Towards a Single Model for Antibiotics against Gram-Negative Bacteria
Johannes Zuegg*
Community for Open Antimicrobial Drug Discovery, University of Queensland, Australia.
Submission: November 30, 2018; Published: December 07, 2018
*Corresponding author: Johannes Zuegg, Community for Open Antimicrobial Drug Discovery, Institute for Molecular Bioscience, University of Queensland, 306 Carmody Road, Brisbane QLD 4072, Australia.
How to cite this article: Johannes Zuegg. Towards a Single Model for Antibiotics against Gram-Negative Bacteria. Nov Appro Drug Des Dev 2018; 4(3): 555640. DOI: 10.19080/NAPDD.2018.04.555640
Abstract
The discovery of novel antibiotics against Gram-negative bacteria is hampered by their highly effective defense mechanism these bacteria developed. These defense mechanism consists of a barrier of two membranes, preventing small molecules to penetrate, and of efficient efflux pump systems, removing any small molecule which succeeded in penetrating the two membranes. These mechanisms have been identified as main reason of failure of many antibiotic discovery efforts. Here we review and combine different approaches that investigated how small molecules penetrate the membranes and accumulate inside Gram-negative bacteria. Despite different approaches, common physicochemical properties have been identified, which overlap with earlier retrospective analysis of existing antibiotics. In general, the electrostatic property is the most significant property in each of the studies, with charged, polar molecules, or molecules with dipole moment, able to better penetrate and accumulate inside Gram-negative bacteria. In addition, the size of molecules displayed a limiting effect in most studies, with smaller molecules able to better penetrate the bacterial membrane. The combination of these studies confirm in general the current guidelines for a good model for Gram-negative antibiotics, while providing more detailed information on mechanism and variation between species.
Keywords: Antibiotics, Gram-Negative Bacteria, Membrane Penetration, Drug Discovery Model
Introduction
The antibacterial drug discovery pipeline has been in constant decline in the past decade, despite the increasing spread of resistant bacteria and the threat of making simple diseases, such as chronic infections after medical interventions, untreatable [1,2]. Especially, against Gram-negative bacteria, which are intrinsically resistant to most existing antibiotics, no new class of antibiotic has been introduced since 1968 [3], even though Gram-negative bacteria possess two cellular membranes, with the outer membrane coated with lipopolysaccharides, making it a formidable barrier for small molecules [4,5]. The penetration of this outer membrane is mainly achieved through narrow protein channels, called porins [6]. In addition, once inside, most small molecules are subjects to efflux pump systems, pumping them out again [7]. Some Gram-negative bacteria, like Pseudomonas sp., possess thereby a quite elaborate and redundant efflux pump systems, proving a highly efficient removal system for small-molecules, before they can reach a concentration high enough to effective [8].
The problem of Gram-negative membrane penetration has been recognized as key problem in the discovery of novel antibiotics, especially in many failed projects using target-based approaches [1]. However, even phenotypic approaches, using whole cell growth inhibition assays, did not yield the desired outcome, as shown by large screening efforts by several large biotech companies. Only recently, specific efforts has been undertaken to collect more understanding on the physicochemical properties required for small molecules to accumulate within Gram-negative bacteria. This review analyses the different studies and combines their outcome to define common key properties affecting the intra-cellular concentration of small molecules, and thereby their potential activity. The aim is to define a comprehensive rule or guideline, assisting medicinal chemists to discover and develop novel antibiotic against Gram-negative bacteria.
Statistical Analysis of Existing Antibiotics
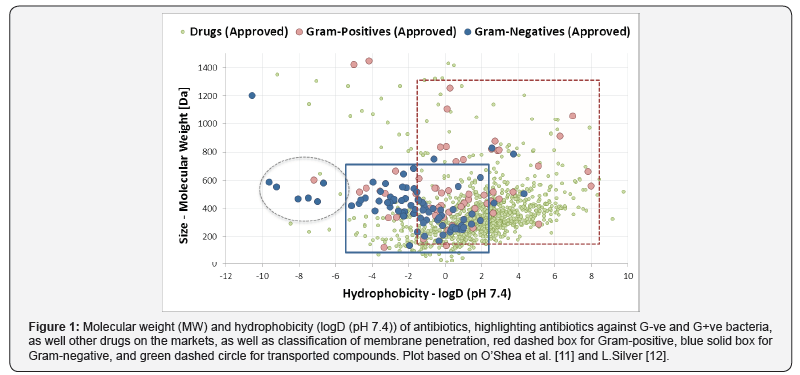
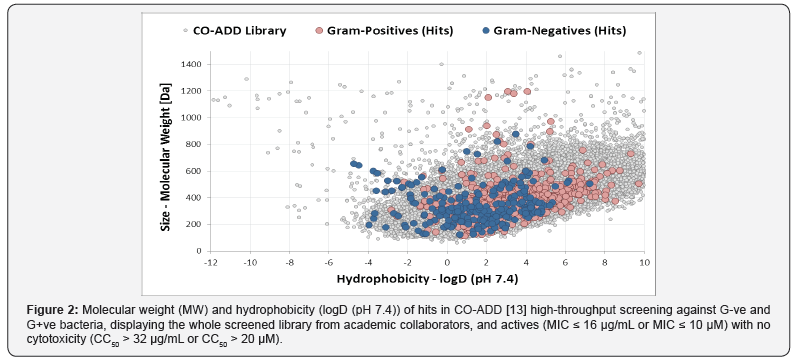
The vast majority of the current antibiotics on the market are natural product or derived from natural products. In this sense, it is of no surprise that antibiotics as a class do not fit within the concept of drug-likeness. Most antibiotics are larger in size and display higher polarity compared to other drugs, in such an extent that, back in 2001, the Rule of Five [9] which tried to define the optimal properties for oral bioavailability, had to exclude the whole class of antibiotics. Two decades on the properties of drugs on the market shifted even further away from antibiotics, further into more hydrophobic properties [10]. This unique property of antibiotics has been explored and defined in 2008 by O’Shea and Moser, comparing in more details antibiotics against other drug [11]. This analysis highlighted that antibiotics are, to a certain extent, larger and, more significantly, more hydrophobic compared to drugs against mammalian targets (Figure 1). An analysis by Lynn [12] recently added information on membrane penetration, but in essence confirming the earlier outcome, that successful drugs against G-ve bacteria have an optimal size below 600 Da and have rather hydrophilic properties, with logD below 2. Recent high-throughput screening data by the Community for Open Antimicrobial Drug Discovery (CO-ADD) [13], testing compounds from 250 academic collaborators around the world, against a range of Gram-negative and Gram-positive bacteria, showed a similar trend for its hits against Gram-negative bacteria, with a preference for more polar compounds (Figure 2). This limitation on hydrophobicity and molecular size remains currently the only guideline to filter potential antibiotics from a library of compounds.
Analysis of Intra-Cellular Concentration
Penetrating the membrane of bacteria has been identified early on as key requirement for successful antibiotics [1,14]. Especially the two-membrane system of Gram-negative bacteria is an efficient barrier [15], with two membrane with orthogonal properties, the outer membrane permeable for polar molecules while the inner membrane permeable for hydrophobic molecules. In addition, membranes of Gram-negative bacteria possess efflux pump systems providing a very efficient removal of compounds from the bacteria with very broad selectivity. One approach to gain more information on how Gram-bacteria are able to remove compounds, is to directly measure the concentration of compounds inside the bacteria.
Measuring intra cellular concentration of compounds in bacteria has so far been limited to radiolabeled [16] or fluorescent [17,18] molecules. While radiolabeling molecules is impractical for large number of compounds, adding a large fluorescent group to a molecule will affect its penetration and efflux properties significantly. Fluoroquinolones, which are the only antibiotics with intrinsic fluorescence, ad have been used in early studies to investigate intracellular accumulation [19-21]. Several studies have been reported using LC-MS/MS methods to measure the accumulation of non-fluorescent molecules in different bacteria [22-26], and while the method is time consuming, recent progress in LC-MS/MS technology has allowed to scale the workflow to analyze larger number of compounds [27]. Most studies have been conducted only on a limited number of different organisms, with most studies applied to Escherichia coli, with a few including Mycobacterium smegmatis, Pseudomonas aeruginosa or Staphylococcus aureus. A common limitation of these studies is the uncertainty on the precise location of the analyzed compounds, if they are located in the cytosol, in the periplasm or within the lipid bilayer of the membranes.
The most recent study [28] analyzed 180, mostly non antibiotic compounds measuring the accumulation of the compounds in Escherichia coli. The study further used the data of the original amines and 80 further selected amines, to build a predictive model, indicating physicochemical properties affecting the intracellular concentration. The properties include globularity, rotatable bonds and the present of a positive charge in the form of an amine. A follow up report [29] provides access to an online tool for their Entry Rule and some analysis on existing cases in which adding an amine made the compound active against Gram-negative bacteria, like Penicillin G, a Gram-positive active antibiotic and Ampicillin (Penicillin G with a amine group) having Gram-negative activity as well.
Structural Analysis and Simulations of Porins
The outer membrane (OM) of Gram-negative bacteria does not only consist of an asymmetric lipid bilayer decorated with large lipopolysaccharides (LPS) on its outside, but the membrane contains a large number protein, including lipoproteins and channels, which among other functions channel nutrients across the membrane [6,15]. These β-barrel proteins, called porins, have been the target of many investigations as they are responsible to get many antibiotics across the outer membrane [30]. In particular, non-specific porins OmpF and OmpC from Escherichia coli have been investigated, with many protein crystal structures available today [31]. Similarly, several Molecular Dynamics (MD) simulations have been conducted to analysis the properties and selectivity of such porins [32-35].
While most these studies were specific for certain porin structure or a compound types, a recent study [36]. used a wider simulation approach of eight prion structures, both trimeric OmpF and OmpC from E. coli, and their orthologues from E. cloacae, K. pneumonia and E. aerogenes. The study was done in combination with nine antibiotics, using MD simulation for each porin and antibiotic combination and building a scoring function for porin penetration based on physicochemical properties of the antibiotics. The properties included molecular size, total charge and transversal dipole, giving a good correlation with experimental penetration. The molecular size is thereby limited by the overall size of the channel, while the charge and dipole requirements are attributed to a mainly polar ‘gate’ in the middle of the channel, controlling the diffusion of the compound through the channel by electro static and electric field interactions. A follow-up study [37] extended the analysis to monomeric porins of Pseudomonas aeruginosa, resulting in the same type of physicochemical properties describing the diffusion, but with different parameters, illustrating similar mode of diffusion but significant structural variations between different types of porin channels.
Cell-Based Analysis Using Membrane Mutant Strains
Bacterial mutant strains are widely used in various mechanistic, mode-of-action and mode-of-resistance studies. Mutant strain affecting the composition and permeability of the bacterial membrane have be used in particular to study the intrinsic resistance of Gram-negative bacteria due to penetration and efflux, by deleting various porin and efflux components [8,17,30,38]. Even though, deletion or addition of major membrane proteins affects the integrity and property of the membrane by itself, such studies are giving valuable information for the development of antibiotics [8].
A recent study used several mutant strains of E. coli and P. aeruginosa to investigate membrane penetration of 64 antibiotics (β-lactams and fluoroquinolones) by comparing the MIC values of wild type and corresponding mutant strains [39]. The strains included highly membrane permeable mutants, deleting either major efflux pumps systems or expressing an additional large nonselective pore. Using the data the study correlated the ratio of MIC between wild type and the different membrane permeable strain, with various physicochemical properties of the antibiotics. While there are some differences between the two species, electrostatic properties were identified as major property affecting the penetration of compound in the wild type. Additional properties included charge, rigidity and lipophilicity. The only limitation in the study was the application to a limited number of antibiotic classes.
Discussion
This review focused on current models or guidelines to select potential antibiotic compounds against Gram-negative bacteria. One of the key requirement for activity against Gram-negative bacteria is that the active molecule is not only able to cross both membranes of the bacteria, but also evades the many efflux pump systems these bacteria possess. From the early analysis of druglikeness is was evident that such antibiotic active compounds have different properties to the drugs against mammalian cells. A more detailed statistical analysis of antibiotics indicated that molecular size and hydrophobicity are two key properties distinguishing the antibiotics from other drugs, in particular antibiotics against Gram-negative bacteria. Low hydrophobicity and low molecular weight was for a while the only guidance for medicinal chemistry to design novel antibiotics.
With the failing of many target-based antibiotic drug development projects, it became more evident that there is a lack of knowledge on how antibiotics penetrate the membranes of bacteria, and especially, how inactive compounds get removed efficiently in many Gram-negative bacteria, even though they display good target inhibition in biochemical assays. Recently, several groups released new studies to fill that gap, providing additional guidance on which properties are specifically beneficial for molecules to enter a bacteria and remain in the bacteria long enough to act.
The studies reviewed here use either statistical methods of successfully marketed antibiotics, biophysical measurements of compound concentration inside a bacteria, in-silico methods to map the structural feature of major porins, or cell-based assays using mutant strain in with altered porins or efflux pump systems. While each method investigates slightly different aspects of membrane penetration, they are surprisingly similar in their outcome, in the sense that similar physicochemical properties have been identified to affect the activity of antibiotics.
The top properties identified by all studies to be important for activity against Gram-negative bacteria are related to the polar and electronic nature of the molecules, and include either charge, presence of amines, dipole moments or hydrophilicity. The porin studies suggest that the porin channels are more open for polar molecules or molecules with a dipole moment. The cell-based membrane mutant approach displays a similar outcome for porins, especially in case of P. aeruginosa, but also adds lipophilicity as positive property for efflux, indicating fast efflux of molecules with high lipophilicity. A further common antibiotic property is molecular size with a limitation in the size of the molecules for good activity. The porin simulation studies and the statistical analysis of drugs on the market identified such size limitation, while the other studies contained molecules with small size variations. Never the less, this limitation on molecular size is most likely related to the size of the prion channels.
Globularity (ratio between number of sp2 and sp3 atoms) and VDW potential (non-bonding interaction potential) are two properties identified in the intra-cellular concentration study and cell-based mutant study to affect the intra-cellular concentration and activity, respectively. For both, smaller values are preferred for small molecules with antibiotic activity. Higher values for VDW potential are thereby related to higher efflux of the molecule. Both properties can be, to a certain extent, related to the size of the molecule and its topography of flat versus nonflat structure. Flexibility, on the other hand, has been identified in with contractive results, in two of the studies, the intra cellular concentration study and the cell-based membrane mutant approach. The former study identified rigidity to be detrimental to the activity, as rigid molecules are more subject to efflux pump system, while the second study showed a benefit with higher intra cellular concentration for rigid molecules.
Conclusion
The penetration of small molecules through the membranes of Gram-negative bacteria and their accumulation inside the bacteria, remains one of the key problems in the discovery of novel antibiotics against these classes of bacteria. Recently, several new approaches have been applied to fill the knowledge gap on how small molecule can penetrate the membrane through their porins, and how these molecules get removed again from the bacteria through their efflux pump system. Each used a different approach, ranging from biophysical measurements of compound concentration inside a bacteria, structure based simulation of porins to cell-based studies using membrane mutant strains. Remarkably, despite the different approaches, common physicochemical properties have been identified for potential antibiotic activity, confirming and expanding ‘antibiotic’ properties from earlier studies.
All studies identify electrostatic properties as the most significant properties, aiding the penetration of the bacteria membrane and the accumulation inside the bacteria. Electrostatic properties, like total charge, polarity, dipole moment or just in the presence of an amine group, increase the penetration and accumulation of a small molecule. From these studies, porins prefer polar molecules, while efflux pump system are more effective against lipophilic molecules. The second most common property class is related to the size and shape of the molecule. The studies indicate a clear limit in molecular for the penetration through porins, with some minor indication that efflux pumps are more tolerant. All recent studies overlap with early analysis of existing antibiotics highlighting similar properties for antibiotic activity. However, they provide more detailed information on the mechanism of penetration and efflux, as well as highlight significant variation between different species of Gram-negative bacteria. While they define in more detailed a common model for antibiotic, they also highlight the need to expand the studies with more chemical diversity and different organisms.
Acknowledgement
The antibacterial screening, presented here as a summary, has been conducted by Community for Open Antimicrobial Drug Discovery, funded by the Wellcome Trust and the University of Queensland, with special acknowledgement to the members of CO-ADD: Alysha Elliott, Ali Hinton, Angela Kavanagh, Gabrielle Lowe, Janet Reid, Maite Amato, Soumya Ramu, Robin Trauer, Karl Hansford, Mark Blaskovich, Ruth Neale, Mathilde Desselle and Matthew Cooper.
References
- Payne DJ, Gwynn MN, Holmes DJ, Pompliano DL (2007) Drugs for bad bugs: confronting the challenges of antibacterial drug discovery. Nat Rev Drug Discov 6(1): 29-40.
- Fischbach MA, Walsh CT (2009) Antibiotics for emerging pathogens. Science 325(5944): 1089-1093.
- Lewis K (2013) Platforms for antibiotic discovery. Nat Rev Drug Discov 12(5): 371-387.
- Nikaido H (1994) Prevention of drug access to bacterial targets: permeability barriers and active efflux. Science 264(5157): 382-388.
- Nikaido H (2003) Molecular basis of bacterial outer membrane permeability revisited. Microbiol Mol Biol Rev 67: 593-656.
- Nikaido H (1994) Porins and specific diffusion channels in bacterial outer membranes. J Biol Chem 269(6): 3905-3908.
- Silver LL (2011) Challenges of antibacterial discovery. Clin Microbiol Rev 24(1): 71-109.
- Schweizer HP (2012) Understanding efflux in Gram-negative bacteria: opportunities for drug discovery. Expert Opin Drug Discov 7(7): 633- 642.
- Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (2001) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 46(1- 3): 3-26.
- Shultz MD (2018) Two Decades under the Influence of the Rule of Five and the Changing Properties of Approved Oral Drugs. J Med Chem.
- O’Shea R, Moser HE (2008) Physicochemical properties of antibacterial compounds: implications for drug discovery. J Med Chem 51: 2871- 2878.
- Silver LL (2016) A Gestalt approach to Gram-negative entry. Bioorg Med Chem 24(24): 6379- 6389.
- Blaskovich MA, Zuegg J, Elliott AG, Cooper MA (2015) Helping Chemists Discover New Antibiotics, ACS Infect Dis 1(1): 285-287.
- Jackson N, Czaplewski L, Piddock LJV (2018) Discovery and development of new antibacterial drugs: learning from experience? J Antimicrob Chemother 73: 1452-1459.
- Zgurskaya HI, Lopez CA, Gnanakaran S (2015) Permeability barrier of Gram-negative cell envelopes and approaches to bypass it. ACS Infect Dis 1: 512-522.
- Piddock LJV, Williams KJ, Ricci V (2000) Accumulation of rifampicin by Mycobacterium aurum, Mycobacterium smegmatis and Mycobacterium tuberculosis. Journal of Antimicrobial Chemotherapy 45: 159-165.
- Piddock LJV, Johnson MM (2002) Accumulation of 10 fluoroquinolones by wild-type or efflux mutant Streptococcus pneumoniae, Antimicrob Agents Chemother 46: 813-820.
- Piddock LJV, Ricci V (2001) Accumulation of five fluoroquinolones by Mycobacterium tuberculosis H37Rv. J Antimicrob Chemother 48: 787- 791.
- Bazile S, Moreau N, Bouzard D, Essiz M (1992) Relationships among antibacterial activity, inhibition of DNA gyrase, and intracellular accumulation of 11 fluoroquinolones. Antimicrob Agents Chemother 36: 2622-2627.
- Asuquo AE, Piddock LJ (1993) Accumulation and killing kinetics of fifteen quinolones for Escherichia coli, Staphylococcus aureus and Pseudomonas aeruginosa. J Antimicrob Chemother 31: 865-880.
- McCaffrey C, Bertasso A, Pace J, Georgopapadakou NH (1992) Quinolone accumulation in Escherichia coli, Pseudomonas aeruginosa, and Staphylococcus aureus, Antimicrob Agents Chemother 36: 1601- 1605.
- Nikaido H, Rosenberg EY, Foulds J (1983) Porin channels in Escherichia coli: studies with beta-lactams in intact cells. J Bacteriol 153: 232-240.
- Zhou Y, Joubran C, Miller-Vedam L, Isabella V, Nayar A, et al. (2015) Thinking outside the “bug”: a unique assay to measure intracellular drug penetration in gram-negative bacteria. Anal Chem 87: 3579-3584.
- Davis TD, Gerry CJ, Tan DS (2014) General platform for systematic quantitative evaluation of small-molecule permeability in bacteria. ACS Chem Biol 9: 2535-2544.
- Cai H, Rose K, Liang LH, Dunham S, Stover C (2009) Development of a liquid chromatography/mass spectrometry-based drug accumulation assay in Pseudomonas aeruginosa. Anal Biochem 385: 321-325.
- Bhat J, Narayan A, Venkatraman J, Chatterji M (2013) LC-MS based assay to measure intracellular compound levels in Mycobacterium smegmatis: linking compound levels to cellular potency. J Microbiol Methods 94: 152-158.
- Gordon LJ, Allen M, Artursson P, Hann MM, Leavens BJ, et al. (2016) Direct Measurement of Intracellular Compound Concentration by RapidFire Mass Spectrometry Offers Insights into Cell Permeability, J Biomol Screen 21: 156-164
- Richter MF, Drown BS, Riley AP, Garcia A, Shirai T, et al. (2017) Predictive compound accumulation rules yield a broad-spectrum antibiotic. Nature 545: 299.
- Richter MF, Hergenrother PJ (2018) The challenge of converting Grampositive- only compounds into broad-spectrum antibiotics. Ann N Y Acad Sci.
- Pages JM, James CE, Winterhalter M (2008) The porin and the permeating antibiotic: a selective diffusion barrier in Gram-negative bacteria. Nat Rev Microbiol 6: 893-903.
- Galdiero S, Falanga A, Cantisani M, Tarallo R, Della Pepa ME, et al. (2012) Microbe-host interactions: structure and role of Gram-negative bacterial porins. Curr Protein Pept Sci 13: 843-854.
- Bond PJ, Sansom MS (2004) The simulation approach to bacterial outer membrane proteins. Mol Membr Biol 21: 151-161.
- Pavlova A, Hwang H, Lundquist K, Balusek C, Gumbart JC (2016) Living on the edge: Simulations of bacterial outer-membrane proteins. Biochim Biophys Acta 1858: 1753-1759.
- Pothula KR, Solano CJ, Kleinekathofer U (2016) Simulations of outer membrane channels and their permeability. Biochim. Biophys Acta 1858: 1760-1771.
- Furini S, Domene C (2016) Computational studies of transport in ion channels using metadynamics, Biochim. Biophys Acta 1858: 1733- 1740.
- Acosta-Gutierrez, Ferrara L, Pathania M, Masi M, Wang J S, et al. (2018) Getting Drugs into Gram-Negative Bacteria: Rational Rules for Permeation through General Porins. ACS Infect Dis 4: 1487-1498.
- Samanta S, Bodrenko , Acosta-Gutiérrez S, D’Agostino T, Pathania M, et al (2018) Getting Drugs through Small Pores: Exploiting the Porins Pathway in Pseudomonas aeruginosa. ACS Infect Dis 4: 1519-1528.
- Kumar A, Schweizer HP (2005) Bacterial resistance to antibiotics: active efflux and reduced uptake. Adv Drug Deliv Rev 57: 1486-1513.
- Cooper SJ, Ganesh Krishnamoorthy, David Wolloscheck, John K Walker , Valentin V Rybenkov, et al. (2018) Molecular Properties That Define the Activities of Antibiotics in Escherichia coli and Pseudomonas aeruginosa, ACS Infect Dis 4: 1223-1234.






























