- Research Article
- Abstract
- Introduction
- Materials and Methods
- Methods
- Software Applications
- Library Design of Ligands
- Preparation of Molecular Target
- Docking Studies
- Binding Analysis
- Result and Analysis of Data
- Discussion
- Predicting Binding Affinity from Docking Scores
- Binding Pattern Analysis
- Conclusion
- References
In vivo Studies of the Effect of a Herbal Preparation on Pharmacokinetic Properties of Rifampicin in Male Swiss Albino Rats and Computer Mediated Mechanism of Interaction
Atuanya Vivian Adaeze1, Poyi Olanike C2, Kolawole Jacob A1*
1Department of Pharmaceutical Chemistry, Faculty of Pharmaceutical Sciences, University of Jos, Nigeria
2Department of Pharmaceutical and Medicinal Chemistry, Faculty of Pharmaceutical Sciences, Kaduna state University, Nigeria
Submission: February 15, 2017; Published: February 23, 2018
*Corresponding author: Jacob A Kolawole, Department of Pharmaceutical Chemistry, Faculty of Pharmaceutical Sciences, University of Jos, Nigeria, Tel: +2348027144522; Email: kolajac@yahoo.com
How to cite this article: Atuanya V A, Poyi Olanike C, Kolawole Jacob A. In vivo Studies of the Effect of a Herbal Preparation on Pharmacokinetic Properties of Rifampicin in Male Swiss Albino Rats and Computer Mediated Mechanism of Interaction. Nov Appro Drug Des Dev 2018; 3(4): 555620. DOI: 10.19080/NAPDD.2018.03.555620
- Research Article
- Abstract
- Introduction
- Materials and Methods
- Methods
- Software Applications
- Library Design of Ligands
- Preparation of Molecular Target
- Docking Studies
- Binding Analysis
- Result and Analysis of Data
- Discussion
- Predicting Binding Affinity from Docking Scores
- Binding Pattern Analysis
- Conclusion
- References
Abstract
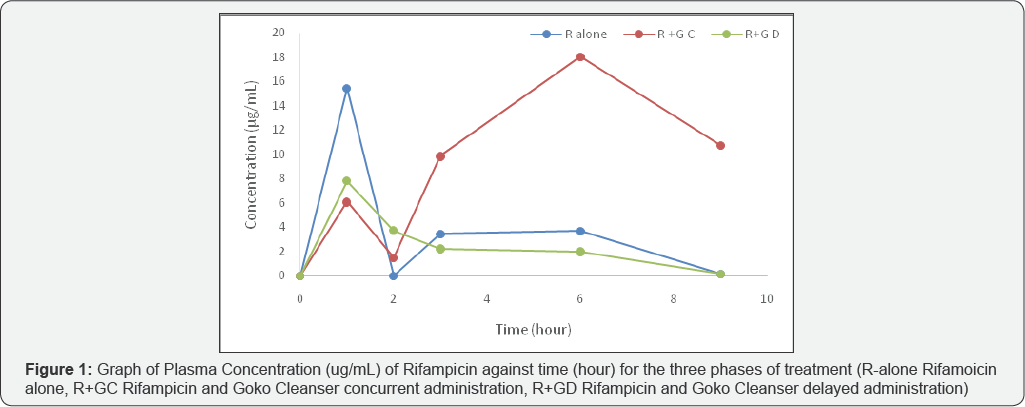
This study investigates the effect of aherbal preparation (‘Goko’ Cleanser], on the pharmacokinetic profile of orally administered Rifampicin in Swiss albino male rats. The In vivo study was carried out in three phases. In phase I only Rifampicin (20mg/kg body) was orally administered to ratsand in phase II, Rifampicin and Goko Cleanser (30ml/kg body weight) were concurrently administered and in phase III Goko Cleanser was administered for 6 days before Rifampicin. Blood samples were collected from rats in each group at 0, 1, 2, 3, 6, 9 and 12 hours in each phase and concentration determined by spectrophotometric method. Pharmacokinetics parameters determined include, Cmax, Tmax, t1/2α, ClT, AUC0-9, AUC0-∞, t1/2β and kβ. The In-silico studies was done by docking molecules contained in the herbal cleanser into rifampicin binding site on the Pregnane X receptor (PXR). The result showed Rifampicin pharmacokinetics profile to be in line with previous reports. Goko Cleanser significantly (P<0.05, 0<0.01) increase the Cmax, Tmax, AUC0-9, AUC0-∞, and t1/2β but decrease ClT, when administer concurrently with Rifampicin (Phase II). Goko Cleanser significantly (P<0.05) decrease the Cmax, Tmax, t1/2α, AUC0-9, AUC0-∞, and t1/2β but increase ClT when Rifampicin was administered after 6 days of taking Goko Cleanser (Phase III). In silico docking results showed a significant interaction of Goko Cleanser constituents with active site binding residues and complex formation between constituents of Goko cleanser and rifampicin when administered together. Taking Goko cleanser with rifampicin at the same or within a week should be avoided..
but decrease ClT, when administer concurrently with Rifampicin (Phase II). Goko Cleanser significantly (P<0.05) decrease the Cmax, Tmax, t1/2α, AUC0-9, AUC0-∞, and t1/2β but increase ClT when Rifampicin was administered after 6 days of taking Goko Cleanser (Phase III). In silico docking results showed a significant interaction of Goko Cleanser constituents with active site binding residues and complex formation between constituents of Goko cleanser and rifampicin when administered together. Taking Goko cleanser with rifampicin at the same or within a week should be avoided..
Keywords: In vivo; In Silico; Rifampicin; Drug-Drug Interaction; Pharmacokinetics
Abbreviations: SD: Standard Derivation; PXR: Pregnane X receptor; MM2: Molecular Mechanics geometry; RSD: Relative Standard Deviation
- Research Article
- Abstract
- Introduction
- Materials and Methods
- Methods
- Software Applications
- Library Design of Ligands
- Preparation of Molecular Target
- Docking Studies
- Binding Analysis
- Result and Analysis of Data
- Discussion
- Predicting Binding Affinity from Docking Scores
- Binding Pattern Analysis
- Conclusion
- References
Introduction
The use of herbs as medicines dates far back to the origin of man civilization with the earliest known record inscribed on six thousand year old clay written by Sumeritans. China and India are the two countries were the use of herbs as medicine has been integrated into the modern health care. There is increasing demand for herbal medicines especially as dietary supplements in Africa, Asia, Europe and USA (Green world, 2010).This makes occurrence of drug interaction between an orthodox drug and herbal preparation more rampant. Drug interactions are broadly classified into two aspects based on the mechanism/mode of interaction-pharmacologic or pharmacokinetic. Pharmacokinetic profiling of a drug molecule enables an accurate assessment of the amount of drug found in the plasma with respect to time (Hadijah, 2013). In silico methods refer to methods involving computational approaches used in fast predictions (Amberg) [1]. To prevent drug-drug interactions, several approaches involving chemoinformatics and physiologically pharmacokinetics based systems have been put in place at different stages of drug development process. Ni A et al. [2], were able to illustrate the usefulness of in silico methods in the prediction of drug- drug interactions and drug interactions with other substances.
Rifampicin is a powerful known inducer of the hepatic cytochrome P450 enzyme system, including iso-enzymes CYP2B6, CYP2C8, CYP2C9, CYP2C19, CYP3A4, CYP3A5, and CYP3A7. Rifampicin has been reported to interact with a number of drugs, atorvastatin, rosiglitazone, pioglitazone, celecoxib, Clarithromycin, caspofungin, voriconazole and lorazepam. The human Pregnane X Receptor is a known inductive regulator of the cytochrome P-450 monooxygenase 3A4 (CYP3A4) which is responsible for the oxidative metabolism of many drugs including rifampicin [3]. The report of Lehmann [4], provided a platform in aiding to predict drug- drug interactions in humans as they provided a molecular explanation for the ability of incongruent chemical entities to induce CYP3A4. Orally administered rifampicin is well absorbed from the gastrointestinal tract and have a reported Cmaxof about 2-4 hours [5]. Food consumption inhibits its absorption from the GI tract, and the drug is more quickly eliminated. About 60% to 90% of the drug is bound to plasma proteins (Rofel). The half-life of rifampicin ranges from 1.5 to 5.0 hours.
Goko Cleanser is an oral herbal preparation labeled to be used in the treatment of many disease conditions including the treatment of infections. The label claim showed that it contained five (5) different herbs which includes Vernoniaamygdalina (12%), Saccharum officinalis (11.5%), Allium sativa (13%), and Cajantus cajan (11.5%). Reported phytochemical composition of Vernonia amygdalina (12%) include; terpenes, saponins, phenolic acids,edotides, xanthones [6]. Saccharumofficinalis (11.5%)commonly known as sugar cane contains phenolic acids, -C- and -O-glycosides,terpenoids, phytosterols, fatty acids and flavonoids responsible for its antioxidant activity [7]. Allium sativa (13%), (garlic) contains phenolic compounds and organosulfur compounds such asallicin, alliin, S-allyl- cysteine sulfoxides which are the source of its antioxidant activity [8], Cajantuscajan (11.5%) (pigeon pea), contains crude proteins(21%), crude fat(1.7%), fiber(2.5%) and moisture(8%) [9]. Label claims of Goko Cleanser include "treatment of fertility, aphrodisiac, urinary tract diseases, infectious diseases, stabilizing menstrual period and other conditions". Also to be administered (30ml/60kg body weight) orally on empty stomach. There is no report in the literature, to our knowledge of drug interaction of a single preparation with five active plant component with a strong CYP 450 inducer and substrate such as Rifampicin. In view of the complexity of Rifampicin in drug interaction with the CYP 450 family and the multiple component of these herbal preparation, this work is aimed at investigating the effect of Goko Cleanser (herbal preparation) on the pharmacokinetic property of oral Rifampicin in male Swiss albino rats and possible mechanism of the interaction (Table 1).
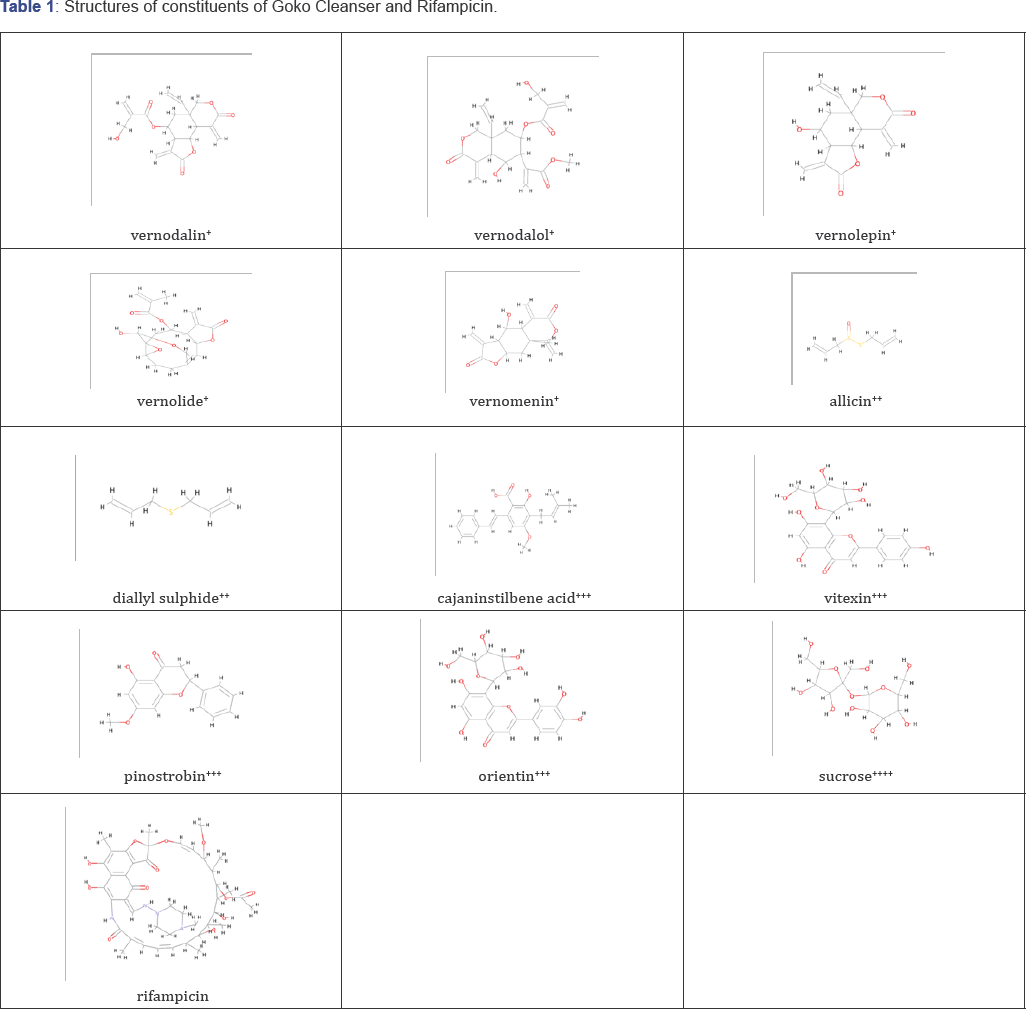
+constituents of Vernonia amygdalina (bitter leaf),
++constituents of Allium sativa (garlic),
+++constituents of Cajantus cajan (pigeon pea)
++++constituents of Saccharum officinalis (sugarcane),
*line structures created in BIOVIA Discovery Studio (Dassault Systemes, 2016).
- Research Article
- Abstract
- Introduction
- Materials and Methods
- Methods
- Software Applications
- Library Design of Ligands
- Preparation of Molecular Target
- Docking Studies
- Binding Analysis
- Result and Analysis of Data
- Discussion
- Predicting Binding Affinity from Docking Scores
- Binding Pattern Analysis
- Conclusion
- References
Material and Methods
The following pieces of equipment and materials were used; UV-Spectrophotometer (JENWAY 6405 UV-VIS Spectrophotometer) connected to a HP computer system loaded with UV-probe, Blood sample bottles pretreated with EDTA, Rifampicin capsule, (at a dose of 20mg/Kg) and Goko Cleanser(at a dose 30ml/60Kg). All Chemical and Reagents were supplied by Dept. of Pharmaceutical Chemistry, University of Jos, and are of analytical grade; Zinc Sulphate powder, Ascorbic acid powder, EDTA, Isopropanol, Acetonitrile, Chloroform. (Baker Analyzed Reagents of J.J. Baker Chemical Co. Phillipsburg)
- Research Article
- Abstract
- Introduction
- Materials and Methods
- Methods
- Software Applications
- Library Design of Ligands
- Preparation of Molecular Target
- Docking Studies
- Binding Analysis
- Result and Analysis of Data
- Discussion
- Predicting Binding Affinity from Docking Scores
- Binding Pattern Analysis
- Conclusion
- References
Methods
Preparation of Reagents
Zinc sulphate and ascorbic reagent. A 0.805g and 0.88g of zinc sulphate and ascorbic acid powdered respectively were weighed and dissolved in 10ml of distilled water. Rifampicin stock solution(200μg/100ml) rifampicin powder (20mg) was dissolved with (500μl) of DMSO and made up to 100ml.
Animal Treatment and Sampling
The study was divided into three phases with each phase having 35 male rats and weighing about 150g each. Each group comprise 5 rats according to the time of collection of blood samples post rifampicin administration time of 0 and 1,2,3,6,9 and 12 hours. The rats were fasted for at least 3 hours prior to rifampicin administrations in each phase. Rifampicin was administered at a dose of 20mg/kg body weight and Goko cleanser was administered at a dose of 30ml/kg body weight of the rats.
Phase I
Rifampicin was administered alone and samples collected at the above time intervals. Blood samples were centrifuged at 4500rpm for 15 minutes and plasma was harvested and stored at -20°C until analysis.
Phase II
Goko Cleanser was administered concurrently with rifampicin followed by post rifampicin samples collections.
Phase III
Goko cleanser was administered alone for 6 days and on the 7th day Goko cleanser was administered 1 hour prior to rifampicin administration, followed by post rifampicin sample collections as in first phase.
Calibration Curve and Method Validation Process
Plasma from ten rats was pooled for the standard calibration curve. The plasma was spiked with Rifampicin solution to obtain 5μg/ml, 10ug/ml, 15ug/ml, 20ug/ml and 25ug/ml concentrations respectively. The method of Milton, (2015) was adopted, modified and validated. Rifampicinwas extracted from plasma (0.5ml) by adding 0.5ml of 0.5M ZnSO4, 2ml of acetonitrile: 2-propanol (1:1 v/v) and 0.5ml of Ascorbic acid and vortex mixed for 2minutes. The mixture was centrifuged at 3500 rpm for 10 minutes and the supernatant collected for immediate UV Spectrophoto metric assay where absorbance was take at 337nm. Absorbance was plotted against concentration for the calibration curve. The calibration curve was repeated on five consecutive days and the average values used to generate a 5-day pooled calibration curve. The regression line equation and correlation coefficient were obtained from the calibration curve using linear regression analysis.
Validation of Analytical Method
The procedure above was repeated five times for the purpose of validation, on the same day (within day precision) and on five different days (day to day precision). A 0.5 ml of plasma each was spiked with the working solutions 5, 10, 15, 20 and 25 Hg/ml of rifampicin and analyzed using the analytical method. Absorbencies were obtained and percentage recovery as well as standard derivation (SD) calculated from the corresponding concentration was obtained.
Recovery
Three selected concentrations of 1ml of stock (rifampicin) as in the calibration curve determination were spiked into 0.5 ml each of pooled plasma. The samples were analyzed as in calibration curve and the absorbencies were measured. The corresponding concentrations were obtained and the percentage recoveries were calculated and shown in table .
Precision and Accuracy
Concentration range of 5.0-25.0 (ng/mL) of rifampicin solution were spiked with 0.5ml each of pooled plasma and analysed using the analytical method. This procedure was repeated for five different days, the corresponding concentrations were obtained from the calibration curve. The standard deviation was calculated to ascertain the precision as shown in table.
Linearity
Three independent readings of the calibration procedure were carried out and the mean obtained.
Plasma Concentration Determination
After collection of samples, they were centrifuged at 4500rpm for 15 minutes for the separation of plasma, relabeled and refrigerated prior to use. A 0.5ml of plasma was decanted into a test tube, 0.5ml of 0.5M ZnSO4, 2ml of acetonitrile: 2-propanol (1:1 v/v) and 0.5ml of Ascorbic acid were added; then vortex mixed for 2minutes and centrifuged at 3500rpm for 10 minutes, the supernatant collected for UV Spectrophotometric assay at a wavelength of 337nm. The measured absorbance for each sample was used to extrapolate the concentration of the drug at the time of collection.
Determination of Pharmacokinetics Parameters
The pharmacokinetic parameters (Cmax, Tmax, AUC, Absorption and Elimination half- life) were determined as follows: Absorption half-life; This was obtained from the graph, the slope was calculated and Kab determined using; S= k/-2.303, T1/2( ab ) =°.693/Kab
Where; S = Slope, Kab= absorption rate constant.
Elimination half-life; The slope of the elimination was calculated, then KEL and T.1/2(el)was determined
Tmax, Cmax were determined from the Pharmacokinetic curve.And trapezoid method used for AUC. This was calculated using AUCto= 1/2(a +b) h
Clearance (Clt), The clearance was calculated using (Clt) = Dose/AUC
Statistical analysis was done using Graphpad online.
Docking Studies
All methods used in the docking study were entirely Insilco, making use of several web interfaces, online databanks and software applications.
Databanks
Protein databank (www.rcsb.org): a database providing information of proteins was used to obtain the x-ray structure of Pregnane X Receptor, 1skx.pdb [10].
- Research Article
- Abstract
- Introduction
- Materials and Methods
- Methods
- Software Applications
- Library Design of Ligands
- Preparation of Molecular Target
- Docking Studies
- Binding Analysis
- Result and Analysis of Data
- Discussion
- Predicting Binding Affinity from Docking Scores
- Binding Pattern Analysis
- Conclusion
- References
Software Applications
BIOVIA Discovery Studio: a programme to examine structure properties of both large and small molecules [11]. ArgusLab 4.0.1: software for molecular modelling and drug designing [12].
- Research Article
- Abstract
- Introduction
- Materials and Methods
- Methods
- Software Applications
- Library Design of Ligands
- Preparation of Molecular Target
- Docking Studies
- Binding Analysis
- Result and Analysis of Data
- Discussion
- Predicting Binding Affinity from Docking Scores
- Binding Pattern Analysis
- Conclusion
- References
Library Design of Ligands
A set of compound library of 12molecules which make up some chemical constituents of goko cleanser and rifampicin structures were downloaded from the Pubchem Compound database and molecular mechanics geometry optimization performed on them in BIOVIA Discovery studio visualize.
- Research Article
- Abstract
- Introduction
- Materials and Methods
- Methods
- Software Applications
- Library Design of Ligands
- Preparation of Molecular Target
- Docking Studies
- Binding Analysis
- Result and Analysis of Data
- Discussion
- Predicting Binding Affinity from Docking Scores
- Binding Pattern Analysis
- Conclusion
- References
Preparation of Molecular Target
The crystal structure of Pregnane X receptor (PXR) with the pdb code 1skx. Pdb was downloaded from the protein databank. The pdb file was opened in WordPad, the water molecules were deleted and the file re-saved. The valance and hybridisation of the PXR structure was checked in Argus Lab. The structure was optimized by locking all atoms except the hydrogens, and Molecular Mechanics geometry (MM2) performed on it. The active site in PXR structure was defined by selecting residues in the hydrophobic pocket of the A chain that were bounded to the rifampicin in the crystral structure and grouped as the binding site.
- Research Article
- Abstract
- Introduction
- Materials and Methods
- Methods
- Software Applications
- Library Design of Ligands
- Preparation of Molecular Target
- Docking Studies
- Binding Analysis
- Result and Analysis of Data
- Discussion
- Predicting Binding Affinity from Docking Scores
- Binding Pattern Analysis
- Conclusion
- References
Docking Studies
The library of molecules were docked into the active site of PXR using Arguslab (spread sheet style batch dockings). A fast Dock settings of flexible ligands in flexible (for goko cleanser constituents alone) and a rigid (for both goko cleanser constituents and rifampicin) active site- conditions in a 3000 step generation using Amber Van der Waals in an active box of 15x 15x 15Ao was performed with a maximum of 150 best poses. AS core scoring function was used in this study. Subsequently, the library of compound constituents of goko cleanser were docked into the binding site of PXR which contained the rifampicin structure and re-docked a second time without the rifampicin in the active site.
- Research Article
- Abstract
- Introduction
- Materials and Methods
- Methods
- Software Applications
- Library Design of Ligands
- Preparation of Molecular Target
- Docking Studies
- Binding Analysis
- Result and Analysis of Data
- Discussion
- Predicting Binding Affinity from Docking Scores
- Binding Pattern Analysis
- Conclusion
- References
Binding Analysiss
Binding interactions of the ligands with the PXR protein active site was analyzed using Argus Lab 4.0.1 and BIOVIA Discovery Studio software's.
- Research Article
- Abstract
- Introduction
- Materials and Methods
- Methods
- Software Applications
- Library Design of Ligands
- Preparation of Molecular Target
- Docking Studies
- Binding Analysis
- Result and Analysis of Data
- Discussion
- Predicting Binding Affinity from Docking Scores
- Binding Pattern Analysis
- Conclusion
- References
Result and Analysis of Data
Calibration Curve of Rifampicin in Plasma of Rats
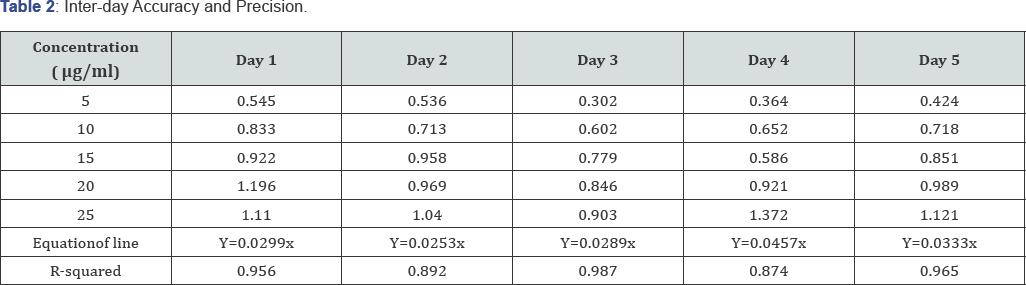
The calibration curve showed a concentration range of 0.025.0 ng/ml, regression equation of y = 0.0346x+ 0.2951 and r2 of 0.9807. The method was found to be reproducible as was observed by the absorbance obtained. The data obtained for accuracy and precision as presented in Table 1, shows that interday precision gave a standard deviation of 0.042 indicating good accuracy. The Relative Standard Deviation (RSD) was less than 2%, which indicates good repeatability. The Mean±S.D is 0.8947±0.0417 (Table 2).
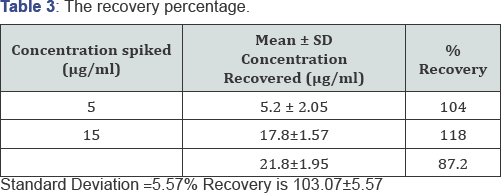
Recovery
Selected concentrations of stock as in the calibration curve determination were spiked into 0.5 ml of plasma. The samples were analyzed as in calibration curve and the absorbencies were measured. The corresponding concentrations were obtained and the percentage recoveries were calculated (Tables 3 & 4) (Figure 1).
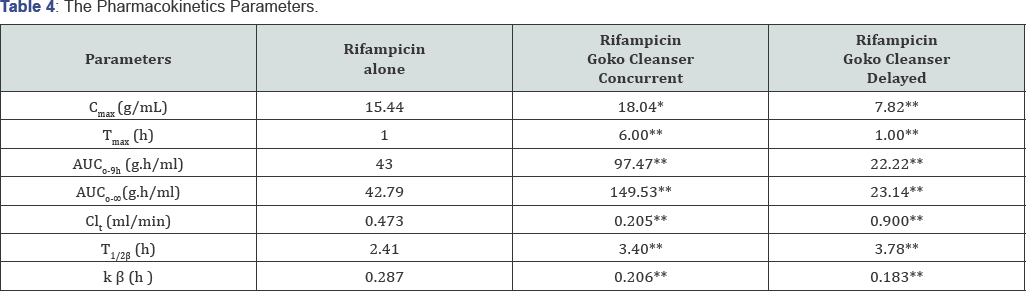
KEY: *P≤ 0.05(quite statistically significant difference); **P extremely < 0.05(extremely statistically significant difference).
BINDING AFFINITY
- Research Article
- Abstract
- Introduction
- Materials and Methods
- Methods
- Software Applications
- Library Design of Ligands
- Preparation of Molecular Target
- Docking Studies
- Binding Analysis
- Result and Analysis of Data
- Discussion
- Predicting Binding Affinity from Docking Scores
- Binding Pattern Analysis
- Conclusion
- References
Discussion
The method validation results are as obtained as presented in Tables 2-4. The calibration curve was found to be linear over a concentration range of 2.5.0-25.0μg/mL, with a correlation coefficient of 0.9857, this shows that Beer Lambert’s law was obeyed. The day to day accuracy and precision gave a relative standard deviation of less than 2% which shows that the modified method adapted is therefore suitable for comparative pharmacokinetics and drug-drug interaction study carried out. It is a reproducible, sensitive, simple, precise and accurate procedure though it’s quite expensive. The rate of oral absorption of rifampicin is not dose related, since the time required for the drug to reach its peak serum concentration has been found to be the same at different dosage levels Acocella, Lecaillon [13,14]. Rifampicin diffuses rapidly from plasma into the other body fluids, tissues and organs. In the liver, gall bladder and urine rifampicin reaches concentration which are higher than those found in the blood and salivary concentration are approximately 20% of the serum concentration [15]. The AUC0-24h of rifampicin in saliva was significantly lower than the serum concentration (i.e. the protein unbound plasma drug) [16]. This shows that rifampicin is rapidly absorbed via oral route. It has a peak plasma concentration between one to four hours; this is significantly altered by 36% by the presence of food [5]. Rifampicin has a mean plasma protein binding rate of 88.9±0.9% in healthy subjects [17,18], though binding has been found to be unrelated to the plasma concentration of the drug [19]. The maximum plasma concentration observed with the corresponding Tmax when rifampicin was given concomitantly with Goko cleanser is 18.04(μg/mL) at a Tmax of 6 h varied with respect to the control when rifampicin was given alone which is15.44(μg/mL) at a Tmax of 1 h, this indicates interaction at the absorption level of rifampicin as the p value (0.065) showed a quite statistically significant difference; where Goko cleanser complexes with rifampicin inhibiting it's absorption thereby prolonging the Tmax from one to four hours of oral administration to the sixth hour, the delay in the gastrointestinal absorption would further delay metabolism rate which is seen by a low clearance rate of 0.21(ml/min) resulting in a larger AUC0-9h as 97.47(μg.h/ml) seen in rifampicin concomitant administration, this shows that Goko cleanser altered both the extent and rate of absorption of rifampicin this confirmed the fact that administration of rifampicin with food altered the rate of absorption although this effect is also enhanced by the plasma protein binding property of rifampicin. From the decreased Cmax observed in the delayed effect where Goko cleanser was administered for 6 days prior to rifampicin administration on the 7th day with a p value extremely less than 0.05 , this shows that Goko cleanser activates this metabolic pathway via activating the following enzymes CYP2B6, CYP2C8, CYP2C9, CYP2C19, CYP3A4, CYP3A5, and CYP3A7 which are involve in the metabolism of rifampicin, this is also seen by a high clearance rate of 0.9(ml/ min) which indicates a higher volume of the drug is being cleared per minute. This also resulted in a smaller AUC0-gli which is 22.22(μg.h/ml) which can possibly results to antibiotic resistant (sub-therapeutic level). The increase in the elimination half-life of the treatment with Goko when administered concomitantly is due to complex formation which serves as reservoir for the drug which now takes a longer time to be reduced to half. And this is also the effect seen in the delayed phase [20-25].
Goko cleanser has both inhibitory effect on absorption and induction effect on the metabolism of Rifampicin although rifampicin is a very potent enzyme inducer which can also enhance its own metabolism (self-induced metabolism). The paired sample test shows the t-value to be less than 0.05 for all the rats at 95% level of confidence, indicating significant difference in treatment before, concurrently and after Goko cleanser. The paired sample correlation has a t-value less than 0.05, indicating significant relationship existing between Goko cleanser and Rifampicin. The human Pregnane X Receptor (PXR) detects a wide variety of structurally distint endogenous compounds with Rifampicin being one of the largest known ligands for the receptor. The work of Chrencik et al. [10], shows that one wall of the PXR binding site cavity can remain flexible even if it is in complex with an activating ligand, in this case rifampicin. Hence co-administration of rifampicin with other endogenous ligands could result in the interaction of PXR with both the rifampicin and the Goko cleanser constituents [26-30].
- Research Article
- Abstract
- Introduction
- Materials and Methods
- Methods
- Software Applications
- Library Design of Ligands
- Preparation of Molecular Target
- Docking Studies
- Binding Analysis
- Result and Analysis of Data
- Discussion
- Predicting Binding Affinity from Docking Scores
- Binding Pattern Analysis
- Conclusion
- References
Predicting Binding Affinity from Docking Scores
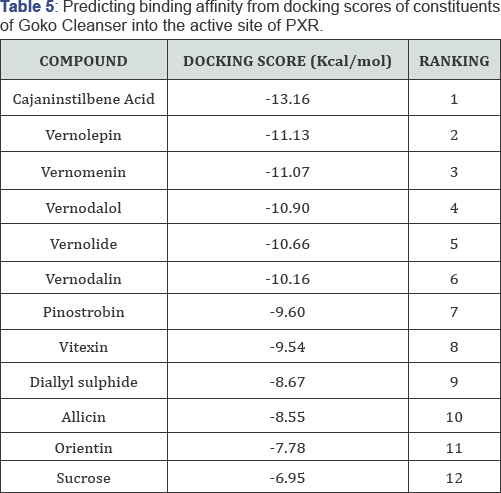
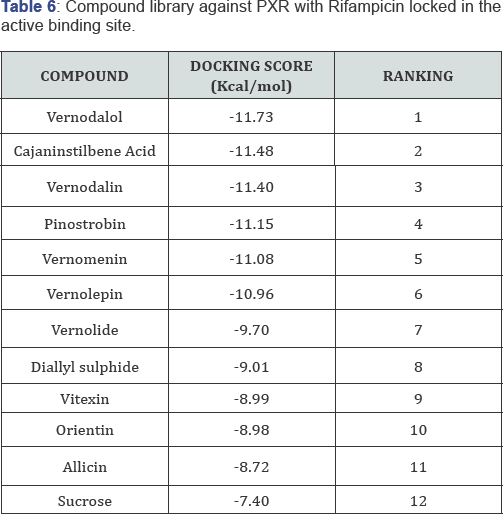
Ligand orientations are scored in relation to intermolecular energy interaction and ranked relative to other poses and ligands in the compound library. Scoring functions such as AScore used in this study provides a fast and predictive tool for the estimation of a ligand's binding affinity from its binding energy. The low energies (Table 5) observed with the binding of these constituents with the PXR active site, shows they are inducers of the receptor. Similar docking scores between the chemical constituents of vernonia amygdalina (Table 5) can be attributed to having similarities in their structures; ranking 2-6 on the table. From this results (Table 5), the goko cleanser constituents are seen to have good binding affinity for the active site of PXR, hence could explain the effect of increase ClT upon administration of goko cleanser 6 days before the administration of rifampicin to the rats. Results from Table 6, shows good binding affinities for PXR biniding site also, but due to complex formation and unfavorable bumps and hindrances, the induction effect of both goko cleanser and rifampicin are greatly affected, hence an increase in Cmax and Tmax during concurrent administration [31-35].
- Research Article
- Abstract
- Introduction
- Materials and Methods
- Methods
- Software Applications
- Library Design of Ligands
- Preparation of Molecular Target
- Docking Studies
- Binding Analysis
- Result and Analysis of Data
- Discussion
- Predicting Binding Affinity from Docking Scores
- Binding Pattern Analysis
- Conclusion
- References
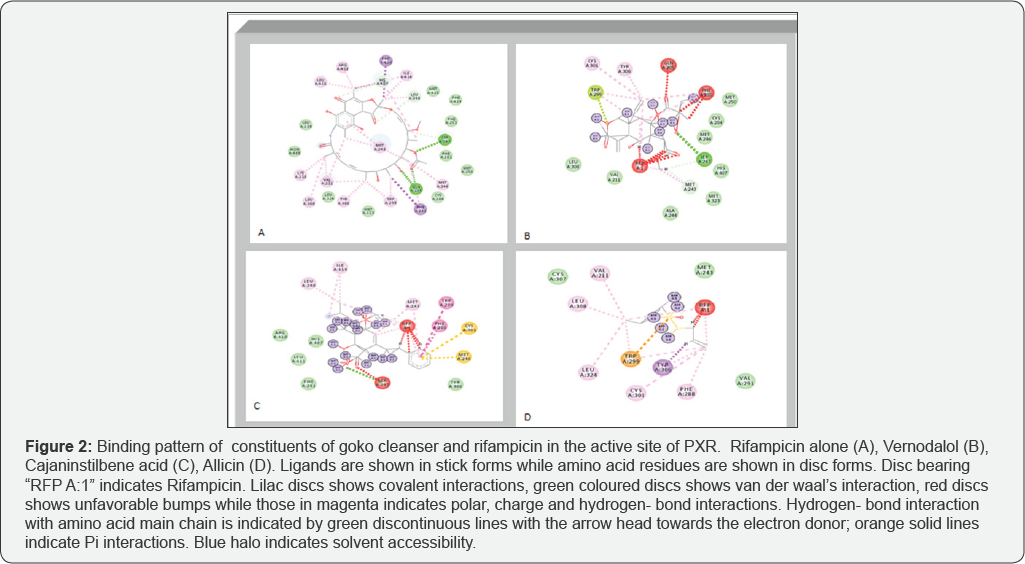
Binding Pattern Analysis
From Figure 2A, Rifampicin can be seen interacting with active site residues , while in the presense of goko cleanser constituents (Figure 2B-2D), there is a shared interaction between Rifampicin and these compounds. From this study, it was observed that interactions of the constituents of goko cleanser (Table 1) was mostly through covalent bonding. The complex of goko cleanser constituents-rifampicin that would be formed through this covalent bonding interactions can explain the decrease in ClT observed in the concurrent administration of rifampicin and Goko cleanser to the rats. Other interaction types such as carbon-hydrogen bonding and some unfavorable bumps were also observed. These interaction can in the long run affect the bioavailability of rifampicin, hence interfering with treatment outcomes [35-43].
- Research Article
- Abstract
- Introduction
- Materials and Methods
- Methods
- Software Applications
- Library Design of Ligands
- Preparation of Molecular Target
- Docking Studies
- Binding Analysis
- Result and Analysis of Data
- Discussion
- Predicting Binding Affinity from Docking Scores
- Binding Pattern Analysis
- Conclusion
- References
Conclusion
Goko Cleanser have effect on the pharmacokinetic profile of Rifampicin in male Swiss albino rats, as indicated by the altered Cmax values in phase 2 and 3. The co-administration of Goko cleanser with rifampicin decrease Tmax , Clt and Kel but increase Cmax, AUC and the T1/2β while in the delayed effect Goko cleanser lowered the C max, AUC and K el with increase in the CIt and the T1/2β, while Tmax remained unchanged as the p value was less than 0.5 for all rats.Constituents of goko cleanser were seen to be involved in covalent bonding interactions amongst other types of interactions with rifampicin in the active site of PXR. This could have resulted in the observed decrease in ClT of rifampicin when administered concurrently as seen in phase II of this study. Also since the constituents of goko cleanser showed a significant binding affinity to the active site residues of PXR, it could have induced its metabolic effect on rifampicin as seen in phase III of this study. These indicates the presence of pharmacokinetic drug interaction between both drugs and they should not be taken concurrently also one should be discontinued at least a week prior to the administration of the other. Goko cleanser enhances the activity of cytochrome P450 enzymes which (are groups of enzymes in the liver responsible for the metabolism of most drugs) leads to a decrease in the concentration and effect of an administered drug, hence possess a clinical problem when co-administered with other drugs. In general orthodox medicine and herbal preparation co-administration should be discouraged.
- Research Article
- Abstract
- Introduction
- Materials and Methods
- Methods
- Software Applications
- Library Design of Ligands
- Preparation of Molecular Target
- Docking Studies
- Binding Analysis
- Result and Analysis of Data
- Discussion
- Predicting Binding Affinity from Docking Scores
- Binding Pattern Analysis
- Conclusion
- References
References
- Amberg A (2013) In Silico Methods. In: Vogel HG, Maas J, Hock FJ, Mayer D (eds) Drug Discovery and Evaluation: Safety and Pharmacokinetic Assays. Springer, Heidelberg, Berlin.
- Ni A, Xiaohui F, Sean E (2015) In silico methods for predicting drug- drug interactions with cytochrome P-450s, transporters and beyond. Advanced Drug Delivery Reviews 86: 46-60.
- Bertilsson G, Heidrich J, Svensson K, Asman M, Jendeberg L, et al. (1998) Identification of a human nuclear receptor defines a new signaling pathway for CYP3A induction. Proc Natl Acad Sci, USA 95(21): 12208-12213.
- Lehmann JM, McKee DD, Watson MA, Willson TM, Moore JT, et al. (1998) The human orphan nuclear receptor PXR is activated by compounds that regulate CYP3A4 gene expression and cause drug interactions. J Clin Invest 102 (5): 1016-1023.
- Rofel, shri GM, Bilakhia (2015) Validated Analytical Methods developed on Anti tubercular Drug, Rifampicin. Journal of pharmaceutical Science and Bio scientific Research 5(3):254-265.
- Ebenezer O, Olatunde O (2011) Antioxidative and chemopreventive properties of (Vernoniaamygdalina and Garcinabiflavonoids). IntJ Environ Res Public health 8(6): 2533-2555.
- Amandeep S, Uma RL, Hayat MM, Prabh SS, Gagan S (2015) Phytochemical profile of sugar cane and its potential health aspects. Pharmacognosy Review Phcog Net 9(17): 45-54.
- Natalia M, Spyridon P, Isabel CFR (2016) Chemical composition and bioactive compounds of garlic (Allium sativum l) as affected by pre and post-harvest composition. Food chemistry 2(11): 41-50.
- Abdel RS, Ali OA, Haron R (2010) The chemical composition of pigeon pea (Cajanus cajan) seed and functional properties of protein isolate. Pakistan Journal of Nutrition 9(11): 1069-1073.
- Chrencik JE, Orans JO, Moore LB, Xue Y, Peng L, et al. (2005) Structural disorder in the complex of human pregnane x receptor and the macrolide antibiotic rifampicin. Mol Endocrinol 19(5): 1125-1134.
- Dassault Systemes BIOVIA (2017) Discovery Studio Modeling Environment, Release (2016) San Diego: Dassault Systemes.
- Thompson MA (2004) Molecular docking using Arguslab, an efficient shape- based search algorithm and the Ascore scoring function. ACS meeting, Pennsylvania.
- Acocella G (1978) Clinical Pharmacokinetics of rifampicin. Clinical Pharmacokinetics 3(2): 108-127.
- Acocella G, Conti R, Liesetti M, Pozzi F, Grassi C, et al. (1985) Pharmacokinetics studies on antituberculosis regimens in humans; Absorption and metabolism of the compound used in the initial intensive phase of the short course regimen: Single administration study. American Review of Respiratory Diseases 132(3): 510-515.
- Pawlowska I, Palewski J (1979) Studies on pharmacokinetics of rifampicin in the body of patients with pulmonary Tb., Arzneimittetforsehung 29:1906-1909.
- Hadijah Semvua (2013) Salivary, Protein Unbound and Total Plasma Concentration of Rifampicin. SYMPOSIUM 1:1-14.
- Boman G (1974) Serum concentration and half- life of rifampicin after simultaneous administration of aminosalicylic acid or isoniazide. European Journal of Clinical Pharmacology 7(3): 217-225.
- Boman G and Ringerber VA (1974) Binding of rifampicin by human plasma proteins. European Journal Of Clinical Pharmacology 7(5): 369-373.
- Kenny MT, and Strates B (1981) Metabolism and pharmacokinetics of the antibiotic rifampicin. Drug metabolism Reviews 12(1): 159218.
- Boehringer Ingelheim Pharmaceutical Ridgrfeild Scientific Discussion (2002) Procedures on Telmisartan documentation 2(5): 17-25.
- Bruguerolle B (1998) Pharmacokinetics: Current status (Review). Clinical pharmacokinetics 35(4): 83-94.
- Cairns D (2008) Essentials of Pharmaceutical Chemistry (3rd edn), Pharmaceutical Press, London, Chicago Pp: 159-162.
- Clinical Pharmacokinetic Studies of pharmaceuticals (2001) (5th edn).
- Craig CR, Stitzel RE, Bostan MA (1997) Modern Pharmacology with Clinical Application. Little brown and company Publication 5:557.
- Desta Z, Soukhova NV, Flockhart DA (2001) Inhibition of cytochrome P450 (CYP450) isoforms by isoniazid: potent inhibition of CYP2C19 and CYP3A4. J Chromatograph B Biomed Sci Appl 45(2): 382-392.
- Esteban Riberia, Carlos A, Rosa ML (2007) Pharmacokinetics interaction between Rifampicin and the once daily combination of saquinavir and low dose ritonavir in HIV infected patients with tuberculosis. J Antimicrob Chemother 59(4): 690-697.
- Fugh Berman A, Ernst E (2001). Herb-drug Interactions: Review and Assessment of Report Reliability. Bri J of Clinical Pharmacol 52(5): 587-595.
- Fugh Berman A. (2003) Herb-drug interactions. The Lancet 355(9198): 134-138.
- General Guideline for Clinical Evaluation.
- Green world products guide and catalogue (2010) Overview and corporate profile. Tianji, China p: 1-32.
- Guidance on the Ethnic Factors to Accept Clinical Data Obtained in a Foreign Country.
- Guideline for Non-clinical Pharmacokinetic Studies.
- ICH Topic Q2 (R1) (2005) Validation of Analytical Procedures: Text and Methodology accessed.
- Iyakushin (1997) Guideline for Bioequivalence Studies of generics pp: 487.
- Jean Pierre, Labaune (1989) Handbook of Pharmacokinetics toxicity assessment of chemicals, Ellis Horwood Limited Chickester, West Sussex, Englan pp: 206-535.
- Katzung BG, Masters SB, Trevor AJ (2007) Basic Clinical Pharmacology, McGraw Hill Publicatin, North America 10: 771-772.
- Kolawole JA (2012) Tollgates to effectiveness and safety of medicines in drug therapy. Uni Jos inaugural lecture series 53:13-29.
- Krishna K Rao (2015) Clinical drug Interactions. JIMSA 1(28): 42-44
- Leccillon JB, Febvre N, Metayer JP, Souppart C (1978) Quantitative assay of rifampicin and three of its metabolites in human plasma. urine and saliva by HPLC: Journal of chromatography; 145(2): 319-324.
- (2001) Official Texts on Conducting Drug Interaction Studies Promulgated in Japanese.
- Rang HP, Dale MM, Ritter JM, Flower RJ, Henderson G (2006) Pharmacology. Elsevier publication, India 5: 649-652.
- Sara Bogenschutz, Angela Bojrab (2003) Biology of Ageing Fall. Drug Interactions (1): 3-11.
- Scott R. Penzak (2007) Clinical Pharmacokinetics Research laboratory. Drug Interaction (1): 16-20.






























