Synthesis and Antibacterial, Antioxidant and DNA Cleavage Evaluation of Triazenes Containing Sulfathiazole Moiety
Mehmet Boga1,2, Yener Tekeli3, Nebih Lolak4, Gorkem Deniz Sonmez5 and Suleyman Akocak4*
1Department of Analytical Chemistry, Faculty of Pharmacy, Dicle University, Turkey
2Dicle University Health Sciences Application and Research Center (DÜSAM), Turkey
3Department of Pharmaceutical Biotechnology, Faculty of Pharmacy, Adıyaman University, Turkey
4Department of Pharmaceutical Chemistry, Faculty of Pharmacy, Adıyaman University, Turkey
5Department of Pharmaceutical Chemistry,, Faculty of Pharmacy, Adıyaman University, Turkey
Submission: December 21, 2021; Published: January 11, 2022
*Corresponding author: Suleyman Akocak, Department of Pharmaceutical Botany, Faculty of Pharmacy, Adıyaman University, 02040 Adıyaman, Turkey
How to cite this article: Mehmet B, Yener T, Nebih L, Gorkem D S, Suleyman A. Synthesis and Antibacterial, Antioxidant and DNA Cleavage Evaluation of Triazenes Containing Sulfathiazole Moiety. Nov Appro Drug Des Dev. 2022; 6(2): 555684. DOI: 10.19080/NAPDD.2022.06.555684
Abstract
Triazenes are one of the most important linker groups that recently investigated as potential enzyme inhibitors. For this reason, in the present work, a series of 1,3-diaryltriazene-substituted sulfathiazole derivatives were re-synthesized by condensation of diazonium salt of sulfathiazole and substituted aromatic amines under optimized conditions. The obtained compounds were tested for their antibacterial, DNA cleavage, and antioxidant properties (including several bioanalytical methods such as DPPH free radical scavenging assay, ABTS cation radical decolarization, metal chelating and cupric reducing antioxidant capacity (CUPRAC)). In general, compounds were more susceptible to gram (-) bacteria strain (P. aeruginosa) with two compounds ST-4 and ST-10 have minimal inhibitory concentration (MIC) values of 15.6μg/mL as compared standard sulfathiazole (MIC: 125μg/mL). On the other hand, all compounds showed weak to moderate antioxidant activity against tested methods.
Keywords: Sulfathiazole; Triazen; Antibacterial; DNA cleavage; Antioxidant
Introduction
The sulfonamide group containing compounds and their isosters (sulfamides, sulfamates) was designed, synthesized and developed as antibacterial agents to treat a broad spectrum of bacterial diseases for decades [1]. Among these studies, many drugs came out and used as potent antimicrobial drugs [1-3]. In the current century, sulfonamides are still a hot topic to discover new and biologically more potent compounds. More specifically, in recent studies, sulfonamides were used as powerful pharmacophore to develop more potent carbonic anhydrase inhibitors (this enzyme is related to many diseases such as, diuretic, cancer, glaucoma, epilepsy, obesity, Alzheimer, etc.) [4-8], antibacterial agents [9,10], and antioxidants [11,12].
Triazenes are a diverse group of compounds contains three contiguous nitrogene atoms that can be easily synthesized from amines or alkyl azides [13,14]. This linker is one of the most important groups for many compounds due to their easy preparation and great pharmaceutical properties in medicinal chemistry research. When the literature studies are examined, it can be seen that the 1,3-diaryl-substituted triazenes gained more importance, more specifically in potent and selective enzyme inhibition studies, in recent years [15,16]. On the other hand, triazene substituted compounds were extensively used in the many drugs design and development studies as anticancer [17,18], antibacterial [19], carbonic anhydrase inhibitors [20-22], acetylcholinesterase inhibitors [23], and antioxidant agents [24].
In our previous study, we have successfully designed and synthesized 1,3-diaryltriazene-substituted sulfathiazole derivatives and enzyme inhibition properties of compounds were evaluated against metabolic enzymes; α-glycosidase (α-GLY), human carbonic anhydrase (hCA I and hCA II), and acetylcholinesterase (AChE) [22]. Among the tested series, some of the compounds greatly inhibited enzymes with nanomolar range [22]. In the present study, we aimed to investigate antibacterial (toward two Gram-positive bacteria and two Gramnegative), antioxidant (four different methods such as, DPPH free radical scavenging assay, ABTS cation radical decolarization, metal chelating and CUPRAC) and DNA cleavage activities of re-synthesized 1,3-diaryltriazene-substituted sulfathiazole derivatives.
Material and Methods
All chemicals and anhydrous solvents were purchased and used without any further purification. Reactions were monitored by thin-layer chromatography (TLC) on Merck silica gel 60 F254 plated visualizing with ultraviolet light and appropriate solvents were used as mobile phase. IR spectra were recorded by using PerkinElmer Spectrum 100 FT-IR spectrum and are expressed in cm-1 (400-4000 cm-1). 1H-NMR and 13C-NMR spectra were recorded using a Bruker Advance III 300 MHz spectrometer using DMSO-d6 as a solvent; the chemical shifts are referenced to tetramethylsilane (TMS). Melting points (mp) were recorded with SMP30 melting point apparatus in open capillary tubes and were uncorrected.
Chemistry
Compounds ST(1-11) were synthesized by diazo reaction as previously described by us [22]. Briefly, the diazonium salt of sulfathiazole was reacted with various substituted aromatic amines to obtain targeted compounds ST(1-11) as summarized in Scheme 1. Physicochemical and spectroscopic characteristic of all the synthesized compounds were previously reported by us [22].
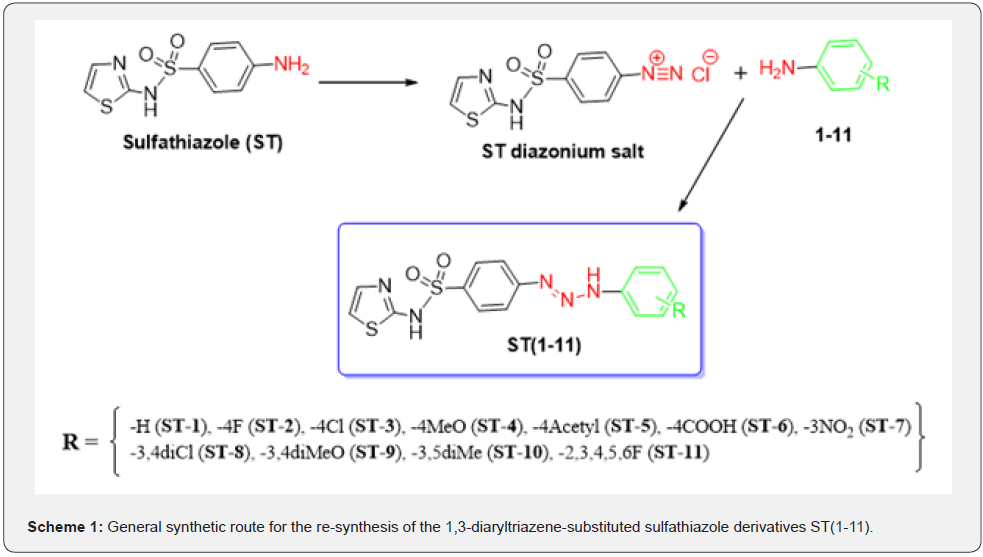
Scheme 1 General synthetic route for the re-synthesis of the 1,3-diaryltriazene-substituted sulfathiazole derivatives ST(1-11).
Antibacterial activities
The microdilution susceptibility tests were carried out in Nutrient broth for the evaluation of antibacterial activity. The freshly synthesized, 1,3-diaryltriazene-substituted sulfathiazole derivatives ST(1-11) were evaluated for their antimicrobial activity against four pathogenic bacterial strains, including Gram-positive (Staphylococcus aureus ATCC 29213, Enterococcus faecalis ATCC 29212) and Gram-negative bacteria (Pseudomonas aeruginosa ATCC 10231, Escherichia coli ATCC 25912).
The antimicrobial analyzes were carried out by modifying according to Al blewi et al. [25]. Briefly, the synthesized compounds were dissolved in DMSO compounds and 5mg/mL stock solution was prepared and were tested at eight different concentrations diluted seven times. The bacterial strain (100μL) was added to each microplate well to be approximately 106 CFU/mL bacteria, adjusting the density with a McFarland densitometer. Sulfathiazole was used as a standard compound for comparison reasons and the same procedures were applied to it. Microplates were incubated for 24 hours at 37°C to measure antibacterial activity, then optical densities were measured at 600 nm (OD600) using a microplate reader (Biotek ELX800 elisa reader). MIC assessments were repeated three times for each microorganism and each compound.
Minimal inhibitory concentrations (MICs) were determined at the end of the 24-hour incubation period. Control experiments with standard antimicrobial agents (positive control) and unvaccinated media (negative control) were performed in parallel and in the same manner as the compounds examined conditions.
Antioxidant activity assays
The in vitro method was followed in the literature and previous studies reported by us, for DPPH free radical scavenging activity assay [26-28], ABTS cation radical decoloriaztion activity assay [29, 30], and CUPRAC activity assay [31]. On the other hand, BHA, BHT and α-TOC were used as standards for comparison. The metal chelating activity assay was also applied as reported in the literature and EDTA was used as standard [32,33].
Plasmid DNA cleavage studies
The plasmid DNA cleavage assay was performed with pET21a (Novagen) plazmid DNA by using the agarose gel electrophoresis method. Escherichia coli Dh5α cells carrying pET21a (Novagen) plasmid were inoculated into fresh LB medium containing 100μg/ mL ampicillin and the culture was grown at 220rpm for overnight at 37°C. Plasmid DNA was extracted by using plasmid DNA isolation kit (K0502, Thermo Fisher Scientific™) and stored at 4°C until needed. In order to examine the plasmid cleavage activity of the re-synthesized compounds; 1μg plasmid DNA, 50μM of each of the derivatives dissolved in 25μM in 10 mM TrisHCl/1mM EDTA buffer pH 8.0 and 0.4M dimethyl sulfoxide (DMSO) were mixed in an Eppendorf tube.
Plasmid DNA with no compound and DMSO as a hydroxy radical scavenger and plasmid DNA and only DMSO were used for control. The samples were incubated at 37°C. After 3h incubation; the samples were mixed with 6X bromophenol blue loading dye (Fermentas) and loaded on a 0.8% agarose gel containing 2.0μg/ mL ethidium bromide. The gel was run for 30 min in 0.5X Tris- Borate-EDTA (TBE) buffer at 120V. The gel was visualized by UV with Quantum ST5 gel documentation system.
Results and Discussion
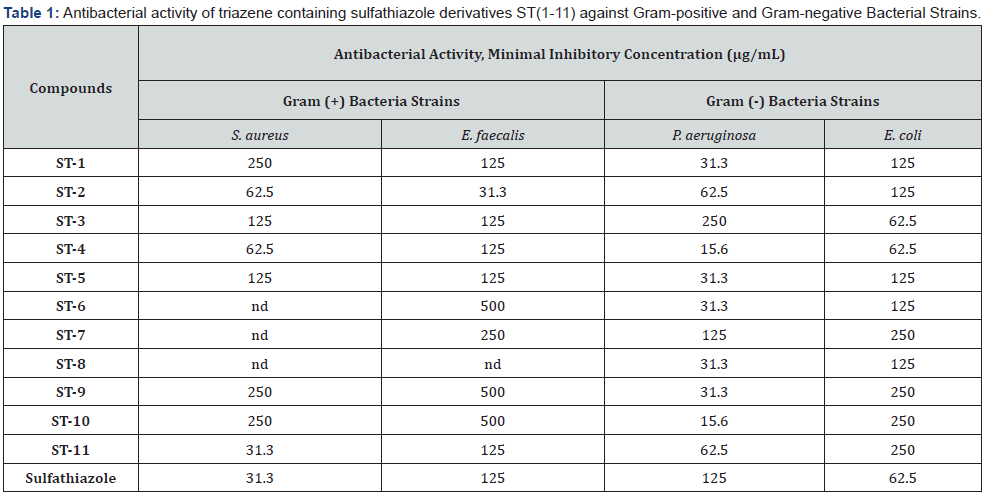
All the re-synthesized triazene containing sulfathiazole derivatives ST(1-11) were assessed for their antibacterial properties against Gram-positive (Staphylococcus aureus ATCC 29213, Enterococcus faecalis ATCC 29212) and Gram-negative bacteria (Pseudomonas aeruginosa ATCC 10231, Escherichia coli ATCC 25912) by obtaining their minimum inhibitory concentration (MIC) values demonstrated in Table 1. The starting compound, sulfathiazole, was used as a reference drug. Among the tested series, some compounds showed greater antibacterial activity, comparing with standard drug sulfathiazole against grampositive bacteria strains including ST-11 (against S. aureus with 31.3μg/mL) and ST-2 (against E. faecalis with 31.3 μg/mL) which is interestingly, both of them have flor atom in their structure (2,3,4,5,6-penta-F, and 4-F, respectively). Another halogeno atom chlor (4-Cl and 3,4di-Cl) did not show same potential effect on gram (+) strains. On the other hand, compounds ST-6, ST-7 and ST-8 against S. aureus strain and compound ST-8 against E. faecalis strain showed any activity. Also, for gram (-) strains, the compounds ST-4 (4-MeO) and ST-10 (3,5di-Me) were more potent than the reference drug with MIC = 15.6μg/mL, while only one compound was less active than the sulfathiazole at MIC = 250μg/ mL against P. aeruginosa (ATCC 10231).
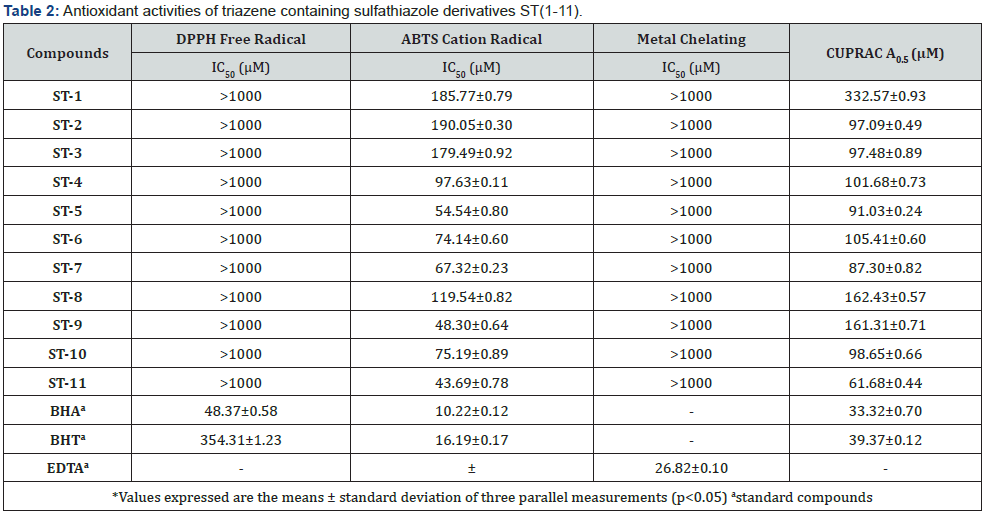
Antioxidant capacities of the re-synthesized compounds were also assessed by using several different bioanalytical methods, including DPPH free radical scavenging assay, ABTS cation radical decolarization, methal chelating, and CUPRAC. In general, all compounds showed weak to moderate antioxidant activity against tested methods. More specifically, any compounds showed DPPH free radical and metal chelating activity by having IC50 value of >1000 μM, which table 2 demonstrates that our compounds not susceptible to these methods. The best antioxidant activity was observed with ABTS cation radical method among the tested series, which is also weak as compared with standards BHT and BHA. The moderate activity was observed with the compounds ST- 5, ST-9 and ST-11 with IC50 values of 54.54±0.80, 48.30±0.64, and 43.69±0.78μM, respectively.
The plasmid DNA cleavage assay was performed to determine the DNA protection ability of each of the re-synthesized compounds ST(1-11) to protect pET21a plasmid DNA from the damaging effects of hydroxyl radicals [34]. Agarose gel electrophoresis method is used frequently in order to reveal the activity of the compounds as chemical nucleases. Circular plasmid DNA is faster than its linear form in agarose gel electrophoresis [35]. If the scission is on one strand of the plasmid, then open circular form (oc DNA; form 2) occurs. If the scission is on on both strands, a linear conformation (form 3) occurs and this configuration will migrate between Form I and II [36]. In the present study, the ability of triazene containing sulfathiazole derivatives ST(1-11) to cleave plasmid DNA was studied by means of gel electrophoresis with supercoiled pET21a in 10mM TrisHCl/1 mM EDTA buffer (pH 8.0) and with DMSO activation.

According to our results presented in Figure 1, the addition of hydroxyl radical scavengers (lane 2) significantly attenuates DNA strand scission. A decrease in the percentage of form I was observed in compounds (Figure 1). ST-2 caused only the formation of Form II. ST-1, ST-4, ST-5, b, and ST-11 caused the formation of Form II and Form III together in the presence of DMSO. On the other hand, ST-3 and ST-6 showed Form I and Form II configuration occured together. ST-10 had an interesting formation in which any DNA fragment was seen was similar to the control group that was the lack of any compounds. The result of the plasmid DNA cleavage activity of our re-synthesized triazene containing sulfathiazole derivatives revealed that our compounds are effective in DNA cleavage with the help of hydroxy radicals.
Conclusion
Herein, we report a series of 1,3-diaryltriazene-substituted sulfathiazole derivatives, which were re-synthesized for their antibacterial, DNA cleavage, and antioxidant properties. Two Gram-positive (Staphylococcus aureus, Enterococcus faecalis) and two Gram-negative bacteria (Pseudomonas aeruginosa, Escherichia coli) were used for antibacterial testing and in general most of the compounds showed moderate to high antibacterial activity as compared with standard sulfathiazole. More specifically, compounds ST-4 and ST-10 have MIC values of 15.6μg/mL against Gram (-) bacteria strain P. aeruginosa. On the other hand, all compounds showed no DPPH free radical scavenging and metal chelating activity by having IC50 value of >1000μM. The resynthesized compounds were more susceptible to ABTS cation radical decolarization and CUPRAC assays. Also, our compounds were efficient in DNA cleavage activity. As a result, these types of compounds might be investigated and improved as potent and effective antibacterial agents.
References
- Tacic A, Nikolic V, Nikolic L, Savic I (2017) Antimicrobial sulfonamide drugs. Advanced Technologies 6(1): 58-71.
- Supuran CT (2008) Carbonic anhydrases: novel therapeutic applications for inhibitors and activators. Nat Rev Drug Discov 7: 168-181.
- Supuran CT, Capasso C (2020) Antibacterial carbonic anhydrase inhibitors: an update on the recent literature. Expert Opin Ther Pat 30(12): 963-982.
- Akocak S, Alam MR, Shabana AM, Sanku RKK, Vullo D, et al. (2016) PEGylated Bis-Sulfonamide Carbonic Anhydrase Inhibitors Can Efficiently Control the Growth of Several Carbonic Anhydrase IX-Expressing Carcinomas. J Med Chem 59(10): 5077-5088.
- Akocak S, Lolak N, Bua S, Turel I, Supuran CT (2018) Synthesis and biological evaluation of novel N,N`-diaryl cyanoguanidines acting as potent and selective carbonic anhydrase II inhibitors. Bioorg Chem 77: 245-251.
- Akocak S, Guzel Akdemir O, Sanku RKK, Russom SS, Iorga BI, et al. (2020) Pyridinium derivatives of 3-aminobenzenesulfonamide are nanomolar-potent inhibitors of tumor-expressed carbonic anhydrase isozymes CA IX and CA XII. Bioorg Chem 103: 104204.
- Lolak N, Akocak S, Turkes C, Taslimi P, Isik M, et al. (2020a) Synthesis, characterization, inhibition effects, and molecular docking studies as acetylcholinesterase, α-glycosidase, and carbonic anhydrase inhibitors of novel benzenesulfonamides incorporating 1,3,5-triazine structural motifs. Bioorg Chem 100: 103897.
- Pacchiano F, Carta F, McDonald PC, Lou Y, Vullo D, et al. (2011) Ureido-substituted benzenesulfonamides potently inhibit carbonic anhydrase IX and show antimetastatic activity in a model of breast cancer metastasis. J Med Chem 54(6): 1896-1902.
- Verma SS, Verma R, Xue F, Thakur PK, Girish YR, et al. (2020) Antibacterial activities of sulfonyl or sulfonamide containing heterocyclic derivatives and its structure-activity relationships (SAR) studies: A critical review. Bioorg Chem 105: 104400.
- Azevedo Barbosa H, Dias DF, Franco LL, Hawkes JA, Carvalho DT (2020) From antibacterial to antitumor agents: A brief review on the chemical and medicinal aspects of sulfonamides. Mini Rev Med Chem 20(19): 2052-2066.
- Abbas A, Murtaza S, Tahir MN, Shamim S, Sirajuddin M, et al. (2016) Synthesis, antioxidant, enzyme inhibition and DNA binding studies of novel N-benzylated derivatives of sulfonamide. J Mol Struct 1117: 269-275.
- Lolak N, Tuneg M, Dogan A, Boga M, Akocak S (2020) Synthesis and biological evaluation of 1,3,5-triazine-substituted ureido benzenesulfonamides as antioxidant, acetylcholinesterase and butyrylcholinesterase inhibitors. Bioorg Med Chem Rep 3(2): 22-31.
- Marchesi F, Turriziani M, Tortorelli G, Avvisati G, Torino F, et al. (2007) Triazene compounds: Mechanism of action and related DNA repair systems. Pharmacol Res 56(4): 275-287.
- Kimball DB, Haley MM (2002) Triazenes: A versatile tool in organic synthesis. Angew Chem Int Ed 41(18): 3338-3351.
- Akocak S, Lolak N, Bua S, Supuran CT (2018) Discovery of novel 1,3-diaryltriazene sulfonamides as carbonic anhydrase I, II, VII, and IX inhibitors. J Enzyme Inhib Med Chem 33(1): 1575-1580.
- Bilginer S, Gonder B, Gul HI, Kaya R, Gulcin I, et al. (2020) Novel sulphonamides incorporating triazene moieties show powerful carbonic anhydrase I and II inhibitory properties. J Enzyme Inhib Med Chem 35(1): 325-329.
- Canakci D, Koyuncu I, Lolak N, Durgun M, Akocak S, et al. (2019) Synthesis and cytotoxic activities of novel copper and silver complexes of 1,3-diaryltriazene-substituted sulfonamides. J Enzyme Inhib Med Chem 34(1): 110-116.
- Tuluce Y, Gorgisen G, Gulacar IM, Koyuncu I, Durgun M, et al. (2018) Antiproliferative and Apoptotic Role of Novel Synthesized Cu(II) Complex with 3-(3-(4-fluorophenyl)Triaz-1-en-1-yl) Benzenesulfonamide in Common Cancer Models. Anticancer Res 38(9): 5115-5120.
- Figueiredo PMS, Filho JCS, Sodre AJS, Junior JRC, Goncalves IS, et al. (2021) Assessment of the biological potential of diaryltriazene-derived triazene compounds. Scientific Rep 11: 2541.
- Lolak N, Akocak S, Bua S, Koca M, Supuran CT (2018) Design and synthesis of novel 1,3-diaryltriazene-substituted sulfonamides as potent and selective carbonic anhydrase II inhibitors. Bioorg Chem 77: 542-547.
- Bilginer S, Gul HI, Anil B, Demir Y, Gulcin I (2020) Synthesis and in silico studies of triazene-substituted sulfamerazine derivatives as acetylcholinesterase and carbonic anhydrase inhibitors. Arch Pharm 354(1): 2000243.
- Isik M, Akocak S, Lolak N, Taslimi P, Turkes C, et al. (2020) Synthesis, characterization, biological evaluation, and in silico studies of novel 1,3-diaryltriazene-substituted sulfathiazole derivatives. Arch Pharm 353(9): 2000102.
- Bilginer S, Gul HI, Hanci H, Gulcin I (2021) Antibacterial and acetylcholinesterase inhibitory potentials of triazenes containing sulfonamide moiety. Pharmaceutical Chem J 55: 284-289.
- Akocak S, Boga M, Lolak N, Tuneg M, Sanku RKK (2019) Design, synthesis and biological evaluation of 1,3-diaryltriazene-substituted sulfonamides as antioxidant, acetylcholinesterase, and butyrylcholinesterase inhibitors. J Turk Chem Soc Sect A: Chem 6(1): 63-70.
- Al-blewi FF, Almehmadi MA, Aouad MR, Bardaweel SK, Sahu PK, et al. (2018) Design, synthesis, ADME prediction and pharmacological evaluation of novel benzimidazole-1,2,3-triazole-sulfonamide hybrids as antimicrobial and aniproliferative agents. Chem Central J 12: 110.
- Blois MS (1958) Antioxidant determinations by the use of a stable free radical. Nature 181: 1199-1200.
- Lolak N, Boga M, Tuneg M, Karakoc G, Akocak S, et al. (2020) Sulphonamides incorporating 1,3,5-triazine structural motifs show antioxidant, acetylcholinesterase, butyrylcholinesterase, and tyrosinase inhibitory profile. J Enzyme Inhib Med Chem 35(1): 424-431.
- Lolak N, Akocak S (2020) Biological evaluation of aromatic bis-sulfonamide Schiff bases as antioxidant, acetylcholinesterase and butyrylcholinesterase inhibitors. Cumhuriyet Sci J 41(2): 413-418.
- Pellegrini RRN, Proteggente A, Pannala A, Yang M, Rice-Evans C (1999) Antioxidant activity applying and improved ABTS radical cation decolorization assay. Free Rad Bio Med 26: 1231-1237.
- Oguz M, Kalay E, Akocak S, Nocentini A, Lolak N, et al. (2020) Synthesis of calix [4] azacrown substituted sulphonamides with antioxidant, acetylcholinesterase, butyrylcholinesterase, tyrosinase, and carbonic anhydrase inhibitory action. J Enzyme Inhib Med Chem 35(1): 1215-1223.
- Apak R, Guclu K, Ozyurek M, Karademir SE (2004) Novel total antioxidant capacity index for dietary polyphenols and vitamine C and E, using their cupric ion reducing capability in the presence of neocuproine: CUPRAC method. J Agric Food Chem 52: 7970-7981.
- Dinis TCP, Maderia VMC, Almedia LM (1994) Action of phenolic derivatives (acetoaminophen, salycilate and 5-aminosalycilate) as inhibitors of membrane lipid preoxidation and as preoxyl radical scavengers. Arc Biochem Biophy 315: 161-169.
- Akocak S, Lolak N, Tuneg M, Boga M (2019) Antioxidant, acetylcholinesterase and butyrylcholinesterase inhibition profiles of histamine Schiff bases. J Turk Chem Soc Sect A: Chem 6(2): 157-164.
- Wang WT, Liao SF, Wu ZL, Chang CW, Wu JY (2020) Simultaneous study of antioxidant activity, DNA protection and anti-inflammatory effect of Vernonia amygdalina leaves extracts. PLoS One 15(7): e0235717.
- Reddy PR, Manjula P (2007) Mixed-ligand copper(II)-phenanthroline-dipeptide complexes: synthesis, characterization, and DNA-cleavage properties. Chemistry & Biodiversity 4(3): 468-480.
- Kirin SI, Happel CM, Hrubanova S, Weyhermüller T, Klein C, et al. (2004) Synthesis, structure and comparison of the DNA cleavage ability of metal complexes M(II)L with the N-(2-ethoxyethanol)-bis(2-picolyl) amine ligand L (M Co, Ni, Cu and Zn). Dalton Trans 21(8): 1201-1207.






























