Localization of the RNA Binding Domain of the IHi Bovine Immunodeficiency Virus Rev Protein
Andrea Gomez Corredor1,2, Claude Marchand1and Denis Archambault1*
1Department of Biological Sciences, University of Quebec, Canada
2McGill University Health Centre, Canada
Submission: July 28, 2017; Published: August 21, 2017
*Corresponding author: Denis Archambault, Department of Biological Sciences, University of Quebec, Succursale Centre-ville, P.O. Box 8888, Montreal, Quebec, H3C 3P8, Canada, Tel: [514} 987-3000; Ext. 4622. Fax: (514) 987-4647; Email: archambault.denis@uqam.ca
How to cite this article: Denis A, Claude M, Andrea G C. Localization of the RNA Binding Domain of the Bovine Immunodeficiency Virus Rev Protein. Adv Biotech & Micro. 2017; 5(2): 555664. DOI: 10.19080/AIBM.2017.05.555664
Abstract
The Rev protein is essential for the replication of lentiviruses. Rev is a shuttling protein that transports unspliced and partially spliced lentiviral RNAs from the nucleus to the cytoplasm. The lentiviral Rev proteins contain at least three central functional domains: a basic arginine- rich domain that mediates RNA binding (RBD) and contains the nuclear/nucleolar localization signals (NLS/NoLS), a multimerization domain and a leucine-rich domain that is necessary for the nuclear exportation of Rev. Recently, were reported the presence of novel types of NLS and NoLS in the Bovine immunodeficiency virus (BIV) Rev protein [186 amino acids (aa) in length] that we different from any lentiviral Rev or retroviral Rev-like proteins. Here we report on the RBD activity of BIV Rev. To indirectly map the RBD domain of BIV Rev, the plasmid pDM138-BIV which contains, between the Human immunodeficiency virus type 1 (HIV-1) donor and acceptor splicing sites, the chloramphenicol acetyltransferase (CAT) reporter gene, and the BIV Rev responsive element (RRE). By using previously described BIV Rev mutants, two regions in BIV Rev were identified to mediate RBD binding, indicating a bipartite RBD in BIV Rev. The first RBD motif was found to localize in the middle region of the protein overlapping with the NoLS and the second motif of the bipartite NLS, whereas the second RBD motif mapped in the C-terminal region of BIV Rev. We just concluded that BIV Rev binds the RRE through a bipartite RBD and that this RBD domain is composed of aa 87 to 107 , and aa 154 to 157.
Keywords: Lentivirus; Bovine immunodeficiency virus; Rev Protein; RNA-binding domain; Rev responsive element (RRE)
Introduction
The Lentivirus life replication cycle involves the synthesis of viral RNAs. These RNAs are divided in three groups: fully spliced, partially spliced and unspliced mRNAs. As described in HIV-1, the fully spliced viral mRNAs encode regulatory proteins including Tat and Rev whereas the partially spliced viral mRNAs produce enzymatic and structural proteins. The unspliced RNAs are template for genomic viral RNAs that will be packaged for the production of progeny viruses [1,2]. The fully spliced viral mRNAs are exported from the nucleus to the cytoplasm of the infected cells through standard cellular machinery. In contrast, nuclear exportation of unspliced and partially spliced viral mRNAs is accomplished with the involvement of the HIV-1 Rev protein[1]. Rev forms a multimeric protein complex on the Rev responsive element (RRE), a secondary RNA structure that is present in unspliced and partially spliced viral mRNAs [3].
From studies reported in the Human immunodeficiency virus type 1 (HIV-1), the lentiviral Rev proteins contain at least three central functional domains: a basic arginine-rich domain that mediates RNA binding (RBD) and contains the nuclear/ nucleolar localization signals (NLS/NoLS), a multimerization domain and a leucine-rich domain that is necessary for the nuclear exportation of Rev. HIV-1 Rev contains a short stretch of 17 basic amino acids (aa), mainly arginine, that serves as nuclear/nucleolar localization signals and viral RNA binding domain (RBD) [4]. The Rev RBD binds the RRE mentioned above for viral mRNA nuclear export [5]. The C-terminal region of HIV-1 Rev contains a nuclear exportation signal (NES) which interacts with CRM1 for exportation of the protein-viral mRNA complexes to the cytoplasm [6].
The Bovine immunodeficiency virus (BIV) is a Lentivirus that infects cattle. Although most cattle remain asymptomatic, persistent lymphocytosis, neurological disorders associated with central nervous system lesions, weight loss, diminished milk production and opportunistic bacterial infections have been associated with BIV infection (reviewed in [7,8]. The BIV genome is structurally analogous to that of other lentiviruses and BIV Rev [186 amino acids (aa) in length] is predicted to be functionally similar to HIV-1 Rev [9]. Unlike HIV-1 Rev protein,the functional domains of BIV Rev have not been widely studied. Here, the RBD domain was localized and the aa residues that compose it were identified, indicating a bipartite RBD.
Results and Discussion
RBDs in proteins are generally composed of arginine- rich motifs (ARM) with a small number of other types of aa residues [10]. To map the residues in BIV Rev associated with RBD function, we used a series of previously described mutant proteins in which selected arginine (R) and lysines (K) in BIV Rev WT sequence were replaced by alanine (A) residues [11] (Table 1).
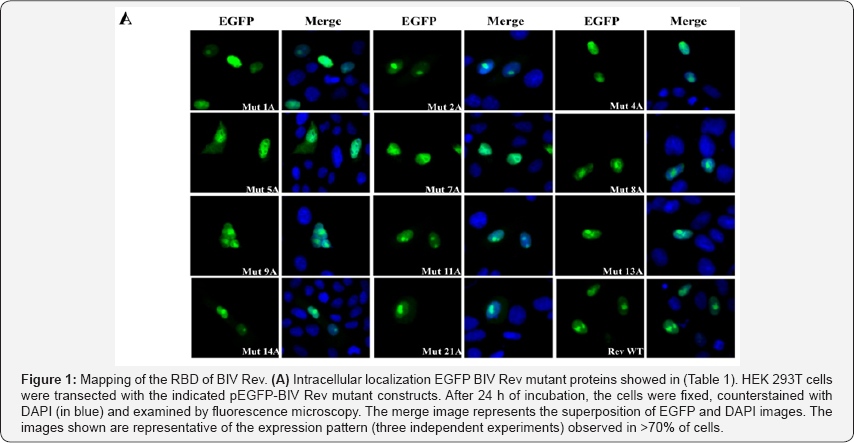
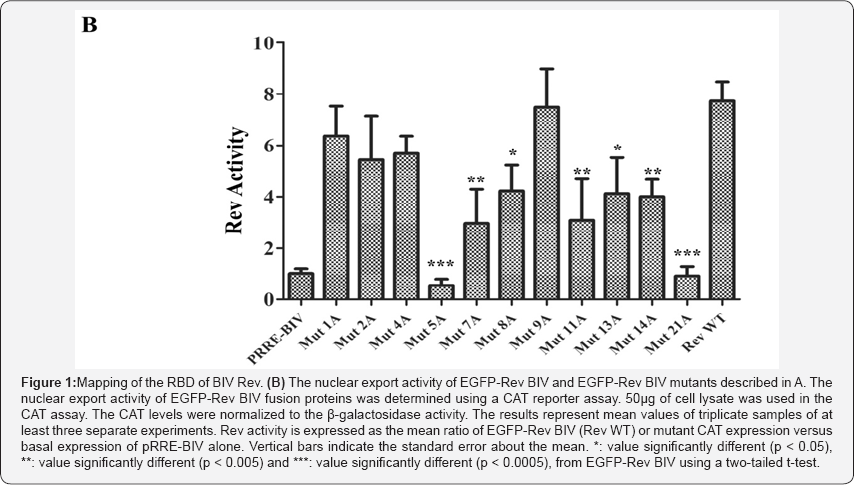
As the mutant proteins were used to map the NLS and NoLS of BIV Rev, we selected in this study only the mutants whose nuclear localization was not impaired [11]. As expected, all selected BIV Rev mutant proteins predominantly enter in the nucleus (Figure 1A). The Rev activity of these mutants was then determined using a CAT reporter assay dependent of the BIV RRE sequence, thereby evaluating both the RNA binding and export capacities of Rev [11]. Among all BIV Rev mutants tested (Figure 1B) Mut 5 and Mut 21 mutants had no activity at all, a finding already reported by us for Mut 5 [11]. Mut proteins 7, 8, 11, 13 and 14 showed a Rev activity which was at least 50% of that of BIV Rev WT. All other mutants tested had an activity comparable to that of BIV Rev WT. Taken together, the regions encompassing aa 87R to 107R (deleted in Mut 5, 8, 9, 11) and 154K to 157R (deleted in Mut 21) are involved in RBD activity of Rev, indicating the presence of a bipartite RBD for BIV Rev.
Viral proteins such as HIV-1 Rev and Tat, HTLV-1 Rex and BIV Tat are typical examples of proteins harboring an ARM that acts as a RBD [10]. In these and in other Rev and Rev-like proteins, RBD is closely associated to the NLS/NoLS [12-14]. These overlapping functions prevent Rev bound to RRE from undesirable return to the nucleus. When Rev binds the RRE, the NLS becomes masked while the NES remains accessible. Here, experiments were conducted to map the RBD of BIV Rev by using a series of mutants harboring alanine scan mutations and an intact NES (Table 1) [11]. All mutant proteins mainly localized to the nucleus (Figure 1A), a prerequisite in our experiment to ensure that the proteins were able to enter into the nucleus for RNA binding. The mutant constructs were then co-transected in cells and analyzed by using in a CAT gene reporter assay that relied on the presence of BIV RRE localized upstream of the CAT-encoding gene [11]. Thus, Rev binding to RNA (RRE) in the nucleus would result in the cytoplasmic transport of the CAT transcript, and, thereof, in CAT expression indicating a Rev activity. By using this assay, two stretches of basic residues localized in the middle and C-terminal regions of BIV Rev were found to mediate RNA binding. In one hand, Mut5 and Mut 21 proteins had no Rev activity at all (Figure 1B). Interestingly, the mutated residue within the Mut 5 protein was part of the BIV Rev NoLS. Mut 7 and Mut 8 proteins (the mutated residues of which were part of the BIV Rev NLS) as well as the Mut 11protein (the mutated residues of which were part of neither BIV Rev NoLS nor NLS) showed a diminished Rev activity. All mutated arginine in the Mut 5, 7, 8 and 11 proteins are localized in the middle region of BIV Rev (87RELVEQNRRKQERRLSGLDRR107). We also found that the integrity of three basic residues localized in the C-terminal region of BIV Rev was necessary for Rev activity. This was shown by the data obtained with the Mut 13 and Mut 14 single mutant proteins that showed a diminished Rev activity (Figure 1B).
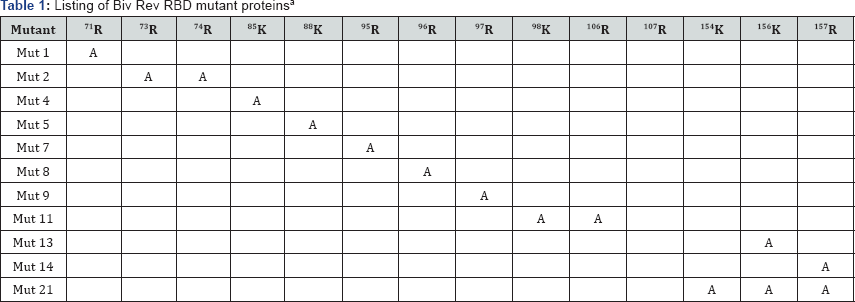
aThe indicated arginine (R) and lysine (K) residues in the BIV Rev sequence were changed to alanine (A) residues, and the numbers indicate the position of residues [11].
These results were confirmed with the use of Mut 21 protein where mutations of 154K-KR157 residues into 154A-AA157 significantly affected the Rev activity. Combined together, these results show that BIV Rev harbors two regions necessary for RNA binding, thereby indicating a bipartite RBD. These finding contrasts with the monopartite nature of the RBD present in HIV- 1 Rev [1,10] but are in agreement with the bipartite RBD present in EIAV Rev [15]. Thus, BIV and EIAV Rev would be the only Rev or Rev-like proteins in retroviruses harboring a bipartite RBD. However, in vitro protein-RNA (RRE) interaction experiments are needed to unequivocally confirm the BIV rev results.
The EIAV Rev RBD is composed of two shorts ARMs. The first ARM is localized in the central region of the protein whereas the second ARM overlaps the NLS sequence in the C-terminal region of the protein [15]. Although the BIV Rev RBD motifs also localize in the central and C-terminal regions of the protein, the aa sequences that compose these motifs differ from those of EIAV Rev RBD. In BIV, the five arginine residues that compose the first ARM are scattered within a 21 aa-long sequence (87RELVEQNRRKQERRLSGLDRR107) overlapping the BIV Rev NoLS/NLS for the first three residues [11]. This motif contrasts with the short RRDRW sequence that composes the first ARM motif of the EIAV Rev RBD [15]. The second motif of BIV Rev RBD is composed of two lysines and one arginine (154KLKR157), and thus does not represent an ARM per se but rather should be only defined as a motif rich in basic residues. This aa composition contrasts with that of the second ARM of the EIAV Rev RBD that is a truly arginine-rich motif (KRRRK) [15].
In summary, we have characterized the RBD domain of BIV Rev that differs from those of HIV-1 and/or EIAV Rev. These results combined with those of our previously published work [11,16] where novel types of NLS and NoLS were found, show that BIV Rev is an atypical protein among Rev or Rev-like proteins in retroviruses.
Acknowledgment
A. Gomez Corredor was supported by a graduate studentship from La Foundation UQAM. C. Marchand was supported by a graduate studentship from the Founds de Recherche Nature et Technologies du Quebec. This work was supported by an operating Discovery grant from the National Sciences and Engineering Research Council of Canada to D. Archambault.
References
- Pollard VW, Malim MH (1998) The HIV-1 Rev Protein. Annu Rev Microbiol 52: 491-532.
- Suhasini M, Reddy TR (2009) Cellular proteins and HIV-1 Rev function. Curr HIV Res 7(1): 91-100.
- Malim MH, Bohnlein S, Fenrick R, Le SY, Maizel JV, et al. (1989) Functional comparison of the Rev trans-activators encoded by different primate immunodeficiency virus species. Proc Natl Acad Sci USA 86(21): 8222-8226.
- Hope T, McDonald D, Huang X, Low J, TG Parslow, et al. (1990) Mutational analysis of the human immunodeficiency virus type 1 Rev Transactivator: essential residues near the amino terminus. J Virol 64(11): 5360-5366.
- Malim MH, Hauber J, Le SY, Maizel V, Cullen BR, et al. (l989) The HIV-l rev trans-activator acts through a structured target sequence to activate nuclear export of unspliced viral mRNA. Nature 338(6212): 254-257.
- Fischer U, Huber J, Boelens WC, Mattaj IW, Luhrmann R, et al. (1995) The HIV1 Rev activation domain is a nuclear export signal that accesses an export pathway used by specific cellular RNAs. Cell 82(3): 475-483.
- St-Louis MC, Cojocariu M, Archambault D (2004) The molecular biology of bovine immunodeficiency virus: a comparison with other lentiviruses. Anim Health Res Rev 5(2): 125-143.
- Corredor AG, St-Louis MC, Archambault D (2010) Molecular and biological aspects of the bovine immunodeficiency virus. Curr HIV Res 8(1): 2-13.
- Oberste MS, Williamson C, Greenwood D, Nagashima K, Copeland D, et al. (1993) Characterization of bovine immunodeficiency virus rev cDNAs and identification and subcellular localization of the Rev protein. J Virol 67(11): 6395-6405.
- Bayer TS, Booth LN, Knudsen SM, Ellington AD (2005) Arginine-rich motifs present multiple interfaces for specifie binding by RNA. RNA 11(12): 1848-1857.
- Gomez Corredor A, Archambault D (2009) The bovine immunodeficiency virus rev protein: identification of a novel lentiviral bipartite nuclear localization signal harboring an atypical spacer sequence. J Virol 83(24): 12842-12853.
- Hope T, Bond BL, McDonald D, Klein NP, Parslow TG, et al. (1991) Effector domains of human immunodeficiency virus type 1 Rev and human T-cell leukemia virus type l Rex are functionally interchangeable and share an essential peptide motif. J Virol 65(11): 6001-6007.
- Yang J, Bogerd HP, Peng S, Wiegand H, Truant R, et al. (1999) An ancient family of human endogenous retroviruses encodes a functional homolog of the HIV-l Rev protein. Proc Natl Acad Sci USA 96(23): 13404-13408.
- Magin C, Hesse J, Lower J, Lower R (2000) Corf, the Rev/Rex homologue of HTDV/HERV-K, encodes an arginine-rich nuclear localization signal that exerts a trans-dominant phenotype when mutated. Virology 274(1): 11-16.
- Lee JH, Murphy SC, Belshan M, Sparks WO, Wannemuehler Y, et al. (2006) Characterization of functional domains of equine infectious anemia virus Rev suggests a bipartite RNA binding domain. J Virol 80(8): 3844-3852.
- Gomez Cooredor A, Archambault D (2012) The bovine immunodeficiency virus Rev protein: identification of a novel nuclear import pathway and nuclear export signal among retroviral Rev/Rev- like proteins. J Virol 86(9): 4892-4905.






























