Hebatitis B Virus Genotypes in Pakistan
Majid Mahmood*
Department of Zoology, University of Poonch, Rawalakot, Pakistan
Submission: May 10, 2017; Published: June 12, 2017
*Corresponding author:Majid Mahmood, Department of Zoology, University of Poonch, Rawalakot, Azad Jammu and Kashmir, Pakistan, Email: majidll33@gmail.com
How to cite this article:Majid Mahmood. Hebatitis B Virus Genotypes in Pakistan. Adv Res Gastroentero Hepatol 2017; 5(5): 555673. DOI: 10.19080/ARGH.2017.03.555673
Mini Review
Hepatitis B virus (HBV) infection is one of the most common causes of human death worldwide [1-3]. HBV has ten genotypes alphabetized as A to J which are not uniformly distributed in different parts of the world. The genotypes are important in treatment of the infection as they influence the disease progression, baseline viral load and the patients' response to treatment.
Distribution of HBV genotypes in the world
According to Bonino et al. [2], who reviewed the global distribution of HBV genotypes, various HBV genotypes are not uniformly distributed all over the world. Further, their distribution is influenced by ethnic and geographical factors.
Genotype A is common in Brazil, North America, Northwest Europe, South Asia, Central Africa, Tunisia and Benin [2,4,5]. Genotype B is common in China, Hong Kong, Indonesia, Japan, Philippines, Taiwan, Thailand, Vietnam and United States of America. Genotype C is prevalent in Australia, Polynesia, Melanesia, Micronesia, Indonesia, China, Hong Kong, Vietnam, Thailand, Japan, Korea, Taiwan, India, Solomon Islands, Brazil and USA. Genotype D is predominant in Mediterranean region, Spain, Albania, Czech Republic, Russia, Turkey, Middle East, Iran, Afghanistan, India, Solomon Islands, Tunisia, Polynesia, Melanesia, Micronesia, Brazil and USA [6,7]. Genotype E is endemic to Africa where it occurs only in some countries of the Western part of the continent [6]. Genotype F is widely distributed in new world; it has been reported from Alaska, Argentina, Bolivia, Brazil, Central America, Polynesia, South America and USA [8,3]. Genotypes G has been recorded from North America, France and Germany while genotype H is confined to Central America, South America and Mexico [9,10]. Genotypes I and J are confined to Vietnam and Japan, respectively [11,12].
A number of studies in India showed that genotypes D and A are present in Western, Northern and mainland India while genotype C is present in the Eastern parts. In a study [13] on 62 HBV patients from Western India, the region along with Pakistani borderline, genotype D was identified in 92% of the isolates while the remaining 8% were determined as genotype A. None of the other genotypes was recorded in the study. According to another study [14], who reviewed the distribution of HBV genotypes in India, the Northern and Western parts of India are prevailed with genotype D which is followed by genotype A but the Eastern parts of India have a different pattern where genotype C infections made almost equal proportion with genotypes A and D. A study having 1096 samples from nine different areas of China reported that genotypes C (53%) and B (41%) are most common in China [15]. They further identified genotype A, D and a mix infection with combination B+C in 1%, 4% and 1% of the samples respectively.
Prevalence and distribution of hbv genotypes in pakistan
Information regarding the various HBV genotypes in Pakistan and their distribution and prevalence is sketchy and somewhat contrasting. A few studies were conducted on the distribution of HBV genotypes in Pakistan and most of them are confined to some specific cities or areas like Karachi and Punjab. Table 1 summarises the available studies on HBV genotype prevalence in Pakistan, their sample size and sites of sampling along with their results. In brief, a study based on just 12 samples from Lahore reported the presence of only two genotypes; genotype C from nine samples which made 75% while remaining three were positive for genotype D [16].
Idrees et al. [17] studied HBV genotypes in 112 samples which they collect from four provinces of Pakistan and reported that genotype C is dominant with 41.1% prevalence. Genotypes A, B, D and mix genotypes were reported in 21.4%, 17.8%, 8.0% and 7.1% of the samples respectively. Similarly, Abbas et al. [18] performed genotyping of 109 samples from Karachi and reported that 107 (98.2%) of them had genotype D while remaining two (1.8%) were infected with a mixture of genotype A and D. None of the other genotypes were reported in this study.
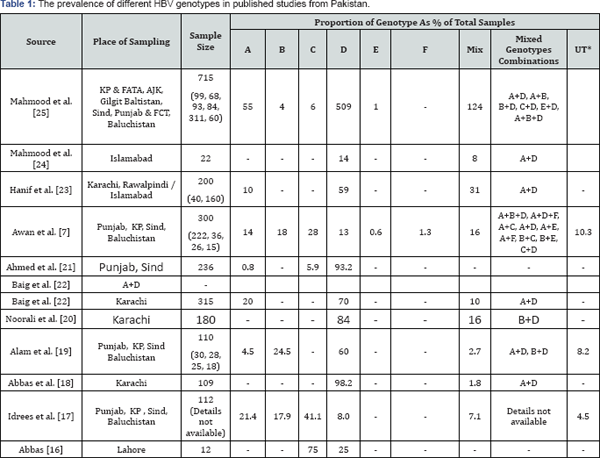
*UT : Un-typeable
Another study described genotyping results for 101 samples which comprised 30 from Punjab [19], 28 from KP, 25 from Sind and 18 from Baluchistan. They identified genotype D in 65%, genotype B in 27% and genotype A in 5% of the samples. Mix genotype infections with combinations A+D and B+D were also reported in this study which were 2.7% of the samples. Yet another study reported that 84% of the 180 female patients in Karachi were infected with genotype D and the rest of 16% had mix infection with genotype B and D [20].
Ahmed et al. [21] conducted genotyping on 236 samples from Punjab and Sind provinces of Pakistan. The study reported that the dominant genotype is D with 93.2% of the samples followed by genotype C with 5.9% while genotype A was rare with only 0.8% of the samples. Sub-genotype D1 was found in 84% and D2 in 8% of the genotype D samples by complete genome sequencing. Genotype C samples belonged to sub genotype C2.
Baig et al. [22] studied 315 samples in Karachi from patients belonging to different ethnic groups. They found that 70% of the samples cARGHied genotype D and 20% of the patients suffered genotype A infection while 10% had mix infection involving genotypes B and D. Using 140 samples from Rawalpindi/ Islamabad, and 60 more from Karachi, a study [23] reported that 59% of the samples have genotype D, 10% have genotype A and 31% have mix infection involving genotype A and D.
However, the genotype profile seems to have been drastically altered by the study of Awan et al. [7] both in terms of the prevalence of genotypes A, B, C, and D and addition of two more genotypes. This study examined 300 samples including 222 from Punjab, 26 from Sind, 30 from Khyber Pakhtunkhwa and 16 from Baluchistan. Out of 300 samples, 269 (89.7%) were successfully genotyped. They identified six HBV genotypes with their prevalence as C(27.7%), B(18%), A(14.3%), D(13%), F(1.3%) and E(0.7%). This percentage is based on 202 single infection samples while mix genotype infections with nine different combinations were observed in 44 samples that make 14.7% of the prevalence.
A recently published smaller study [24] described that about 64 % of the studied HBV samples belong to genotype D while remaining 36 % had a mixture of genotypes A and D. The study however, confirmed sub genotype D1 with complete genome sequencing of 4 out of the total 22 samples. The prevalence and distribution of HBV genotypes is almost confirmed in a conclusive study with a large number of samples from all over the Pakistan [25]. With a total of 715 samples from all 6 geopolitical regions of Pakistan, this study reported that genotype D is the predominant genotype in all regions of Pakistan with an overall prevalence of about 71 %. However, the mix infection with the combination of genotypes A+D was reported to be the second most prevalent infection in Pakistan with a percentage of 13.7. Genotype A, with an infection percentage of 7.7 was reported to be the third most common HBV genotype in Pakistan while about 3.6 % of the HBV infections were reported to be a mixture of genotypes with combinations other than A+D.
In conclusion, It is now clear that HBV/D is the most common genotype in all regions of Pakistan followed by the mix genotype infection with combination of A+D while the third most common HBV genotype in Pakistan is HBV/A.
References
- Zhang Q, Wu G, Richards E, Jia SG, Zeng C (2007) Universal primers for HBV genome DNA amplification across subtypes: a case study for designing more effective viral primers. Virol J 4(1): 92.
- Wiegand J, Hasenclever D, Tillmann HL (2008) Should treatment of hepatitis B depend on hepatitis B virus genotypes? A hypothesis generated from an explorative analysis of published evidence. Antivir Ther 13(2): 211
- Bonino F, Piratvisuth T, Brunetto MR, Liaw YF (2010) Diagnostic markers of chronic hepatitis B infection and disease. Antivir Ther 15(3): 35.
- LJUNGGRE K, Öberg M, Kidd AH (1995) The hepatitis B virus X gene: analysis of functional domain variation and gene phylogeny using multiple sequences. J Gen Virol 76(9): 2119-2130.
- Bowyer SM, van Staden L, Kew MC, Sim JG (1997) A unique segment of the hepatitis B virus group A genotype identified in isolates from South Africa. J Gen Virol 78(7): 1719-1729.
- LJUNGGRE K (2002) Genetic variability in hepatitis B viruses. J Gen Virol 83: 1267-1280.
- Awan Z, Idrees M, Amin I, Butt S, Afzal S, et al. (2010) Pattern and molecular epidemiology of Hepatitis B virus genotypes circulating in Pakistan. Infect Genet Evol 10(8): 1242-1246.
- Nakano T, Lu L, Hu X, Mizokami M, Orito E, et al. (2001) Characterization of hepatitis B virus genotypes among Yucpa Indians in Venezuela. J Gen Virol 82(2): 359-365.
- Stuyver L, Gendt S, Van G, Zoulim F, Fried M, et al. (2000) A new genotype of hepatitis B virus: complete genome and phylogenetic relatedness. J Gen Virol 81(1): 67-74.
- Magnius LO, Norder H (1995) Subtypes, genotypes and molecular epidemiology of the hepatitis B virus as reflected by sequence variability of the S-gene. Intervirol 38: 24-34.
- Huy TT, Ngoc TT, Abe K (2008) New complex recombinant genotype of hepatitis B virus identified in Vietnam. J Virol 82(11): 5657-5663.
- Tatematsu K, Tanaka Y, Kurbanov F, Sugauchi F, Mano S, et al. (2009) A genetic variant of hepatitis B virus divergent from known human and ape genotypes isolated from a Japanese patient and provisionally assigned to new genotype J. J Virol 83(20): 10538-10547.
- Gandhe SS, Chadha MS, Arankalle VA (2003) Hepatitis B virus genotypes and serotypes in western India: lack of clinical significance. J Med Virol 69(3): 324-330.
- Datta S (2008) An overview of molecular epidemiology of hepatitis B virus (HBV) in India. Virol J 5: 156.
- Zeng G, Wang Z, Wen S, Jiang J, Wang L, et al. (2005) Geographic distribution, virologic and clinical characteristics of hepatitis B virus genotypes in China. J Vir Hepat 12(6): 609-617.
- Abbas N (2001) Variability analysis in immunogenic region of surface gene of Pakistani isolates of Hepatitis B virus (Doctoral dissertation, University of the Punjab, Lahore).
- Idrees M, Khan S, and Riazuddin S (2004) Common genotypes of hepatitis B virus. J Coll Phy Sur Pak 14: 344-347.
- Abbas Z, Muzaffar R, Siddiqui A, Naqvi SA, Rizvi SA (2006) Genetic variability in the precore and core promoter regions of hepatitis B virus strains in Karachi. BMC gastroenterol 6(1): 20.
- Alam MM, Zaidi SZ, Malik SA, Shaukat S, Naeem A, et al. (2007) Molecular epidemiology of Hepatitis B virus genotypes in Pakistan. BMC Infect dis 7(1): 115.
- Nooali S, Hakim ST, McLean D, Kazmi SU, Bagasra O (2008) Prevalence of hepatitis B virus genotype D in females in Karachi. Pakistan. J Infect Developing Countries 2: 373-378.
- Ahmed CS, Wang ZH, Bin Z, Chen JJ, Kamal M, et al. (2009) Hepatitis B virus genotypes, subgenotypes, precore, and basal core promoter mutations in the two largest provinces of Pakistan. J Gastroenterol Hepatol 24(4): 569-573.
- Baig S, Siddiqui A, Chakravarty R, Moatter T (2009) Hepatitis B virus subgenotypes D1 and D3 are prevalent in Pakistan. BMC Res Notes 2(1): 1.
- Hanif M, Zaidi P, Habib S, Ahmed A, Raza A, Murtaza SA, Irfan J (2013) Study of genotypes and subgenotypes of Hepatitis B virus prevalent in big cities of Pakistan. Afric J Microbiol Res 7(3): 152-157.
- Mahmood M, Raza A, Anwar MA, Qayyum M, Zaman N, et al. (2016) Analysis of Complete and Partial Genome Sequences of Hepatitis B Virus and Determination of its Genotypes and Sub-Genotypes from Pakistan. Pak J Zool 48(3): 747-753.
- Mahmood M, Anwar MA, Khanum A, Zaman N, Raza A (2016) Distribution and clinical significance of hepatitis B virus genotypes in Pakistan. BMC Gastroenterol 16: 104.






























