electrical engineering and her PhD degree in computer science for the work on "Data Reduction Methods for Industrial Robots with Direct Teach-in- Programing”. Her habilitation-thesis was about "A Methodology for the Development of Knowledge-Based Image-Interpretation Systems”. She has been the principal investigator of various national and international research projects. She received several research awards for her research work and has been awarded with three business awards for her work on bringing intelligent image interpretation methods and data mining methods into business. Her research interest is image analysis and interpretation, machine learning, data mining, big data, machine learning, image mining and case-based reasoning. She was running the Committee TC3 Neuronal Nets and the Technical Committee TC 10 of Machine Learning and Data Mining for IAPR. Recently, she is working on various medical, chemical and biomedical applications, information management applications, technical diagnosis and e-commerce applications. Most of the developments are protected by legal patent rights and can be licensed to qualified industrial companies. She has published numerous scientific publications and patents and is often requested as a plenary speaker in distinct research fields as well as across disciplines. She is developing efficient data mining and big data methods and parallel computer architectures such as clouds. Her vision is to build intelligent flexible and robust data-interpreting systems that are inspired by the human case- based reasoning process.
References
- Rao AR (1990) A Taxonomy for Texture Description and Identification, Springer Verlag, Berlin.
- Haralick RH, Shanmugam K, Dingstein I (1973) Textural Features for Image Classification. IEEE Transactions on Systems. Man and Cybernetics 3(6): 610-621.
- Perner P, Perner H, Muller B (2002) Mining Knowledge for Hep-2 Cell Image Classification. Artif Intell Med (26): 161-173.
- Chuan C, Ting WC, Chen YF, Pu Q, Jiang X (2008) Colorectal Polyps Detection Using Texture Features and Support Vector Machine, In: P. Perner, Advances in Mass Data Analysis of Images and Signals in Medicine, Biotechnology, Chemistry and Food Industry Lecture Notes in Computer Science 5108: 62-72.
- Perner P (1998) Classification of HEp-2 Cells Using Fluorescent Image Analysis and Data Mining, 14th Intern. Conference on Pattern Recognition. Brisbane Austr., IEEE Computer Society Press 2: 16771679.
- Matheron G (1975) Random Sets and Integral Geometry. J Wiley & Sons, New York, London.
- Stoyan D, Kendall WS, Mecke J (1987) Stochastic Geometry and Its Applications. Akademie Verlag.
- www.ibai-solutions.de
- Klette R, Zamperoni P (1996) Handbook of image processing operators, Chichester; New York: Wiley, USA.
- Perner P (2002) Data Mining on Multimedia Data, lncs 2558, Springer Verlag.
- Perner P, Trautzsch S (1998) Multinterval Discretization for Decision Tree Learning, In: Advances in Pattern Recognition, A Amin, D Dori, P Pudil (Eds.), Springer Verlag 1998, LNCS 1451, pp. 475-482.
- P Perner, U Zscherpel, C Jacobsen (2001) A Comparision between Neural Networks and Decision Trees based on Data from Industrial Radiographic Testing. Pattern Recognition Letters 22: 47-54






























