Enzymes: The Biological Accelerators
Sundeep Kaur Manjal, Ramandeep Kaur, Rohit Bhatia and Ravindra K Rawal*
Department of Pharmaceutical Chemistry, ISF College of Pharmacy, India
Submission: April 06, 2017; Published: May 26,2017
*Corresponding author: Ravindra K. Rawal, Department of Pharmaceutical Chemistry, ISF College of Pharmacy, GT Road Moga, Punjab 142001, India, Tel: 91-9878882520 Email: rawal.ravindra@gmail.com
How to cite this article: Sundeep K M, Ramandeep K, Rohit B & Ravindra K. Rawal. Enzymes: The Biological Accelerators.Organic & Medicinal Chem IJ. 2017; 2(4): 555594. DOI: 10.19080/OMCIJ.2016.01.555594
Abstract
Enzymes are the macromolecular biological catalysts which tend to exhibit tremendous biological value for the human society. These are known to accelerate and catalyze the chemical reactions many times faster than ordinary. Generally, they are known to catalyze more than 5,000 types of biochemical reactions. Numerous enzymes are produced inside the human body which tend to play crucial role in the functioning of biological activities. Some of the enzymes are used commercially such as for the synthesis of antibiotics and also used for household purpose like in manufacturing of washing powder. Apart from this, enzymes serve a variety of functions inside the living organisms. Recently pro-drug approach has gained wide popularity, it is mainly utilised in the lead optimization of the drug molecule. In this review we have tried to discuss different aspects of enzymes, enzyme kinetics, enzyme inhibitors, pro-drugs and their utility with suitable examples.
Keywords: Enzyme; Serine; Trypsinogen; Heme; Pro-drug
Introduction
Enzymes act as catalysts for almost all of the chemical reactions that occur in all living organisms [1]. They require mild conditions for the enzymatic action. They have a large efficiency because only minute amount of enzyme is required in the production of many biotechnology products like food, beverages, paper and textile industries, clothing, etc [2] (Figure 1).
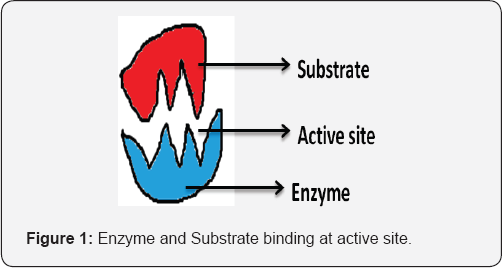
The structures of a number of enzymes contain groups of metal ions, known as clusters, coordinated to the peptide chain. These enzymes are often known as metalloenzymes. Enzymes with transition metal cofactors are named metalloenzymes. Metalloenzymes can be divided on the basis of presence or absence of heme and known as heme and non heme metalloenzyme. Metalloenzyme show many functions and this function are determined by four structural features that are - the metal core, the metal binding motif, the second sphere residues in the active site and the electronic statistics [3]. Enzymes which require non-protein components to get functionally active, it is generally accepted and known as haloenzyme, which refers to the complete catalytic entity which consists of a protein part known as the apoenzyme and a non-protein constituent. Cofactors are those small organic or inorganic molecules that the enzyme requires for the activity. The prosthetic group is similar to the cofactor but is tightly bound to the apoenzyme. A coenzyme is defined as a molecule that possesses physicochemical properties not found in the polypeptide chain of the enzyme and that acts together with the enzyme to catalyze biochemical reaction. Tightly bound coenzymes are sometimes referred to as prosthetic groups [4] (Figure 2).

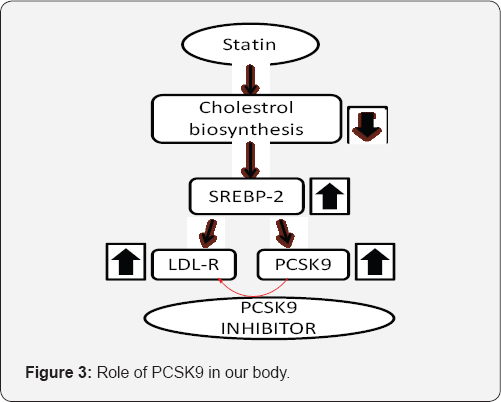
A number of enzymes are produced by the body from inactive protein precursors, known as proenzymes/zymogens. For example, the active form of enzyme trypsin is produced from the proenzyme trypsinogen by the loss of a 6-amino acid residue chain from the N-terminal of trypsinogen. This loss is accompanied by a change of conformation of the resultant protein to form the active form of the enzyme. Proenzymes allow the body to produce and control enzymes that could be harmful in the wrong place. E.g., trypsin catalyzes the breakdown of proteins [5]. There are many other examples which show role of proenzyme or zymogen in body. Tibolla Norata etal. [6], describe that PCSK9 is a secretory serine proteinase. Pro-PCSK9 is a 73 kDa zymogen and secreted as a 63 kDa mature protein. Then the secreted PCSK9 binds specifically to the epidermal growth factor (EGF) of the LDLR at the cell surface. As it has direct binding to and degradation of LDLR, PCSK9 is regarded as a valid and novel target for the treatment of hypercholesterolemia. PCSK9 exhibit higher levels of liver LDLR and reduced serum cholesterol, while over expression of PCSK9 reduces LDLR and increases serum cholesterol. So, those individuals with loss of function mutations of PCSK9 have lower levels of LDL cholesterol and are protected from cardiovascular diseases. PCSK9 is one of the best genetically validated targets for treatment of heart disease (Figure 3).
Inoue et al. [7] described about the proenzyme of the protein kinase. The proteolysis was done by calcium-dependent protease occurring in the tissue, the proenzyme was converted to an active protein kinase. The proenzyme and protease were separated by DEAE-cellulose (DE52). Then differently the proenzyme was resolved into two components. These components showed a sedimentation coefficient. But proenzyme showed no glycogen phosphorylase kinase activity but was always associated with activity to phosphorylate protamine. Ca2+-dependent protease was also purified. The protease showed a sedimentation coefficient and Ca2+ being the most active and the maximum activity was obtained.
The active protein kinase thus produced from the proenzyme in vitro. Although these values were slightly different from those of the protein kinase which was obtained from bovine cerebellum stored frozen. The active protein kinases were indistinguishable from each other. So it was clear that such proenzymes and proteases were distributed in many other tissues including lungs, liver, kidney, cerebellum, heart, skeletal muscle, and adipose tissue. Guroff described that many proteases act on casein, oxidized ribonuclease, and the oxidized β-chain of insulin [8]. In muscle, Ca2+-dependent neutral protease has been described by Meyer who acts on muscle glycogen phosphorylase kinase. The protease has act not only on muscle glycogen phosphorylase kinase but also on casein, muscle glycogen synthetase, and many other components of myofibrils such as Z-discs, troponin, tropomyosin, and C-protein [9-13]. Abita et al. [14] describe the transformation of prophospholipase A into phospholipase A is triggered by the tryptic hydrolysis of the Arg,-Ma, and bond of the zymogen. This bond is a better substrate for trypsin as compared to others of other zymogens such as trypsinogen and chymotrypsinogen. Calcium ions do not play any role in prophospholipase activation. The physicochemical analysis of the zymogen and of the enzyme shows that activation produces only very little changes in the overall folding of prophospholipase. At N-terminal end of the precursor rearrangement take place. The conformational modification was at side-chain of tryptophan-3 (in phospholipase sequence) which passes from a polar to an apolar environment during activation. The α-amino group of alanine-1 (in the phospholipase sequence) which appears on activation is essential for enzyme activity. Selective chemical modification of function by a number of reagents abolishes phospholipase activity. By analogy with the chymotrypsinogen-chymotrypsin and the proelastase-elastase transformations, it was proposed that the a-amino group was involved in the formation of a salt-bridge which stabilizes the adequate geometry of the active site (Figure 4).
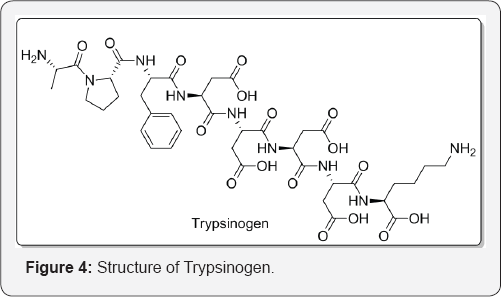
Enzymes are usually isolated from different tissues but can be found in the same tissue and have same catalytic activity, therefore are referred to as isoenzyme/isozymes. Isoenzyme structures often contain the same protein subunits but either in a different order or ratio. For example LDH catalysing the conversion of lactate to pyruvate and vice versa, is a tetramer consisting of 4 H and M subunits. The ratio of H and M subunits in the enzyme will depend on its source. LDH isolated from heart contains mainly H subunits whilst that from liver and skeletal muscle contains M subunit. Isoenzymes apart from catalysing the same reaction can also exhibit thermal stability, electrophoretic mobolities, the effect of inhibitors, and also as a diagnostic aid (H type LDH) in the blood for indication of heart attack, since heart attacks cause the death of heart muscle with a subsequent release of H type LDH in the circulatory system [15] (Figure 5).
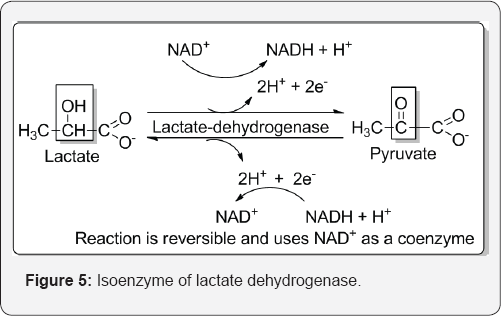
Houddane et al. [16] described that proliferating cells depend on glycolysis mainly to supply precursors for macromolecular synthesis. Fructose-2, 6-bisphosphate was the most potent positive allosteric effectors of 6-phosphofructo-1-kinase. Mitogen stimulation of thymocytes with concanavalin A (ConA) led to increases in lactate accumulation Fru-2,6-P2 content, 6-phosphofructo-2-kinase (PFK-2)/fructose-2,6-bisphosphatase isoenzyme 3 and 4 (PFKFB3 and PFKFB4) protein levels and cell proliferation and protein synthesis. This support a role for Fru-2,6-P2 in coupling glycolysis to cell proliferation and protein. Annesley et al. [17] describe that thermally injured rats reveals an isoenzyme profile for lactate dehydrogenase that was compatible with origin from lung.
The appearance of LDH-4 isoenzyme in serum of thermally injured rats correlates linearly with indices of lung damage, supporting the results that thermal trauma to the skin can cause oxygen radical production by complement-activated blood neutrophils with resultant acute microvascular injury in the lung interstitium. Furthermore, interventions that protect from oxidant-mediated lung injury (catalase, scavengers of hydroxyl radical, iron chelators or neutrophil depletion) result in significant reductions in serum levels of the LDH-4 isoenzyme following thermal injury to the skin. Thus, measurements of LDH isoenzyme patterns in serum appear to be useful in monitoring tissue damage such as oxygen radical-mediated acute lung injury [17]. Enzymes that catalyse the same reactions but have significantly different structures are known as isofunctional enzymes.
Isofunctional enzymes are produced by different tissues and different organisms. Their different origins and structures mean that they can act as selective targets for drugs. Example: the enzyme DHFR catalyses the interconversion of FAH2 to FAH4 in both humans and bacteria. Trimethoprim is more effective in inhibiting this conversion in bacteria than in humans, which makes it useful for the treatment of bacterial infections in humans. Variation in the enzymes structures can also occur between and within ethnic groups of the same species. For example, a number of different dehydrogenase isoenzymes have been observed in some groups of Asians [18] (Figure 6).

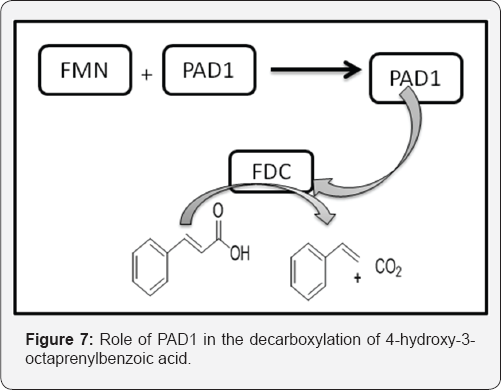
Lin et al. [19] described that the decarboxylation of antimicrobial aromatic acids such as phenylacrylic acid (cinnamic acid) and ferulic acid by yeast requires two enzymes described as phenylacrylic acid decarboxylase (PAD1) and ferulic acid decarboxylase (FDC). These enzymes are of interest for various biotechnological applications, such as the production of chemical feed stocks from lignin under mild conditions. The PAD1 and FDC were both examined. The PAD1 was a flavin mononucleotide (FMN)-containing protein. PAD1 catalyzes the formation of a novel, diffusible cofactor required by FDC for decarboxylase activity. Co-expression of FDC and PAD1 results in the production of FDC with high levels cofactor bound. Holo-FDC catalyzes the decarboxylation of phenylacrylic acid, coumaric acid and ferulic acid. The E. coli enzymes UbiX and UbiD are related by sequence to PAD1 and FDC respectively and are involved in the decarboxylation of 4-hydroxy-3-octaprenylbenzoic acid, an intermediate in ubiquinone biosynthesis. This implies that the same cofactor was required for decarboxylation of 4-hydroxy- 3-polyprenylbenzoic acid by UbiD and suggests a wider role for this cofactor in metabolism (Figure 7).
Enzyme Kinetics
Enzyme kinetics it is main fundamental tool to know about cellular metabolism and for synthetic biology applications. From in vitro enzyme assay was done to examine enzyme characterization. Kinetic parameters are only available for less of reactions, and even this data tells less limits of predictive power of metabolic models. Nowadays quantitative proteomics was in-sighted into in vivo enzyme kinetics. They studied between in vivo and in vitro enzyme catalysis and proteomics can be used to characterize the efficiency of enzyme utilization across conditions [20].
Single substrate reactions
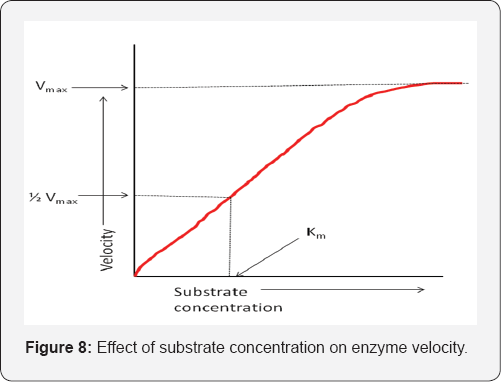
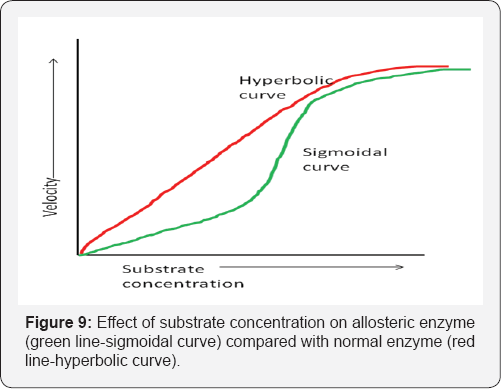
In these reactions a single substrate (S) is converted by the enzyme (E) to the products (P) (Figure 8). The mathematical relationship between the rate of the enzyme catalysed reaction and the conc. of the single substrate depends on the nature of the enzyme process. Many processes exhibit 1st order kinetics at low conc. of the substrate, which changes to 0 order as the concentration increases, that is, a hyperbolic relationship between rate and substrate conc. Processes under allosteric control (inhibition or activation of an enzyme by a small regulatory molecule that interacts at a site (allosteric site) other than the active site (at which catalytic activity occurs) have a sigmoid relationship between reaction rate and the substrate concentration. This suggests 2nd and higher orders of reaction, that is, the rate is proportional to [S]n where n>1 [21] (Figure 9).
The kinetics of enzyme processes with a single substrate that exhibit a hyperbolic rate/substrate concentration relationship may often be described by the Michaelis-Menten equation [22].
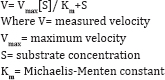
Obach and Reed-Hagen et al. [23] described that the Michaelis constant (Km) for cytochrome P 450-mediated drug biotransformation reactions is an important parameter in understanding the potential for a drug to exhibit saturable metabolism in both in vivo and non-linear dose-exposure relationships. The determinations were made using an approach of monitoring substrate loss at multiple substrate concentrations, withthe objective of comparing KM values. The reactions examined were CYP2C9-catalyzed diclofenac 4-hydroxylation, CYP2D6- catalyzed dextromethorphan O-demethylation and thioridazine S-oxidation, CYP2C19-catalyzed imipramine N-demethylation, CYP3A4-catalyzed midazolam 1'-hydroxylation, and CYP1A2- catalyzed tacrine 1-hydroxylation (Figure 10).

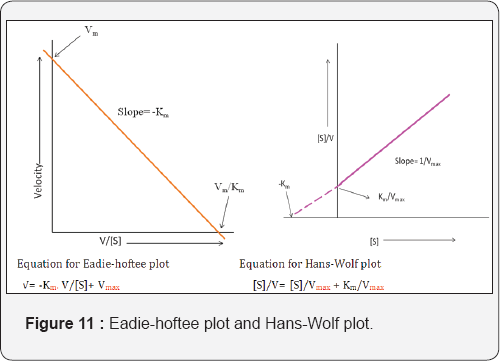
Various rearrangements of the Michaelis-Menten equation have been developed to convert equation into a straight line format. Popular rearrangements are the Eadie-Hofstee and Hanes-Wolf plot that are used to obtain accurate values of Km and Vmax. The Eadie-Hofstee plot is particularly useful for detecting deviations from a straight line, i.e., deviations from Michaelis- Menton kinetics. These deviations may indicate that the enzyme is under feedback control. The product of the final reaction in that pathway reacts with an enzyme somewhere along the pathway at the enzyme's allosteric site, changing the conformation of the enzyme. That enzyme can no longer binds to its substrate as effectively due to the conformational change, closing down that pathway and stopping the final product from synthesizing [24] (Figure 11).
Multiple substrate reactions
Enzymes often catalyse reactions that involve more than 1 substrate. The kinetics of these reactions will depend on the number of substrates and the general mechanism of enzyme action. Example: Reactions involving A and B substrate (bisubstrate reactions) often proceed by one of the 2 general routes (Figure 12).
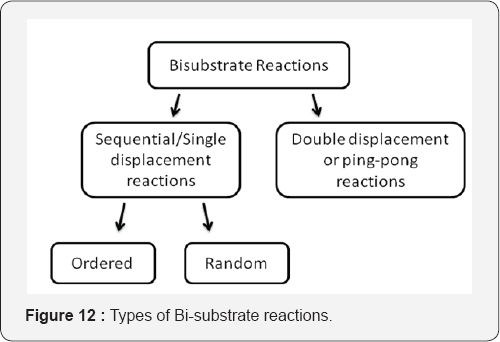
I. The sequential/Single displacement reactions:These reactions can follow 2 routes. 1st case: substrates A and B bind to the enzyme that has no effect (random order) 2nd case: leading substrate can bind to the enzyme before the other substrate can bind to the enzyme (ordered process). Once both substrates are bound to the enzyme, reaction to the products occurs [25].
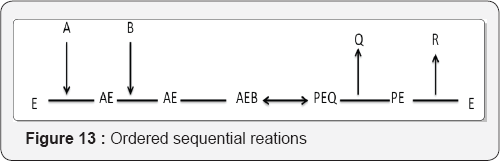
A. Ordered Sequential reactions: Here the leading substrate binds first, followed by the other substrate (Figure 13).
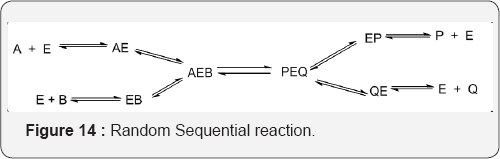
B. Random Sequential reaction: Where either any of the substrate may bind first followed by the other substarte (Figure 14).
Both of these types of sequential reaction exhibit the characteristic type of Line weaver-Burk plot, when the conc. of 1 of the substrates and the enzyme are kept constant and the conc. of the other substrate is varied (Figure 15).
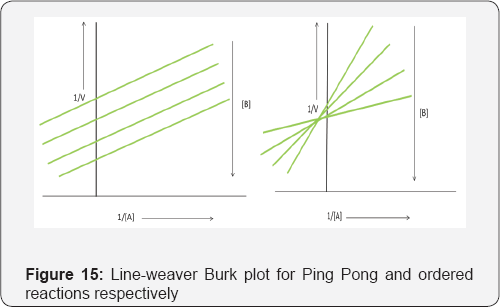
II. Double displacement or ping-pong reactions: A substrate binds to the enzyme and forms a complex, which forms a product P and a modified form of the enzyme. At this point the 2nd substrate binds to the modified enzyme and reacts to form a 2nd product and regenerate the original form of the enzyme [26].
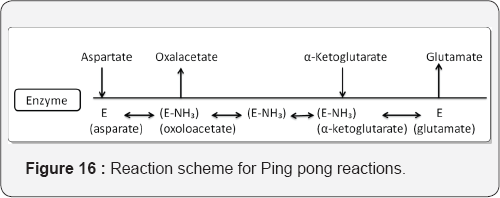
A. Example: Reaction between the amino group between amino acids and α-keto acids (Figure 16).
Acid � Base Catalysis
In any reaction where proton transfer occurs, general acid catalysis and/or general base catalysis can be an important mechanism for specificity and rate enhancement. There are 2 kinds of acid-base catalysis.
Specific catalysis
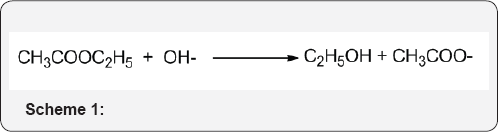
If catalysis occurs by a hydronium ion or hydroxide ion and is determined only by the pH, not the buffer concentration, it is referred to as specific acid or specific base catalysis. Example: The hydrolysis of ethyl acetate is an exceedingly slow reaction at neutral pH because both the nucleophile (H2O) and the electrophile (the carbonyl group) of ethyl acetate have low reactivity (Scheme 1). The reaction rate could be accelerated, however, if the reactivity of either the nucleophile or the electrophile could be enhanced. An increase in the pH increases the concentration of hydroxide ion, which is a much better nucleophile than is water, and, in fact, the rate of hydrolysis at higher pH increases. Likewise, a decrease in the pH increases the concentration of the hydronium ion, which can protonate the ester carbonyl, thereby increasing its electrophilicity, and this also increases the hydrolysis rate. That being the case, then the hydrolysis rate should be doubly increased if base and acid are added together, right? of course not. Addition of an acid to a base would only lead to neutralization and loss of any catalytic effect [27](Figure 17) .
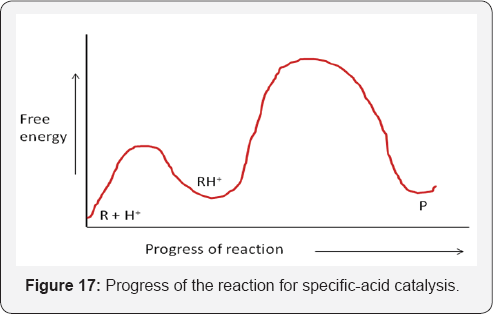
Unlike reactions in solution, however, an enzyme can utilize acid and base catalysis simultaneously for even greater catalysis. The protonated base is either an acidic amino acid side-chain or the basic side chain in the protonated form and the free base is either a basic residue or an acidic residue in the deprotonated form. It is important to appreciate the fact that the pKa values of amino acid side-chain groups within the active site of enzymes are not necessarily the same as those measured in solution. Also, pKa values can change drastically in hydrophobic environments.
General acid base catalysis
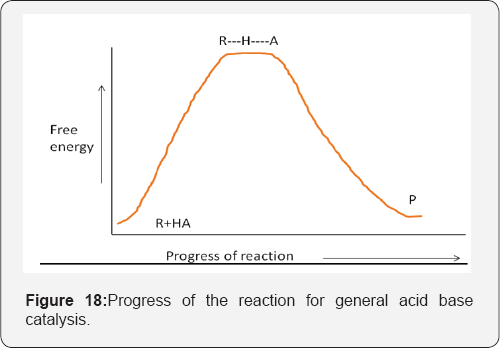
General acid catalysis occurs when acids other than hydronium ion accelerate the reaction rate. Similarly, general base catalysis occurs when bases other than hydroxide ion accelerate the rate. In solution, general acid-base catalysis can be demonstrated when the reaction rate increases with increasing buffer concentration, at a constant pH and ionic strength, and shows a larger increase with a buffer that contains a more concentrated acid or base component. Because the hydronium or hydroxide ion concentration is not increasing (the pH is constant), it must be the buffer that is catalyzing the reaction. In enzyme mediated catalysis, general acid-base catalysis occurs when an acidic or basic residue at the active site is used to facilitate proton transfers in the reaction [28] (Figure 18).
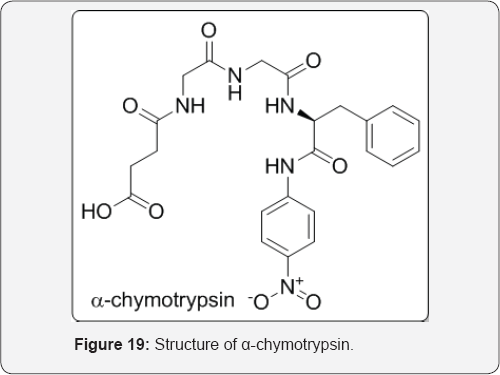
As an example of general acid-base catalysis, consider the enzyme α-chymotrypsin, a member of serine protease family (Figure 19). The general accepted mechanism of action of this enzyme, which features nucleophilic attack by serine hydroxyl
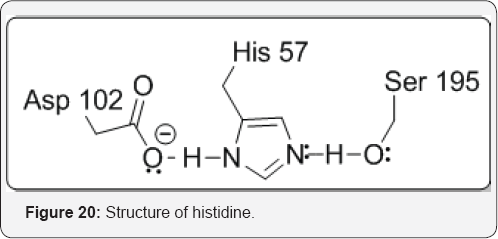
group onto the carbonyl group of a peptide bond. Although the serine hydroxyl group is not normally regarded as a potent nucleophile, aspartic acid and histidine residues nearby have been implicated in the conversion of serine to an alkoxide by a mechanism called the charge relay system by Blow and co-workers, the discoverers of the existence of the hydrogen bonding network involving Asp-102, His-57, and Ser-195. This catalytic triad involves the aspartate carboxylate (pKa of the acid is 3.9 in solution) removing a proton from the histidine imidazole (pKa 6.1 in solution), which, in turn, removes a proton from the serine hydroxyl group (pKa 14 in solution). Now, how can a base such as aspartate, whose conjugate acid is 2 pKa units lower than that of the histidine imidazole, remove the imidazole proton efficiently. The equilibrium is only 1% in favor of the 1st proton transfer, and the equilibrium for the 2nd proton transfer favors the back direction by a factor of 108. One explanation could be that the pKa values of some of these acids and bases at the active site are different from those in solution. Furthermore, a full deprotonation is likely not occurring ; rather, because these groups are held close together at the active site, as the proton is beginning to be removed from the serine hydroxyl group, the charge density proceeds to the next step (attack of alkoxide at the peptide carbonyl), thereby driving the equilibrium in the forward direction. This is the beauty of enzyme-catalyzed reactions; the approximation of the groups and the fluidity of the active site residues working in concert permit reactions to occur that would be nearly impossible in solution [29] (Figure 20).
Covalent catalysis
Some enzymes can use nucleophilic amino acid side-chains or cofactors in the active site to form covalent bonds to the substrate; in some cases a 2nd substrate then can react with this enzyme substrate intermediate to generate the product. This is known as nucleophilic catalysis, a subclass of covalent catalysis that involves covalent bond formation as a result of attack by an enzyme nucleophile at an electrophilic site on the substrate. For example, if Y is an amino acid or peptide and Z- is a hydroxide ion, then the enzyme would be a peptidase or protease. For nucleophilic catalysis to be most effective, Y should be converted into a better leaving group than X (for example, by protonation), and the covalent intermediate should be more reactive than the substrate. The most common active site nucleophiles are the thiol group of cysteine, the hydroxyl group of serine, the imidazole of histidine, the amino group of lysine, and the carboxylate group of aspartate or glutamate.
These active site nucleophiles are generally activated by deprotonation, often by a neighbouring histidine imidazole or by a water molecule that is deprotonated in a general base reaction. The principal catalytic advantage of using an active site residue instead of water directly is that the former leads to a unimolecular reaction (because the substrate is bound to the enzyme, attack by the serine residue is equivalent to an intramolecular reaction), which is entropically favored over the bimolecular reaction with water. Also, alkoxides (ionized serine) and thiolates (ionized cysteine) are better nucleophiles than hydroxide ions. Amide bonds of peptides have low reactivity, but the nucleophile in the active site could be made more nucleophilic in a nonpolar environment. Nucleophilic catalysis is the enzymatic analogy to anchimeric assistance. Anchimeric assistance is the process by which a neighboring functional group assists in the expulsion of a leaving group by intermediate covalent bond formation. This results in accelerated reaction rates. Typical enzymatic reactions where nucleophilic catalysis is important include the proteolytic enzymes, for example, the serine proteases (proteases that utilizes a serine residue at the active site as the nucleophile) such as elastase (degrades elastin, a connective tissue prevalent in the lung) or plasmin (lyses blood clot), and the cysteine proteases (utilize an active site cysteine residue as the nucleophile) such as papain (found in papaya fruit and used in digestion) [30].
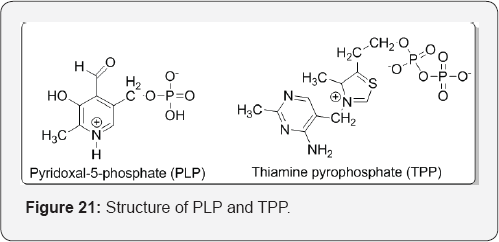
Pratt et al. [31] described that some enzymes use nonamino acid cofactors such as pyridoxal phosphate (PLP) or thiamine pyrophosphate (TPP) to form covalent intermediates. Such covalent intermediates function to reduce the energy of later transition states, similar to how covalent intermediates formed with active site amino acid residues allow stabilization, but the capabilities of cofactors allow enzymes to carryout reactions that amino acid side residues alone could not. Enzymes utilizing cofactors include the PLP-dependent enzyme aspartate transaminase and the TPP-dependent enzyme pyruvate dehydrogenase (Figure 21).
Coenzyme catalysis
A coenzyme, or cofactor, is any organic molecule or metal ion that is essential for the catalytic action of the enzyme. The usual organic coenzymes are generally derived as products of the metabolism of vitamins. Other organic molecules that are involved in essential enzyme functioning, but which are not derived from vitamins, include coenzyme A, which is used to make the CoA thioesters of carboxylic acid substrates; heme (protoporphyrin IX) and the tripeptide glutathione, which are very important to enzymes involved in drug metabolism; adenosine triphosphate (ATP), which supplies the energy required to activate certain substrates during enzyme-catalyzed reactions; and lipoic acid and ascorbic acid (Vit. C), which are involved in oxidation and reduction systems, respectively [32] (Figure 22).
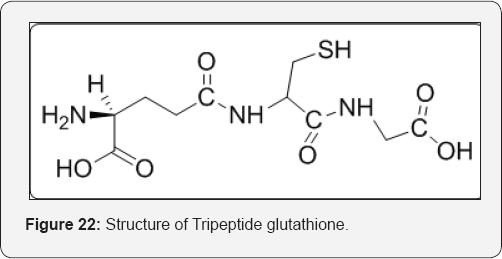
Vitamins are by definition, essential nutrients; human metabolism is incapable of producing them. Deficiency in a vitamin results in the shutting down of the catalytic activity of various enzymes that require the coenzyme made from the vitamin or results from the lack of other activities of the vitamins. This leads to certain disease states.
Chowdhury and Banerjee et al. [33] described that structures of cobalamin-dependent methionine synthase and methylmalonyl-CoA mutase have revealed a striking conformational change that accompanies cofactor binding to the proteins. Alkylcobalamins have octahedral geometry in solution at physiological pH, and the lower axial coordination position is occupied by the nucleotide, dimethylbenzimidazole ribose phosphate, that is attached to one of the pyrrole rings of the corrin macrocycle via an aminopropanol moiety. The active sites of these two B12-dependent enzymes, the nucleotide tail is held in an extended conformation in which the base is far removed from the cobalt in cobalamin. Instead, a histidine residue donated by the protein replaces the displaced intramolecular base.
The two truncated cofactor analogues: adenosylcobinamide and adenosylcobinamide phosphate methyl ester, lacking the nucleotide and nucleoside moieties are studied and revealed that the nucleotide tail has a modest effect on the strength of cofactor binding. The nucleotide has a profound influence on organizing the active site for catalysis, as evidenced by the retention of the base off conformation in the truncated cofactor analogues bound to the mutase and by their inability to support catalysis. In contrast, the truncated cofactors associate very rapidly with the enzyme at rates that are too fast to measure. So it was concluded that a model in which the base-on to base-off conformational change is slow and is assisted by the enzyme, and is followed by a rapid docking of the cofactor in the active site [33] (Figure 23).
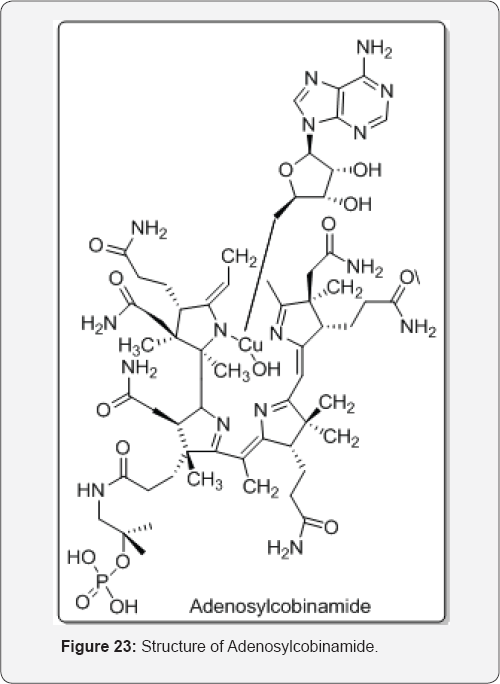
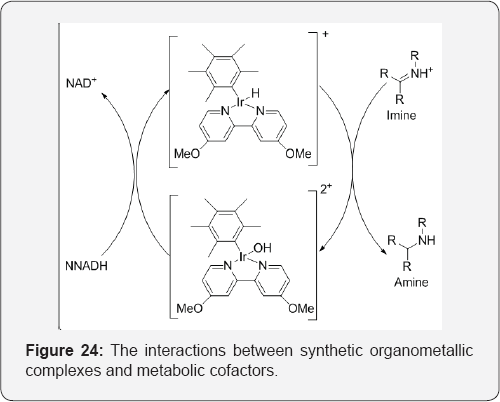
The interactions between synthetic organometallic complexes and metabolic cofactors proved to be a newly emerging topic in bio-organometallic chemistry. The first cationic Cp*lr- catalyzed imine reduction in neutral buffered aqueous medium was reported. The reaction proceeds via a hydride transfer from NADH as the hydride source at room temperature and under air. Cationic Cp*lr complexes proved to be the most efficient catalysts for this transformation. The method was subsequently applied on cyclic and non-cyclic imines. It was extended to one example of reductive alkylation of an amine [34] (Figure 24).
Experimental and clinical investigations reveal that vitamin-B deficiency (folate—vitamin B9, vitamin B6 or vitamin B12) contributes to the risk of neurodegeneration because of an increase in homocysteine. Vitamins are crucial cofactors in the metabolism of carbohydrates, fat and proteins, and vitamin deficiency is associated with an increased risk for Parkinson's disease (PD).4,5 Low intake of vitamin B6, but not of folate (B9), vitamin B12 or riboflavin (B2), was independently associated with an increased risk for PD. Sufficient dietary vitamin B6 intake decreases the risk for PD, probably through mechanisms unrelated to homocysteine metabolism [35].
Heme
Heme, or protoporphyrin IX, is an iron (III)-containing porphyrin cofactor for a large number of liver microsomal mixed function oxygenases principally in the cytochrome P450 family of enzymes. These enzymes, like flavin monooxygenases, are important in the metabolism of xenobiotics, including drugs. As in the case of the flavin monooxygenases, molecular oxygen binds to the heme cofactor (after reduction of the Fe3+ to Fe2+), and is converted into a reactive form that is used in a variety of oxygenation reactions, especially hydroxylation and epoxidation reactions. The hydroxylation reactions often occur at seemingly inactivated C atoms. In heme, the peripheral nitrogens represent the 4 pyrrole nitrogens. The axial ligands in the case of CYP450 are a cysteine thiolate from the protein and water. The electrons for reduction of the heme of CYP-450 (the 2nd and 4th steps of 4.35) come from an enzyme complexed with CYP450 called NADPH-cytochrome P450 reductase, which contains NADPH and 2 different flavin coenzymes (FAD and FMN) [36] (Figure 25).
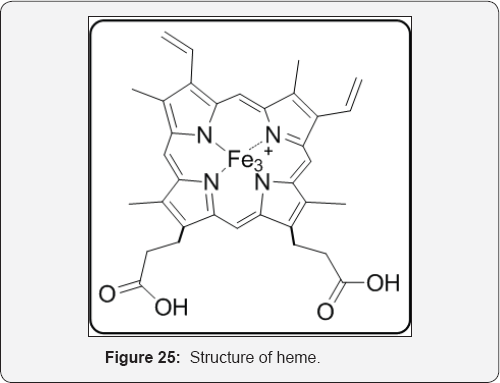
Heme is an essential molecule found in many tissues where it plays key roles as the prosthetic group of several proteins involved in vital physiological and metabolic processes such as gas and electron transport. Structurally, heme is a tetrapyrrole ring containing an atom of iron (Fe) in its center. When released into the extracellular milieu, heme exerts several deleterious effects, which make it an important player in infectious and noninfectious hemolytic diseases where large amounts of free heme are observed such as malaria, dengue fever, β-thalassemia, sickle cell disease and ischemia-reperfusion. The heme degradation by HMOX1/HO-1 (heme oxygenase 1) is required and that Fe is essential for the formation of ALIS, as heme analogs lacking the central atom of Fe are not able to induce these structures. ALIS formation is also observed in vivo, in a model of phenylhydrazine (PHZ)-induced hemolysis, indicating that it is an integral part of the host response to excessive free heme and that it may play a role in cellular homeostasis [37].
Flavin-containing monooxygenases (FMOs) also metabolize foreign chemicals, including drugs, pesticides and dietary components. The mechanism of action of FMOs and insights gained from the structure of yeast FMO. The three FMOs (FMOs 1, 2 and 3) that is most important for metabolism of foreign chemicals in humans, focusing on the role of the FMOs and their genetic variants in disease and drug response. Loss-of-function mutations of FMO3 cause the disorder trimethylaminuria [38]. Flavin-containing monooxygenases (FMOs) also catalyze NADPH- dependent monooxygenation of soft-nucleophilic nitrogen, sulfur, and phosphorous atoms contained within various drugs, pesticides, and xenobiotics. Flavin-containing monooxygenase 3 (FMO3) is responsible for the majority of FMO-mediated xenobiotic metabolism in the adult human liver. Mutations in the FMO3 gene can result in defective trimethylamine (TMA) N-oxygenation, which gives rise to the disorder known as trimethylaminuria (TMAU) or ''fish-odour syndrome''[39].
Enzyme Inhibitors
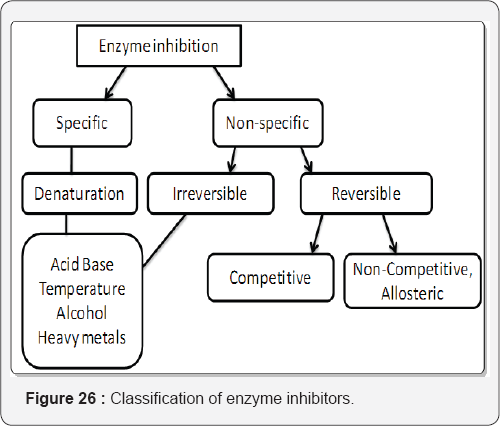
Enzyme inhibitors play an important role in our body As mainly all the drugs act through this mechanism example: The active site of angiotensin-converting enzyme, based on known chemical and kinetic properties of the enzyme, has enabled us to design a new class of potent and specific inhibitors. These compounds, carboxy alkanoyl and mercapto alkanoyl derivatives of proline, inhibit the contractile response of guinea pig ileal strip to angiotensin I and augment its response to bradykinin. When administered orally to rats, these agents inhibit the pressor effect of angiotensin I, augment the vasodepressor effect of bradykinin, and lower blood pressure in model of reno vascular hypertension [40] (Figure 26).
Reversible inhibitors
These are inhibitors that tend to bind to an enzyme by electrostatic bonds, hydrogen bonds and van der Waals forces and so tend to form a dynamic equilibrium system with the enzyme (exception: A few reversible inhibitors bind by weak covalent bonds). These are normally time dependent because the removal of unbound inhibitor from the vicinity of its site of action will disturb this equilibrium. This allows more enzyme sites to become available, which decreases the inhibition of the process, hence it is a time dependent process.
The neutrophil enzyme myeloperoxidase (MPO) promotes oxidative stress in numerous inflammatory pathologies by producing hypohalous acids. Its inadvertent activity is a prime target for pharmacological control. Previously, salicylhydroxamic acid (SHA) was reported to be a weak reversible inhibitor of MPO. We aimed to identify related hydroxamates that are good inhibitors of the enzyme. We report on three hydroxamates as the first potent reversible inhibitors of MPO.
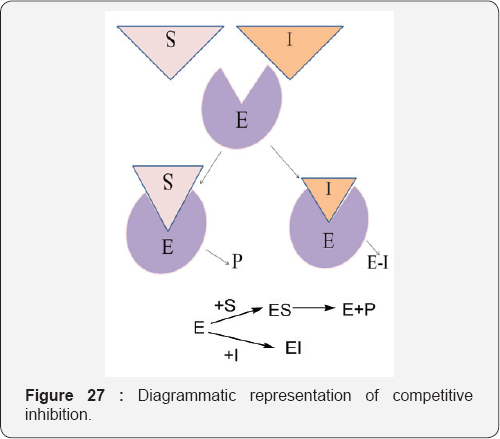
The chlorination activity of purified MPO was inhibited by 50% by 5 nM of a trifluoromethyl-substituted aromatic hydroxamate, HX1. The hydroxamates were specific for MPO in neutrophils and more potent toward MPO compared to a broad range of redox enzymes and alternative targets. Surface plasmon resonance measurements showed the strength of binding of hydroxamates to MPO correlated with the degree of enzyme inhibition. The crystal structure of MPO-HX1 revealed the inhibitor was bound within the active site cavity above the heme and blocked the substrate channel. HX1 was a mixed-type inhibitor of the halogenation activity of MPO with respect to both hydrogen peroxide and halide. Spectral analyses demonstrated that hydroxamates can act variably as substrates for MPO and convert the enzyme to a nitrosyl ferrous intermediate. This property was unrelated to their ability to inhibit MPO. We propose that aromatic hydroxamates bind tightly to the active site of MPO and prevent it from producing hypohalous acids. This mode of reversible inhibition has potential for blocking the activity of MPO and limiting oxidative stress during inflammation [41] (Figure 27).
Competitive inhibition: The inhibitor binds by a reversible process to the same active site of the enzyme as the substrate. This competition between the inhibitor and substrate for the same site results in some enzyme molecules being inhibited whilst others function as normal, resulting in an overall reduction in the rate of conversion of the substrate to the products. Since the binding of the inhibitor is reversible, its effect will be reduced as the concentration of the substrate increases until, at high substrate concentrations, the inhibitory effect is completely prevented. This allows the Vmaxfor the process to be reached.However, it also follows that as the concentration of the inhibitor increases its inhibiting effect increases.
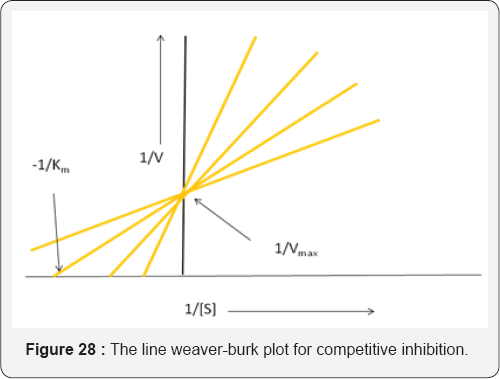
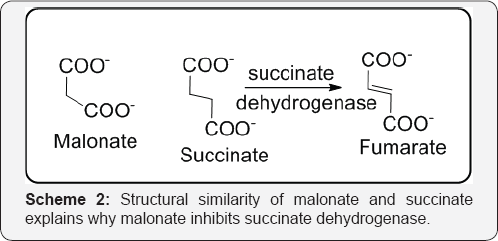
Consequently, for an inhibitor to be effective a relatively high concentration has to be maintained in the region of the enzyme. For single substrate enzyme processes exhibiting simple Michaelis-Menten kinetics the Line weaver-Burk plots for competitive inhibition show characteristic changes when different concentrations of the inhibitor (I) are used. These changes do not affect the value of Vmax but can be used to diagnose the possible presence of a competitive inhibitor [42] (Figure 28). Since the substrate and inhibitor compete for the same active site it follows that they will probably be structurally similar. For e.g., succinate dehydrogenase, which catalyses the conversion of succinate to fumarate, is inhibited by malonate, which has a similar structure to succinate [43]( Scheme 2).
Non-competitive inhibition: Non-competitive inhibitors bind reversibly to an allosteric site on the enzyme. In pure non-competitive inhibition, the binding of the inhibitor to the enzyme doesn't influence the binding of the substrate to the enzyme. However, this situation is uncommon and the binding of the inhibitor usually causes conformational changes in the structure of the enzyme, which in turn affects the binding of the substrate to the enzyme. This is known as mixed non-competitive inhibition (Figure 29).
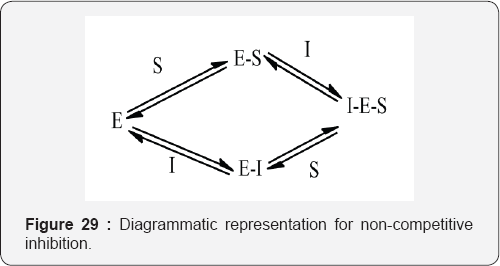
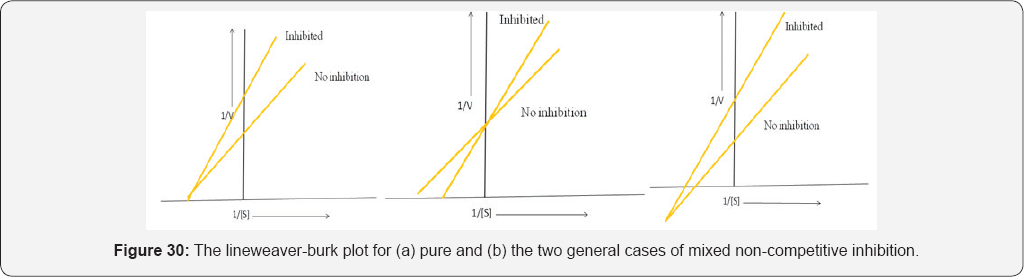
In both types of non-competitive inhibition the binding of the inhibitor to the enzyme ultimately result in the formation of an enzyme-substrate inhibitor (I-E-S) complex, which will not yield the normal products for the enzyme catalysed process. However, some normal products will be formed since not all the E-S complex will be converted to the I-E-S complex. Furthermore, as non-competitive inhibition is reversible the E-S complex can be reformed by the loss of the inhibitor (Figure 30).
Where single substrate enzyme processes exhibit Michaelis- Menten kinetics both types of non-competitive inhibition exhibit characteristic changes to their Line weaver-Burk plots. Both types of non-competitive inhibitors decrease the value of V max .Furthermore, Km remains the same for pure non-competitive inhibition but changes for mixed non-competitive inhibition. The fact is that in non-competitive inhibition the inhibitor doesn't bind to the active site of the enzyme means that the structure of the substrate cannot be used as the basis for designing new drugs that act in this manner to inhibit enzyme action [42].
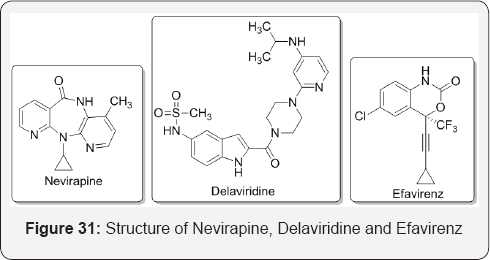
Non-competitive inhibitors have been used to treat AIDS (caused by the HIV virus. This is a retrovirus that uses an enzyme reverse transcriptase (RT) to reproduce). One approach to controlling HIV and hence AIDS is to inhibit RT, which would prevent viral replication. Nevirapine, delaviridine and efavirenz are NNRTIs (Non-nucleoside reverse transcriptase inhibitors) and as such are used in combination with other AIDS drugs to control HIV levels in AIDS cases [44] (Figure 31).
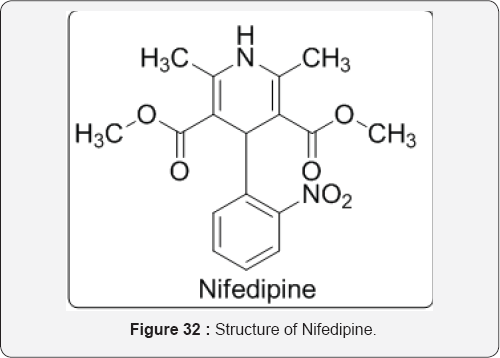
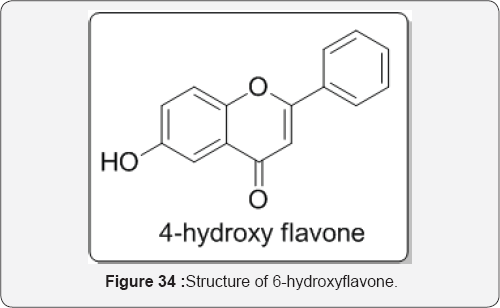
Noncompetitive inhibitors of CYP2C9 enzyme include nifedipine, tranylcypromine, phenethyl isothiocyanate, and 6-hydroxyflavone. Computer docking simulation and constructed mutants substituted indicate that the noncompetitive binding site of 6-hydroxyflavone is the reported allosteric binding site of CYP2C9 enzyme [45] (Figure 32).
The Human Immunodeficiency Virus, or HIV, is the cause of AIDS, an immune syndrome that continues to have profound effects in the United States and around the world. Mainly there is no cure for AIDS, but there are several classes of medications that inhibit the reproduction of HIV, one of which are non-nucleoside reverse transcriptase inhibitors, or NNRTIs. This class of medications suppresses the enzyme reverse transcriptase, a HIV enzyme which converts viral RNA into human-compatible DNA and NNRTIs act as non competitive inhibitor [46].
Uncompetitive inhibition: Uncompetitive inhibitors are believed to form a complex with the enzyme-substrate complex.
E + S↔ES + I ↔IES
The substrate residue in this complex is unable to react and form its normal product. An uncompetitive inhibitor doesn't change the slope of its Line weaver-Burk plot of 1/V against [S] but moves it towards the left hand side of the plot as the concentration of the inhibitor increases. The substrate is often used as the lead compound when designing new uncompetitive inhibitors [42] (Figure 33).
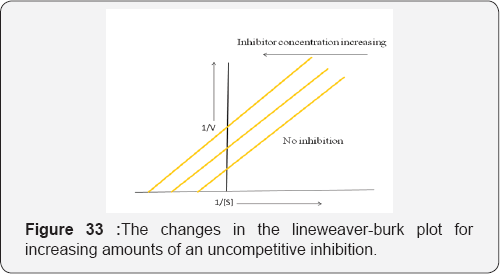
Example: Finasteride is a relatively selective non-competitive inhibitor of 5α-reductase type II (occurs mainly in the prostate gland and testes). The MOA is believed to involve it bonding to the NADH cofactor of the enzyme to form a NADPH-finasteride complex. This complex has a half life of about 30 days and so, as a result, 5α-reductase is almost irreversibly inhibited by the finasteride. Finasteride is also used to treat male pattern baldness. It is believed that 5α-reductase type I (mainly found in liver) also plays a part in the progression of hormone-dependent prostate cancer. As a result, dutasteride, a drug that inhibits both types of 5α-reductase, was developed, and has been approved for use in treating BPH (benign prostate hyperplasia, an enlargement of the prostate that affects many older men) [47]. The inhibition of cytochrome P450 (P450) 2C9 by a series of flavonoids made up of flavones (flavone, 6-hydroxyflavone, 7-hydroxyflavone, chrysin, baicalein, apigenin, luteolin, scutellarein, and wogonin) and flavonols (galangin, fisetin, kaempferol, morin, and quercetin). With the exception of flavone, all flavonoids were shown to inhibit CYP2C9- mediated diclofenac 4'-hydroxylation in the CYP2C9 RECO system. The mechanism of inhibition, 6- hydroxyflavone was found to be a noncompetitive inhibitor of CYP2C9, whereas the other flavonoids were competitive inhibitors [48] (Figure 34).
Irreversible inhibition
These bind to the enzyme by either strong non-covalent or strong covalent bonds. Compounds bound by strong non-covalent bonds will slowly dissociate, releasing the enzyme to carry out its normal function. However, whatever the type of binding the enzyme will resume its normal function once the organism has synthesised a sufficient number of additional enzyme molecules to overcome the effect of the inhibitor [42] (Figure 35).

Classification
Active site directed inhibitors
These are compounds that bind at or near to the active site of the enzyme. These inhibitors usually form strong covalent bonds with the functional groups that are found at the active site or close to that site. The inhibitor effectively reduces the concentration of the active enzyme, which in turn reduces the rate of formation of the normal products of the enzyme process.
The action of many irreversible inhibitors depends on the inhibitor possessing a functional group that can react with a functional group at the active site of the enzyme. As the functional groups found at the active sites of enzymes are usually nucleophiles, the incorporation of strongly electrophilic groups in the structure of a substrate can be used to develop new inhibitors.
Many of the inhibitors developed in this way are too reactive, resulting to be too toxic to be used clinically. However, they have been used to identify the amino acid residues forming the active site of an enzyme. The inhibitor, when it irreversibly binds to a functional group at the active site, effectively acts as a chemical label for the amino acid residue containing that group. Identification of the labelled residue indicates which residues are involved in forming the active site of the enzyme. Example: Aspirin irreversibly inhibits both COX-1 and COX-2 (isozymes of COX, catalysing the metabolism of arachidonic acid to PGG2 which is further oxidised to PGH2 and other PGs. Excessive concentrations of these biologically active metabolic products are associated with inflammation, pain, swelling and fevers) by acetylating serine hydroxy groups at the enzyme's active site, probably by a transesterification mechanism [49]. As thrombin also bind as by four active-site directed inhibitors to human thrombin. The inhibitors are benzamidine, PPACK, NAPAP, and MD-805, of which the last three are compounds evolved specifically to inhibit thrombin. Inhibitors bind to thrombin in a similar way as to trypsin, but thrombin has an extra loop, the "Tyr-Pro-Pro-Trp loop,” not present in trypsin, which gives further binding interactions and is seen to move somewhat to accommodate binding of the different inhibitors [50]. The essential catalytic elements of bacterial collagenases, including the zinc-coordinating residues, as well as their typical domain organization with relevance to collagenolysis and on the structural basis for the substrate specificities, both towards linear and triple-helical peptides. These enzymatic properties shape the structural framework for the discovery and development of competitive, active site-directed inhibitors [51].
Suicide or irreversible mechanism-based inhibitors
Alternatively known as Kcat, these are irreversible inhibitors that are often analogues of the normal substrate of the enzyme. The inhibitor binds to the active site where it is modified by the enzyme to produce a reactive group that reacts irreversibly to form a stable inhibitor-enzyme complex. This subsequent reaction may or may not involve functional groups at the active site. These mechanisms mean that suicide inhibitors are likely to be reasonably specific in their action since they can only be activated by a particular enzyme. Consequently, this specificity means that drugs designed as suicide inhibitors could exhibit a lower degree of toxicity (Scheme 3).
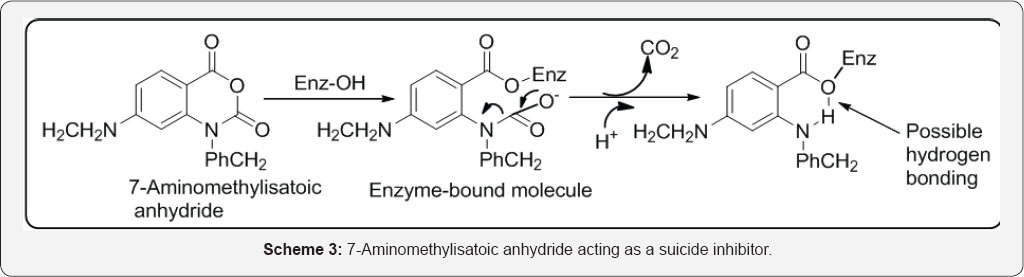
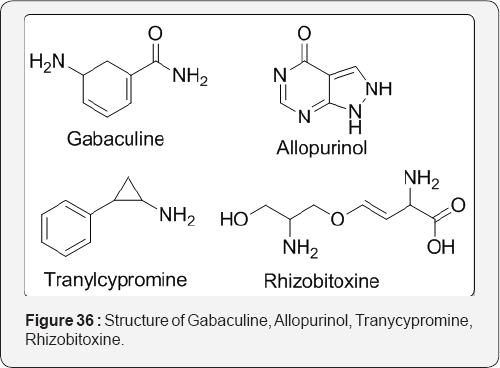
The action of suicide inhibitors usually involves either the prosthetic group of the enzyme or its coenzyme. Example: 7- aminomethylisatoic acid anhydride is a suicide inhibitor of serine protease thrombin, which catalyses the hydrolysis of peptides and proteins. This catalysis involves the rapid hydrolysis of acyl-enzyme intermediates. The inhibitor reacts with the hydroxy group of a serine residue of thrombin to form an acyl derivative, which rapidly decarboxylates to form a hydrolysis-resistant complex stable enough to inhibit the enzyme. It is possible that the resistance to hydrolysis may be due to hydrogen bonding between the o-amino group and the O atom bound to the enzyme. The electrophilic groups formed by many suicide inhibitors often take the form of α, β-unsaturated carbonyl compounds and imines. These electrophiles react by a type of Michael addition with Nucleophilic groups, such as OH of serine residues, the SH of cysteine residue and the M-NH2 of lysine residues frequently found at the active sites of enzymes. Example: The diuretic tienilic acid acts as a suicide inhibitor of CYP-450. Tienilic acid is oxidized by CYP-450 to its corresponding sulphoxide derivative, which acts as the suicide inhibitor [52]. The effect of suicide inhibitors increases as the concentration of the inhibitor increases relative to the concentration of the enzyme. For processes involving single substrates that follow simple Michaelis-Menten kinetics, the Line weaver-Burk plot for different concentrations of the inhibitor is a series of straight lines. Mapping Human Brain Monoamine Oxidase A and B with 11C-Labeled Suicide Inactivators and PET MONOAMINE OXIDASE (MAO) is responsible for the oxidative deamination of endogenous neurotransmitter amines as well as amines from exogenous sources. It exists in two forms, MAO A and MAO B, which are identified by their inhibitor sensitivity and by their substrate selectivity [53]. Walsh et al. [54] defines that suicide substrates as a class of irreversible inactivators of specific target enzymes where the target enzyme participates in its own destruction by catalytic unmasking of a latent functional group at some stage in the catalytic cycle of the enzyme. The example includes are gabaculine, secobarbital, allopurinol, clavulanate and penicillin sulfones, tranylcypramine, rhizobitoxin, fluoroalanine, S-fluorodeoxyuridylate and mitomycin (Figure 36).
Transition State Inhibitors
The substrate in an enzyme-catalysed reaction is converted to the product through a series of transition state structures. Although these transition state structures are transient, they bind to the active state of the enzyme and therefore must have structures that are compatible with the structures of the active site. Consequently, it has been proposed that stable compounds with structures similar to those of these transition state structures could bind to the active site of an enzyme and act as inhibitors for that enzyme. Compounds that fulfil this requirement are known as transition state inhibitors. They can act in a reversible/irreversible manner.
Example
i. The antibiotic coformycin, which inhibits adenosine deaminase, has a structure similar to that of the transition state of adenosine in the process for the conversion of adenosine to inosine which is catalysed by adenosine deaminase.
ii. A consideration of the transition state structure offers a possible method of approach for new drugs discovery: To design the experimental anticancer drug PALA (N-phosphonoacetyl-L- aspartate) to be effective against some cancers in rats.
iii. Condensation of aspartic acid with carbamoyl phosphate to form N-carbamoyl aspartic acid, reaction is being catalysed by aspartate transcarbamoylase.
iv. The transition state for this conversion involves the simultaneous loss of phosphate with the attack of the nucleophilic amino group of the aspartic acid on the carbonyl group of the carbamoyl phosphate.
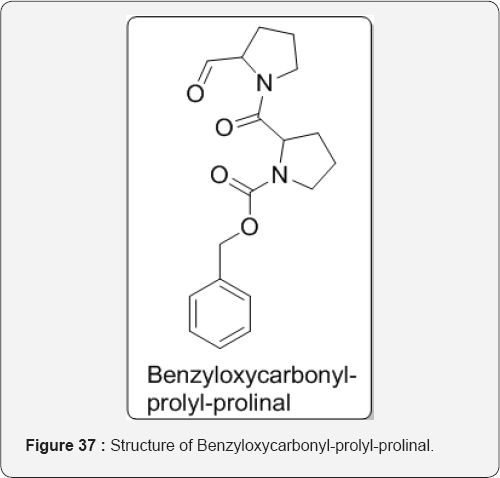
v PALA was designed to have a similar structure to this transition state but without the amino group necessary for the next stage in the synthesis, which is the conversion of N-carbamoyl aspartic acid to dihydroorotic acid, hence inhibiting the formation of uracil. It was found that PALA bound 103 times more tightly to the enzyme than the normal substrate [42]. Wilk and Orlowski et al. [55] describe that prolyl endopeptidase cleaves the peptide bonds on the carboxyl side of proline residues within a peptide chain. The enzyme readily degrades a number of neuropeptides including substance P, neurotensin, thyrotropin-releasing hormone, and luteinizing hormone releasing hormone. Finding that the enzyme is inhibited by benzyloxycarbonyl-prolyl-proline and prompted the synthesis of benzyloxycarbonyl- prolyl-prolinal as a potential transition state analog inhibitor. Even other transition state inhibitors, can be assumed that binding of the prolinal residue to the S, subsite and the formation of a hemiacetal with the active serine of the enzyme greatly contribute to the potency of inhibition (Figure 37).
Schramm et al. [56] described that enzymatic transition-state structures uses kinetic isotope effects (KIEs) to bonding and geometry differences between reactants and the transition state. Computational correlation of experimental values with chemical models permits three dimensional geometric and electrostatic assignment of transition states formed at enzymatic catalytic sites. The combination of experimental and computational access to transition-state information permits (a) the design of transition-state analogs as powerful enzymatic inhibitors, (b) exploration of protein features linked to transition-state structure, (c) analysis of ensemble atomic motions involved in achieving the transition state, (d ) transition-state lifetimes, and (e) Separation of ground-state (Michaelis complexes) from transition-state effects. Transition-state analogs with picomolar dissociation constants have been achieved for several enzymatic targets. Transition states of closely related isozymes indicate that the protein's dynamic architecture is linked to transition- state structure. Fast dynamic motions in catalytic sites are linked to transition-state generation. Enzymatic transition states have lifetimes of femtoseconds, the lifetime of bond vibrations. Binding isotope effects (BIEs) reveal relative reactant and transitionstate analog binding distortion for comparison with actual transition states.
Enzyme Activation of Drugs
Prodrug term involves chemically modified inert compound which upon administration make inactive compound into an active drug to releases the active parent drug to elicit its pharmacological response within the body. Prodrug has been developed for main strategy that is to solve many unwanted drug properties. In most cases, prodrugs contain a promoiety (linker) that is removed by enzymatic or chemical reactions, while other prodrugs release their active drugs after molecular modification, such as an oxidation or reduction reactions. Codrug is defines as two biologically active drugs can be linked together in a single molecule. In a codrug, each drug acts as a linker for the other. Prodrug is used to overcome many pharmaceutical and pharmacokinetic problems such as, low bioavailability by increasing or decreasing lipophilicity of the parent drug, site selectivity for higher absorption and less toxicity, short duration of action to increase patient compliance, rapid metabolism to increase oral bioavailability and masking bitter sensation of commonly used drugs, which is mainly needed for patient compliance [57].
Utility of Prodrugs
Prodrug design is considered as an option after lead optimization has been carried out and when the desired drug candidate has failed in preclinical studies, or prodrug approach should be carried out in early stages of lead optimization when there are pharmacokinetics defects in the molecule, such as poor absorption, very short or very long half life, high first pass effect or off-target inhibition [58].
Reasons for utilizing a prodrug strategy in drug design
A. Aqueous solubility: Consider an injectable drug that is so insoluble in water that it would need to be taken up in more than a litre of saline to administer the appropriate dose. Or what if each dose of your ophthalmic drug required a litre of saline for dissolution but it was to be administered as eye drops. These drugs could be safe, effective and potent, but they would not be viable for their applications. In these cases a water solubilizing group could be attached to the drugs, which is metabolically released after drug administration [59].
B. Absorption and distribution: If the desired drug is not absorbed and transported to the target site in sufficient concentration, it can be made more water soluble or lipid soluble depending on the desired site of action. Once absorption has occurred or when drug is at the appropriate site of action, the water or lipid soluble group is removed enzymatically [60].
C. Site specificity: Specificity for a particular organ or tissue can be made if there are high concentrations of or uniqueness of enzymes present at that site that can cleave the appropriate appendages from the prodrug and unmask the drug [61].
D. Instability: A drug may be rapidly metabolized and rendered inactive prior to when it reaches the site of action. The structure may be modified to block that metabolism until the drug is at the desired site [62].
E. Prolonged release: It may be desirable to have a steady low concentration of a drug released over a long period of time. The drug may be altered so that it is metabolically converted to the active form slowly [63].
F. Toxicity: A drug may be toxic in its active form and would have a greater therapeutic index, if it were administered in a nontoxic inactive form that was converted into the active form only at the sites of action [63].
G. Poor patient acceptability: A active drug may have an unpleasant taste or odor, produce gastric irritation, or cause pain when administered (e.g., when injected). The structure of the drug could be modified to alleviate these problems, but once administered, the prodrug would be metabolized to the active drug [63].
H. Formulation problems: If the drug is a volatile liquid, it would be more desirable to have it in a solid form so that it could be formulated as a tablet. An inactive solid derivative could be prepared that would be converted in the body to the active drug [63].
Types of Prodrugs
A. A carrier linked prodrug is a compound that contains an active drug linked to a carrier group that can be removed enzymatically, such as an ester which is hydrolyzed to an active carboxylic acid containing drug. The bond to the carrier group must be labile enough to allow the active drug to be released efficiently in vivo, and the carrier group must be nontoxic and biologically inactive when detached from the drug. Example, the Ester Prodrug of Acyclovir Interactions with Peptides, Organic Anions and Organic Cations [64].
i. Bipartate prodrug: is a prodrug comprised of one carrier attached to the drug [65].
ii. Tripartate prodrug: When a carrier is connected to a linker that is connected to the drug [65].
iii. Mutual prodrug: Consists of two, usually synergistic drugs attached to each other [65].
B. A bio precursor prodrug is a compound that is metabolized by molecular modification into a new compound which is the active principle or which can be metabolized further to the active drug. e.g., if the drug contains a carboxylic acid group, the bio precursor may be a primary amine that is metabolized by oxidation to the aldehyde, which is further metabolized to the carboxylic acid drug [65].
Mechanisms of Drug Activation
A.Carrier linked prodrugs
i.Carrier linkages for various functional groups
a. Alcohols, carboxylic acids and related groups: Alcohol- containing drugs can be acylated with aliphatic or aromatic carboxylic acids to decrease water solubility or with carboxylic acids containing amino or additional carboxylate groups to increase water solubility. Conversions to phosphate or sulphate esters also increases water solubility. By using these approaches a wide range of solubilities can be achieved that will affect the absorption and distribution properties of the drug. These derivatives also can have an important effect on the dosage form, that is, whether used in tablet form or in aqueous solution.
b. Amines: The pKa of amines can be lowered by approximately 3 units by conversion to their N-Mannich bases). This lowers the basicity of the amine so that at physiological pH few of the prodrug molecules are protonated, thereby increasing its lipophilicity.
c. Sulfonamides: Just like amines, sulfonamides can be acylated, but generates an acidic proton, which makes these compounds amenable to conversion to water-soluble sodium salts. E.g., the 2nd generation anti-inflammatory drug and COX-2 inhibitor valdecoxib has been converted into parecoxib sodium, an injectable analgesic drug.
d. Carbonyl compound: The most important prodrug forms of aldehyde and ketones are Schiff bases, oximes, acetals, enol esters, oxazolidines, and thiazolidines [66].
Conclusion
In nut-shell, enzymes are known to play an important role directly as well as indirectly for the living organisms. The enzymes are the key regulators of several biochemical processes and elevation or decrease in their concentration affects adversely and sometimes lead to diseases also. Several drugs has been designed and synthesized which directly inhibit or modify the actions of the enzymes to prevent the disease. Hence through the understanding of various mechanisms of enzyme action and inhibition, several newer drugs can be designed and biochemical reactions can be made more efficient.
Acknowledgment
We sincerely thank of ISF College of pharmacy for their guidance and advice at all times.
References
- Smith A, Datta S, Smith G, Campbell P, Bentley R, et al. (1997) Oxford dictionary of biochemistry and molecular biology. Oxford: Oxford University Press, India.
- Sanchez S, Demain AL (2009) Microbial Primary Metabolites: Biosynthesis and Perspectives. Encyclopedia of Industrial Biotechnology.
- Chen Y (2013) The Structure and Function Study of Three Metalloenzymes That Utilize Three Histidines as Metal Ligands.
- Hashim OH, Adnan N A (1994) Coenzyme, cofactor and prosthetic group—ambiguous biochemical jargon. Biochemical education 22: 93-94.
- Stroud RM, Krieger M, Koeppe RE, Kossiakoff AA, Chambers J L (1975) Structure-Function Relationships in the Serine Proteases. Proteases and biological control pp: 13-32.
- Tibolla G, Norata GD, Artali R, Meneghetti F, Catapano A (2011) Proprotein convertase subtilisin/kexin type 9 (PCSK9): From structure-function relation to therapeutic inhibition. Nutrition. Metabolism and Cardiovascular Diseases 21(11): 835-843.
- Inoue M, kishimoto A, Takai Y, Nishizuka Y (1977) Studies on a cyclic nucleotide-independent protein kinase and its proenzyme in mammalian tissues. II. Proenzyme and its activation by calcium- dependent protease from rat brain. Journal of Biological Chemistry 252(21): 7610-7616.
- Guroff G (1964) A neutral, calcium-activated proteinase from the soluble fraction of rat brain. The Journal of biological chemistry 239: 149-155.
- Huston R, Krebs E (1968) Activation of skeletal muscle phosphorylase kinase by calcium ions. II. Identification of the kinase activating factor as a proteolytic enzyme. Biochemistry 7(6): 2116-2122.
- Drummond GI, Duncan L (1968) On the mechanism of activation of phosphorylase b kinase by calcium. Journal of Biological Chemistry 243: 5532-5538.
- Reddy M K, Etlinger JD, Rabinowitz M, Fischman DA, Zak R (1975) Removal of Z-lines and alpha-actinin from isolated myofibrils by a calcium-activated neutral protease. Journal of Biological Chemistry 250: 4278-4284.
- Dayton WR, Reville W, Goll De, Stromer M (1976) A Ca2+-activated protease possibly involved in myofibrillar protein turnover. Partial characterization of the purified enzyme. Biochemistry 15(10): 21592167.
- Belocopitow E, Appleman MM, Torres HN (1965) Factors Affecting the Activity of Muscle Glycogen Synthetase II. THE REGULATION BY Ca++. Journal of Biological Chemistry 240(9): 3473-3478.
- Abita JP, Lazdunski M, Bonsen PP, Pieterson WA, Haas GH (1972) Zymogen-Enzyme Transformations. European Journal of Biochemistry 30: 37-47.
- Augoff K, Hryniewicz-Jankowska A, Tabola R (2015) Lactate dehydrogenase 5: an old friend and a new hope in the war on cancer. Cancer letters 358: 1-7.
- Houddane A, Bultot L, Novellasdemunt L, Johanns M, Gueuning MA, et al. (2017) Role of Akt/PKB and PFKFB isoenzymes in the control of glycolysis, cell proliferation and protein synthesis in mitogen- stimulated thymocytes. Cellular Signalling 34: 23-37.
- Annesley T M, Till GO, Ward PA (1985) Cutaneous thermal burn and oxidant-mediated acute lung injury: Appearance in serum of lung- related LDH isoenzyme. Journal of free radicals in biology & medicine 1: 409-414.
- Bock R A, Soulages JL, Barrow WW (2007) Substrate and inhibitor specificity of Mycobacterium avium dihydrofolate reductase. FEBS Journal 274(13): 3286-3298.
- Lin F, Ferguson KL, Boyer DR, Lin XN, Marsh ENG (2015) Isofunctional enzymes PAD1 and UbiX catalyze formation of a novel cofactor required by ferulic acid decarboxylase and 4-hydroxy-3-polyprenylbenzoic acid decarboxylase. ACS chemical biology 10(4): 1137-1144.
- Davidi D, Milo R (2017) Lessons on enzyme kinetics from quantitative proteomics. Current Opinion in Biotechnology 46: 81-89.
- Chopra H (2007) Chemical Kinetics and Catalysis.
- Dimitrov S (2017) A modeling approach for cell growth based on enzyme kinetics. Biomath Communications 3.
- Obach R S, Reed-Hagen AE (2002) Measurement of Michaelis constants for cytochrome P450-mediated biotransformation reactions using a substrate depletion approach. Drug Metabolism and Disposition 30(7): 831-837.
- Seibert E, Tracy TS (2014) Fundamentals of enzyme kinetics. Enzyme Kinetics in Drug Metabolism: Fundamentals and Applications 9-22.
- Zhou Y, Zhuang X (2007) Kinetic analysis of sequential multi-step reactions. The journal of physical chemistry. B 111(48): 13600.
- Cha S (1968) A simple method for derivation of rate equations for enzyme-catalyzed reactions under the rapid equilibrium assumption or combined assumptions of equilibrium and steady state. Journal of biological chemistry 243: 820-825.
- Kirby AJ, Acid-Base Catalysis byEnzymes. University of Cambridge, Cambridge, UK.
- Mcnaught AD (1997) Compendium of chemical terminology, Blackwell Science Oxford.
- Botos I, Wlodawer A (2007) The expanding diversity of serine hydrolases. Current opinion in structural biology 17: 683-690.
- Kenyon G L, Covalent Nucleophilic Catalysis.
- Pratt CW, Cornely K (2011) Essential Biochemistry, Wiley
- Larbie C, Metabolism in Health And Diseases I Vitamins and Coenzymes.
- Chowdhury S, Banerjee R (1999) Role of the dimethylbenzimidazole tail in the reaction catalyzed by coenzyme B12-dependent methylmalonyl- CoA mutase. Biochemistry 38((46): 15287-15294.
- Soetens M, Drouet F, Riant O (2017) (r|5-Pentamethylcyclopentadienyl) iridium Complex Catalyzed Imine Reductions Utilizing the Biomimetic 1, 4-NAD (P) H Cofactor and N-Benzyl-1, 4-dihydronicotinamide as the Hydride-Transfer Agent. ChemCatChem 9: 929-933.
- Haglin L, Johansson I, Forsgren L, Backman L (2017) Intake of vitamin B before onset of Parkinson's disease and atypical parkinsonism and olfactory function at the time of diagnosis. European Journal of Clinical Nutrition 71: 97-102.
- Aughey WS, Smythe G, O'keeffe D, Maskasky J, Smith M (1975) Heme A of cytochrome c oxicase. Structure and properties: comparisons with hemes B, C, and S and derivatives. Journal of Biological Chemistry 250: 7602-7622.
- Travassos LH, Vasconcellos LR, Bozza MT, Carneiro LA (2017) Heme and iron induce protein aggregation. Autophagy 13.
- Phillips IR, Shephard EA (2008) Flavin-containing monooxygenases: mutations, disease and drug response. Trends in pharmacological sciences 29(6): 294-301.
- Zhou J, Shephard EA (2006) Mutation, polymorphism and perspectives for the future of human flavin-containing monooxygenase 3. Mutation Research/Reviews in Mutation Research 612(3): 165-171.
- Ondetti MA, Rubin B, Cushman DW (1977) Design of specific inhibitors of angiotensin-converting enzyme: new class of orally active antihypertensive agents. Science 196: 441-444.
- Forbes LV, Sjogren T, Auchere F, Jenkins DW, Thong B, et al. (2013) Potent reversible inhibition of myeloperoxidase by aromatic hydroxamates. Journal of Biological Chemistry 288: 36636-36647.
- Sharma R (2012) Enzyme inhibition: mechanisms and scope. INTECH Open Access Publisher.
- Kimura T, Hauber J, Singer TP (1967) Studies on Succinate Dehydrogenase XIII. Reversible Activation of The Mammalian Enzyme. Journal of Biological Chemistry 242: 4987-4993.
- Usach I, Melis V, Peris JE (2013) Non-nucleoside reverse transcriptase inhibitors: a review on pharmacokinetics, pharmacodynamics, safety and tolerability. Journal of the International AIDS Society 16.
- Schonauer E, Brandstetter H (2016) Inhibition and Activity Regulation of Bacterial Collagenases. Medicinal Chemistry 22: 69-94.
- Jacob S (2013) An Examination of Non-Nucleoside Reverse Transcriptase Inhibitors.
- Azzouni F, Godoy A, Li Y, Mohler J (2011) The 5 alpha-reductase isozyme family: a review of basic biology and their role in human diseases. Advances in urology pp: 18.
- Si D, Wang Y, Zhou YH, Guo Y, Wang J, et al. (2009) Mechanism of CYP2C9 inhibition by flavones and flavonols. Drug metabolism and disposition 37(3): 629-634.
- Baigent C, Patrono C (2003) Selective cyclooxygenase 2 inhibitors, aspirin, and cardiovascular disease: a reappraisal. Arthritis & Rheumatism 48: 12-20.
- Banner DW, Hadvary P (1991) Crystallographic analysis at 3.0-A resolution of the binding to human thrombin of four active site-directed inhibitors. Journal of Biological Chemistry 266(30): 20085-20093.
- Schonauer E, Brandstetter H (2016) Inhibition and Activity Regulation of Bacterial Collagenases.
- Farady CJ, Craik CS (2010) Mechanisms of macromolecular protease inhibitors. Chembiochem 11(17): 2341-2346.
- Fowler J, Macgregor R, Wolf A, Arnett C, Dewey S, et al. (1987) Mapping human brain monoamine oxidase A and B with carbon eleven-labeled suicide inactivators and PET. Science 235: 481-486.
- Waleh C (1982) Suicide substrates: mechanism-based enzyme inactivators. Tetrahedron 38: 871-909
- Wilk S, Orlowski M (1983) Inhibition of Rabbit Brain Prolyl Endopeptidase by N-Benzyloxycarbonyl-Prolyl-Prolinal, a Transition State Aldehyde Inhibitor. Journal of neurochemistry 41: 69-75.
- Schramm VL (2011) Enzymatic transition states, transition-state analogs, dynamics, thermodynamics, and lifetimes. Annual review of biochemistry 80: 703-732.
- Rafik Karaman (2014) Prodrugs Design (Pharmacology - Research, Safety Testing and Regulation), Edition: 1, Chapter: II, Publisher: Nova Science Publishers Inc pp: 77-102.
- Boa A (2003) Introduction to drug discovery. Department of Chemistry, University of Hull. UK.
- Stella VJ, Nti-Addae KW (2007) Prodrug strategies to overcome poor water solubility. Advanced drug delivery reviews 59: 677-694.
- Rautio J, Kumpulainen H, Heimbach T, Oliyai R, Oh D, et al. (2008) Prodrugs: design and clinical applications. Nature Reviews Drug Discovery 7: 255-270.
- 6> Friend DR, Pangburn S (1987) Site-specific drug delivery. Medicinal research reviews 7: 53-106.
- Zawilska J B, Wojcieszak J, Olejniczak AB (2013) Prodrugs: a challenge for the drug development. Pharmacological reports 65: 1-14.
- Huttunen KM, Raunio H, Rautio J (2011) Prodrugs—from serendipity to rational design. Pharmacological reviews 63: 750-771.
- Sinko PJ, Balimane PV (1998) Carrier-mediated intestinal absorption of valacyclovir, the L-valyl ester prodrug of acyclovir. 1. Interactions with peptides, organic anions and organic cations in rats. Biopharmaceutics & drug disposition 19(4): 209-217.
- Karpagavalli L, Vigneshwar M, Monisha M, Prabavathi M, Prasanth Pet al. ( review on prodrugs.
- Rautio J, Kumpulainen H, Heimbach T, Oliyai R, Oh D, et al. (2008) Prodrugs: design and clinical applications. Nature Reviews Drug Discovery 7: 255-270.






























