Bacteria Isolated from Hospital Wastes in Owerri Municipality: Antibiotic Susceptibility and Resistance Profile
Ohalete CN1*, Uwaezuoke JC1, Emeh AA2 and Dunga KE3
1Department of Microbiology, Imo State University, Owerri, Nigeria
2Department of Biology & Microbiology, Federal Polytechnic Nekede, Owerri, Nigeria
3Department of Medical Laboratory Science Rhema University, Aba
Submission: September 12, 2023; Published: October 17, 2023
*Corresponding author: Ohalete CN, Department of Microbiology, Imo State University, Owerri, Nigeria
How to cite this article:Ohalete CN, Uwaezuoke JC, Emeh AA, Dunga KE. Bacteria Isolated from Hospital Wastes in Owerri Municipality: Antibiotic Susceptibility and Resistance Profile. JOJ Pub Health. 2023; 8(2): 555734. DOI: 10.19080/JOJPH.2023.08.555734
Abstract
Hospital waste samples from the Federal Medical Center, Anagbaoso Hospital, and Amanda Hospital in Owerri, Nigeria were haphazardly collected from April to July 2020. These wastes included hand gloves, urine infusion bags, blood infusion bags, bandages, and syringes. Standard procedures were used for the bacteriological and antibiotic susceptibility testing. The spread plate method of aerobic bacterial count revealed bacterial loads ranging from 4.22 x 104 to 5.54 x 106 cfu/ml. The isolates that were found were Salmonella typhi, Staphylococcus aureus, Escherichia coli, Klebsiella pneumoniae, and Pseudomonas aeruginosa.
The antibiotic susceptibility test, which employed Levofloxacin, Erythromycin, Streptomycin, Chloramphenicol, Ceporex, Gentamycin, Tarivid, Augmentin, Reflacine, Ciproflox, and Ampicillin, revealed zones of inhibition ranging from 6 mm to 25 mm. The isolates’ antibiotic sensitivity percentages ranged from 36.36% to 63.64%. Staphylococcus aureus and Salmonella typhi from the three hospitals had the lowest percentage sensitivity to the medicines, whereas Pseudomonas aeruginosa isolated from Amanda Hospital had the highest percentage sensitivity (63.64%). All of the isolates had Ampicillin resistance and had decreased sensitivity to Gentamycin and Erythromycin. To lessen exposure to dangerous hospital acquired infections, better hospital waste management is required.
2. Keywords:Hospital wastes; Isolates; Antibiotics Susceptibility; Resistance
Introduction
Regardless of ethnicity, sex, age, or religion, the hospital is a crucial location for human life, health, and wellbeing. But due to its high risk of disease transmission, the waste produced by hospitals and other medical facilities can be dangerous, poisonous, and even fatal. If not properly handled, disposed of, or permitted to mix with other municipal waste, the hazardous and toxic components of waste from healthcare facilities, which includes infectious, biomedical, and radioactive material as well as sharp objects (hypodermic needles, knives, scalpels, etc.), pose a serious risk. As a result, it promotes the development of numerous infections and vectors, contaminates other municipal waste that isn’t poisonous or dangerous, and jeopardizes attempts to manage municipal waste as a whole [1,2] Hospital solid waste (SW) can have extremely high levels of disease-causing pathogenic organisms. (Madhya Pradesh Pollution Control Board [3,4] When trying to rescue products for sale or reuse, waste pickers and employees are frequently the ones that suffer the most. They sift through a variety of dangerous materials.
Unbeknownst to them, such unethical and unlawful reuse poses serious public health concerns and can lead to epidemic or even endemic forms of diseases like cholera, typhoid, diarrhea, dysentery, and skin conditions. Making reference to the research carried out in Kenya [4] on medical waste and health care settings in hospitals, the bacteria isolated from their study included; Providentia rettgeri, Staphylococcus aureus, Escherichia coli, Pseudomonas aeruginosa, Coagulase negative Staphylococcus, Serratia marcescens, Klebsiella pneumoniae, Proteus vulgaris and Enterobacter cloaca. However, in their investigation of biomedical waste [5] recovered B. subtilis, S. aureus, K. pneumoniae, and E. coli.
Traditional cleaning techniques, such as dry cleaning and dusting, can distribute dust particles, encouraging contamination across the space surrounding the patient and adversely affecting the air quality in medical and healthcare facilities. Traditional guidelines are insufficient to safeguard people at risk for serious and fatal illnesses [6,7] Inadequate treatment of these wastes exacerbates this effect. Adoption of new technology was advised to augment traditional approaches in order to improve desirable levels of surface disinfection since the appearance and spread of multi-drug resistant germs in hospital wards, which have been on the rise, represent a severe threat [8] Infectious hospital waste may contain germs that can spread to hospital patients, healthcare professionals, and members of the general public by a variety of methods, including punctures, abrasions, or lacerations of the skin, via the mucous membranes, by inhalation, or by ingestion. To safeguard both humans and the environment, hazardous waste needs to be appropriately packaged, transported, and disposed of.
It is now a major public health concern because pathogenic organisms are becoming increasingly resistant to antibiotics. Through intentional or spontaneous genetic mutation, or from other bacterial species via horizontal gene transfer, drug-resistant organisms may have developed resistance to first-line antibiotics, second-line antibiotics, third-line antibiotics, and even broad range antibiotics. The majority of the time, antibiotic resistance genes are found in intransmissible plasmids, which promotes their transfer and allows them to proliferate throughout a bacterial community and confer resistance to a variety of antibiotics [9]. The misuse and overuse of antibiotics is a major factor in the creation of drugresistant organisms. Increased antibiotic resistance is also a result of the practice of adding antibiotics to livestock feeds to promote faster growth and any unethical behavior in the pharmaceutical manufacturing sector [10,11]. Therefore, the goal of this study is to identify the patterns of isolates’ antibiotic susceptibility from hospital wastes in Owerri, Nigeria.
Method and Materials
Collection of Samples
In Owerri, Nigeria, the Federal Medical Center, Anagbaoso Hospital, and Amanda Hospital provided samples of hospital wastes that were collected in sterile polythene bags. Bandages, syringes, blood infusion bags, urine infusion bags, and hand gloves were among the samples gathered using the random sampling technique. These samples were gathered between the months of April and July 2020 and transported to the Imo State University’s Microbiology Laboratory in Owerri for analysis.
Processing and bacterial analysis of hospital waste samples
The technique as described by [5] and used on the hospital waste samples. In 99ml of sterile, distilled water, 1gm of each type of sample was suspended, and the mixture was violently agitated for 2 minutes. The varied suspensions were then taken out of the samples and put into sterile conical flasks for collection. The study’s media, including MacConkey agar, Nutrient agar, Blood agar, and Salmonella-Shigella agar (SSA), were produced in accordance with the instructions provided by the manufacturers. The produced samples underwent bacterial examination in accordance with accepted textual descriptions [12-14] The produced suspensions were serially diluted by one milliliter (1.0 ml), and a few of the dilutions were streaked onto Blood agar, MacConkey agar, and Salmonella-Shigella agar before being incubated at 37°C for 24 hours. To obtain a pure culture of each potential isolate for biochemical identification, the colonies were subsequently subcultured.
The spread plate method was used to do the aerobic plate counts of the waste samples in accordance with accepted practices. To obtain dilutions between 10-1 and 10-5, each sample suspension was serially diluted in 1.0 ml (0.1% buffer peptone water). The surface of prepared sterile nutritional agar plates, Blood agar plates, MacConkey agar plates, and Salmonella- Shigella agar were then covered with 0.1 ml of the diluted sample (10-1 - 10-5) using a sterile spreader. The plates were sprayed with an inoculant in triplicate and incubated for 24 hours at 37°C. Following incubation, colonies on plates ranging in size from 30 to 300 were counted using a colony counter, and the average value obtained was recorded as cfu/ml.
Identification of organisms
As indicated in standard texts, the various bacteria isolates were identified by their cal, morphological, and biochemical features (gram staining, spore staining, motility test, catalase test, citrate utilization test, indole test, oxidase test, sugar fermentation test) [15,16].
Antibiotic Sensitivity Test
According to the standards of the Clinical and Laboratory Standard Institute, the antibiotic resistance of the isolates to certain conventional antibiotics was tested in triplicate on Mueller Hinton agar plates using the standard Kirby- Bauer disk diffusion technique [17]. Levofloxacin (20 mcg), Erythromycin (30 mcg), Streptomycin (30 mcg), Chloramphenicol (30 mcg), Ceporex (10 mcg), Gentamycin (10 mcg), Tarivid (10 mcg), Augmentin (30 mcg), Reflacin (10 mcg), Ciproflox (10 mcg), and Ampicillin were the antibiotics employed against the isolates (30mcg). The test bacteria was suspended in sterile, universal vials containing nutrient broth for a 24-hour culture period. In order to compare with 0.5 MacFariane standards (108 cells/ml), normal saline was gradually added. After thoroughly mixing the inoculums, any surplus fluid was removed by dipping a clean, sterile cotton tipped swab into the inoculums. The surface of the solidified Mueller Hinton agar was equally covered with the swab. Agar plates with seeds already on them were gently and aseptically covered with antibiotic discs, which were then incubated at 37°C for 24 hours. Measurements and comparisons with the NCCLS break point were made between the zones of inhibition that appeared in the triplicate plates [18].
Statistical Analysis
The analysis of variance (ANOVA) was used to statistically examine all the data collected from the various parameters, and the least significant difference (LSD) test was employed to compare the means (Steel et al., 1997).
Results
Table 1 displayed the results of the aerobic bacterial loads from the hospital wastes that were collected, which ranged from 4.22 x 104 cfu/ml (waste syringes from Federal Medical Center - Hospital A) to 5.54 x 106 cfu/ml. (blood infusion bags from Amanda hospital - Hospital C). The urine infusion bag from Amanda Hospital had an aerobic bacterial load of 3.4x105, which was higher than that of the urine infusion bag samples from the other two hospitals. Bacterial load on the hand gloves ranged from 4.22x104 to 4.78x104. Bandages ranged in size from 5.68x104 (Amanda Hospital - Hospital C) to 3.04x105 (Anagbaoso Hospital - Hospital B) Table 1.
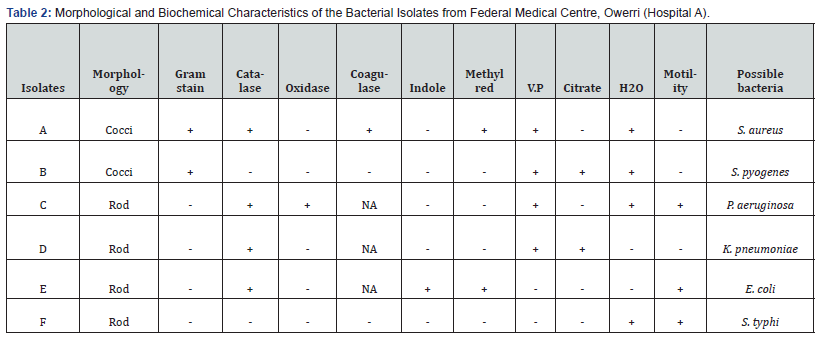
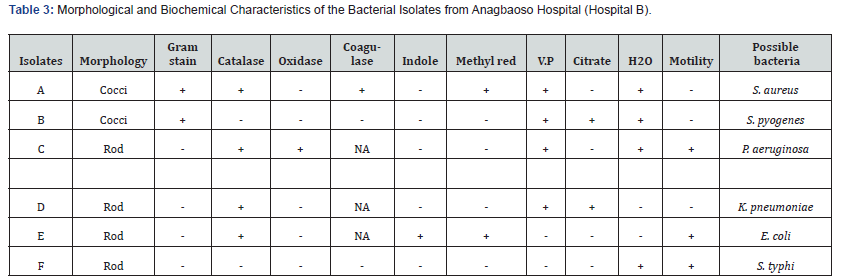
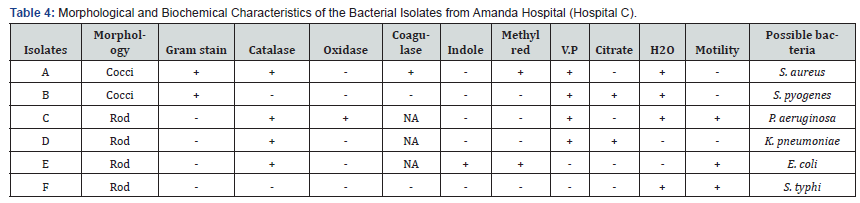
The results of the morphology and biochemical characteristics of the bacterial isolates from the three different hospital wastes were provided in detail in Tables 2-4. The tables showed that the six bacteria isolated from the wastes of the three hospitals under study were identical and included Pseudomonas aeruginosa, Staphylococcus aureus, Escherichia coli, Streptococcus pyogenes, Klebsiella pneumoniae, and Salmonella typhi. The isolated patterns of antibiotic sensitivity to various antibiotics were shown in Tables 5-7. While Tables 8 & 9 displayed the sensitivity and resistance percentages of each isolate from the three hospitals. According to the tables, Staphylococcus aureus and Salmonella typhi had the lowest percentage sensitivity (36.36%) from the three hospitals, while Pseudomonas aeruginosa isolated from Amanda hospital wastes had the highest percentage sensitivity (63.64%) Additionally, Staphylococcus aureus and Salmonella typhi had the highest percentage resistance (63.64%), while Pseudomonas aeruginosa isolates from Amanda hospital had the lowest percentage resistance (36.36%).

Discussion
Hospital trash is generated both by the patients themselves and by the staff as they provide services to the patients. These wastes may be generated directly in conjunction with an upstream, downstream, or service process (such as blood or urine culture in a laboratory). (e.g. injection). According to the hospital involved and the categories of hospital waste, there were different levels of contamination, according to the aerobic bacterial loads from the hospital wastes collected for this study. The Federal Medical Center’s syringes had the lowest bacterial load, 4.22 x 104 cfu/ml, while Amanda Hospital’s blood infusion bag samples had the highest aerobic bacterial load, 5.54 x 106 cfu/ml. Compared to other blood infusion bag samples from FMC and Anagbaoso hospitals, the Amanda hospital’s (Hospital C) bags showed a higher load. Blood is a fertile environment for bacterial development, and any organisms that come into touch with it have a propensity to reproduce luxuriantly. In a similar vein, pee infusion bag samples from Amanda hospital showed the greatest aerobic bacterial load, with 3.4x105, compared to urine infusion bag samples from the other two hospitals Table 1. Inadequate hospital waste management practices may be to blame for the elevated bacterial loads.
In the current study, the findings from the bacteriological analysis of wastes in the three chosen hospitals in Owerri Tables 2-4 showed that the isolates (bacteria), which included P. aeruginosa, S. aureus, E. coli, S. pyogenes, K. pneumoniae, and S. typhi, were the same in the three hospitals. These six bacteria—P. aeruginosa, S. aureus, E. coli, S. pyogenes, K. pneumoniae, and S. typhi—were examined as part of this investigation and are significant germs frequently linked to numerous nosocomial infections Some of the microorganisms that were isolated for this investigation came from the hospital waste studies done by [4]. They reported that different percentages of Staphylococcus aureus, Escherichia coli, Pseudomonas aeruginosa, Klebsiella pneumoniae, and other gram negatives were isolated. In their investigation of biomedical waste, [5] also isolated S. aureus, K. pneumoniae, and E. coli.
While some of these isolates were sensitive to some of the antibiotics, others were resistant to some of the medicines (Table 5-7). The dumpsite area could be held responsible for the growth and spread of germs that are easily transported by air. Furthermore, given that the dumpsites are adjacent to hospitals and may have an impact on persons who visit or work there, the environment there may have had a direct or indirect impact on the isolates. Increased illness severity or an increase in infections picked up in hospitals could result from this.
According to [19] the widespread use of antibiotics gives bacteria a selection advantage, and it is conceivable for resistant strains to spread from patient to patient within hospitals. P. aeruginosa, S. aureus, E. coli, S. pyogens, K. pneumoniae, and S. typhi, six bacteria explored in this investigation, are significant microorganisms frequently associated with several nosocomial diseases [20]. Because both patients and hospital employees are exposed to these bacteria, the presence of resistant strains of these six pathogens in hospitals and the environment should be of great concern. These microorganisms are easily transported by foot or other methods into the hospital environment, raising the possibility of infection. Additionally, they may be washed into the hospitals’ subsurface water supply by rain, contaminating it and spreading infections that are transmitted by water [21]. It is important for public health that these bacteria isolates were found in the waste samples from the three hospitals that were chosen. Since the majority of these organisms are linked to a number of illness problems, they could seriously endanger both the public and hospital patients’ health [3].
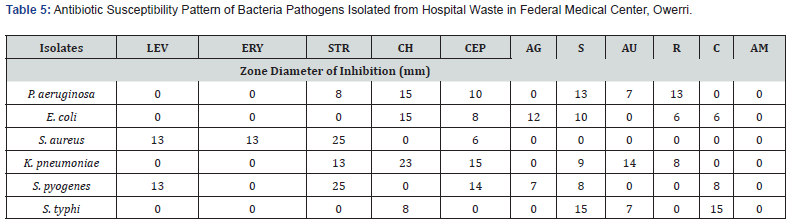
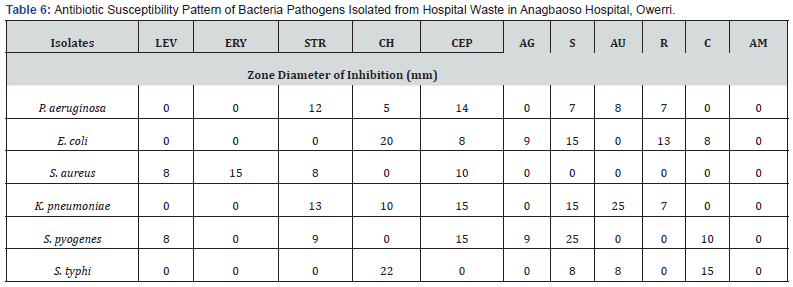
As shown in Tables 5-7 the antibiotic susceptibility pattern used in this study revealed variable degrees of zones of inhibition on the hospital isolates. The zone diameter of inhibition in the case of FMC Owerri ranged from 6 to 25mm Table 5.
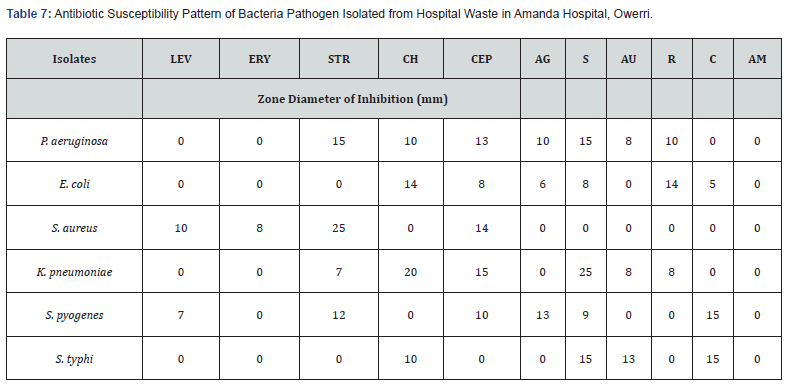
Streptomycin was found to have a zone of inhibition of 25 mm against S. aureus and S. pyogenes, which were the most susceptible bacteria to the antibacterial agent, while zones of inhibition of 13 mm and 8 mm were recorded for K. pneumoniae and P. aeruginosa, respectively. E. coli and S. typhi were resistant to streptomycin. The second most active agent, chloramphenicol, had zones of inhibition of 23The antibiotics Ceporex (14mm), Streptomycin (12mm), Augmentin (8mm), Gentamycin (8mm), Tarivid and Reflacine (7mm), and Chloramphenicol (5mm) zones of inhibition were effective against P. aeruginosa in the Anagbaoso hospital Table 6 but Levofloxacin, Erythromycin, Ciproflox, and Ampicillin were resistant. Additionally, Streptomycin and Tarivid had the maximum zones of inhibition at 25 mm against S. aureus and K. pneumoniae in Amanda Hospital Table 7.
Chloramphenicol had zones of inhibition at 20 mm against K. pneumoniae, 14 mm against E. coli, and 10 mm against P. aeruginosa and S. typhi. Additionally, Tarivid showed zones of inhibition of 15 mm against P. aeruginosa, 15 mm against S. typhi, and 9 mm and 8 mm respectively against S. pyogenes and E. coli. Additionally, Ceporex demonstrated zones of inhibition against K. pneumoniae, S. aureus, P. aeruginosa, S. pyogenes, and E. coli of 15 mm, 14 mm, 13 mm, 10 mm, and 8 mm, respectively Table 7. It is interesting to notice that every isolate of bacteria from the three hospital wastes used in this investigation was ampicillin resistant.
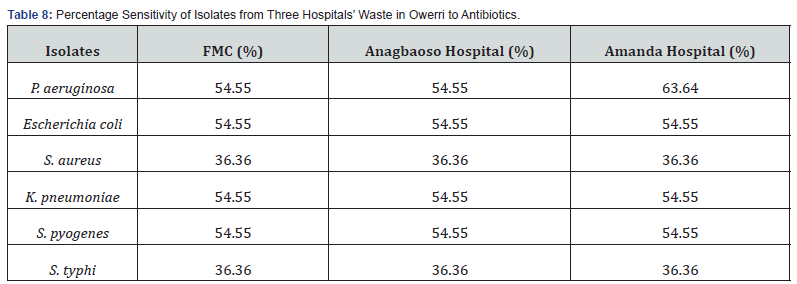
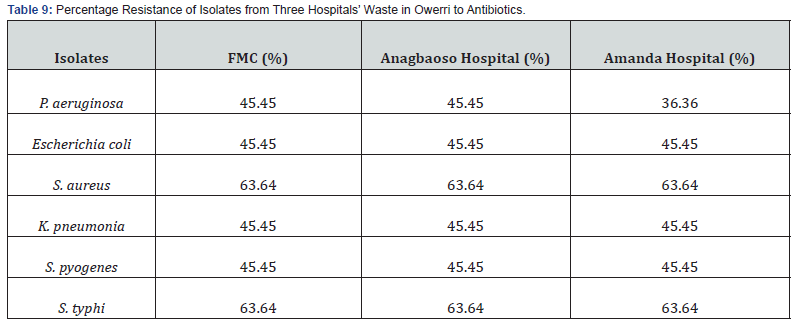
Table 8 further revealed that the isolates’ percentage sensitivity ranged from 36.36% to 63.64%. The proportion of sensitivity for Pseudomonas aeruginosa isolated from Amanda Hospital was 63.64%, whereas the percentages for Staphylococcus aureus and Salmonella typhi from the three hospitals were 36.36% and 36.36%, respectively. In contrast, as shown in Table 9, Staphylococcus aureus and Salmonella typhi had the highest percentage resistance, with Pseudomonas aeruginosa in Amanda Hospital having the lowest percentage resistance of 36.36% and 63.64%, respectively.
Inadequate or excessive use of the antibiotics by patients who purchase their medication from local pharmacy shops may be the cause of some hospital waste isolates’ resistance to some of the antibiotics utilized in this investigation. (popularly called chemists). It was found by [22] that, despite the high rate of non-compliance, the number of people who received antibiotic prescriptions that were not necessary is a bigger contributor to the rise in bacterial resistance than non-compliance among those who were given the drugs. People insisting on specific antibiotics, doctors prescribing antibiotics because they feel they don’t have time to explain why such drugs are not necessary, or doctors being overly cautious for legal or medical reasons are a few examples of factors that have been linked to inappropriate antibiotic prescribing [23]. The likelihood of developing strains resistant to antibiotics can also be increased by unethical pharmaceutical production practices. It encourages the transfer and dissemination of antibiotic resistance genes within an ecosystem of bacteria in scenarios where they are likely to be present in transmissible plasmids, resulting in the development of resistance to a number of antibiotics [7]. If an organism has plasmid and that organism is resistant to an antibiotic, that antibiotic may not be able to treat the infection, especially if that antibiotic is the medicine of choice. This is very concerning since managing and treating infections brought on by this plasmid-carrying species could result in therapeutic failure.
A similar study by [24] concluded that the demand for alternate treatments increases as antibiotic resistance spreads. Salmonella and other human harmful bacteria have been proven to be treatable with bacteriophages rather than antibiotics, and they may even be able to be controlled by them [25]. Additionally, it has been demonstrated that using medicinal herbs is an excellent substitute for applications of antibiotics [26]
Conclusion
The study’s high microbial load densities, pathogenic isolates, and some resistant strains indicate that hospital wastes in the environment pose a serious hazard to human health and the ecosystem. In order to prevent nosocomial infections and potential epidemics, it is necessary to control infection within the hospital. This necessitates the creation of an appropriate legislative framework for the disposal of hospital wastes as well as a pressing need for management strategies, education campaigns, and increased public knowledge of medical waste issues. as a result, the following suggestions: The government should design, plan, implement as well as insist on policies and regulations for proper waste management to minimize risks relevant to human health and environment.
Proper financial support should also be available to the health sector so that they can comply with those rules and regulations. Each hospital should constitute a hospital waste management committee having wide representation from all major departments, responsible for making specific action plan for hospital waste management and its supervision, monitoring and implementation. Adoption of modern technologies should be encouraged to supplement traditional methods in order to achieve desired goals.
References
- WHO (2018) Health-care waste. World Health Organization.
- Shareefdeen ZM (2012) Medical waste management and control. Journal of Environmental Protection 3(12): 1625-1628.
- Nworie A, Ayeni JA, Eze UA, Azi SO (2012) Bacterial contamination of door handles/knobs in selected public conveniences I Abuja metropolis, Nigeria: A public health threat. Continental Journal of Medical Research 6(1): 7-11.
- Maina SM, Nyerere AK, Ngugu CW (2018) Isolation of bacterial diversity present in medical waste and health care settings in hospitals in Kenya. African Journal of Microbiology Research 12(26) :606-615.
- Anitha J, Indira JA (2012) Isolation and identification of bacteria from biomedical waste (BMW). International Journal of Pharmacy and Pharmaceutical Sciences 4(5): 386-388.
- Al Hamad A, Maxwell S (2008) How clean is clean? Proposed methods for hospital cleaning assessment. Journal of Hospital Infection 70(4): 328–334.
- WHO (2019) Wash in health care facilities: global baseline report 2019. World Health Organization.
- Louw GE, Warren RM, Gey Van Pittius NC, McEnvoy CR, Van Helden PD, et al. (2009) A Balancing Act: Efflux/Influx in Mycobacteri Drug Resistance. Antimicrobial Agents and Chemotherapy 53(8): 3181-3189.
- Wright JD (2010) Antibiotic resistance in the environment: A link to the clinic? Current Opinion in Microbiology 13(5): 589-594.
- Matthew AG, Cissel R, Liamthong S, Cissel R (2007) Antibiotic Resistance in bacteria associated with food animals: A United States perspective of livestock production. Foodborne Pathogenic Diseases 4(2): 115-133.
- Larsson DG, Fick J (2009) Transparency throughout the production chain: A way to reduce pollution from the manufacturing of pharmaceuticals?. Regul Toxicol Pharmacol 53(3): 161-163.
- Baron EJ, Peterson LR, Finegold SM (1994) Bailey and Scott’s Diagnostic Microbiology 9th Edt Pp: 177-180 St.
- Fawole MO, Oso BA (2004) Laboratory Manual of Microbiology 2nd Edt Spectrum Books Limited, Lagos.
- Cheesbrough M (2006) District Laboratory Practice in Tropical Countries Part 2. 15 McNulty CA, Boyle P, Nichols T, Clappison P, Davey P (2007) The public’s attitudes to and compliance with antibiotics. Journal of Antimicrobial Chemotherapy 60(1): 163-168.
- Li JZ, Winston LG, Moore DH, Bent S (2007) Efficacy of short-course antibiotic regimens for community-acquired pneumonia: A meta-analysis. Amplified Journal of Medicine 120(9): 783-790.
- Onyeleke SB, Istifanus N (2009) The microbiological effects of hospital wastes on the environment. African Journal of Biotechnology 8(22): 6253-6257.
- Bauer AW, Kirby WM, Sherris JC (1996) Antibiotic susceptibility testing by standardized single disk method. American Journal of Pathology 45: 493-496.
- CLSI (2006) Methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically: Approved standard – 7th CLSI document M7 – A7 (ISBN 1 – 56238-587-9). Clinical and Laboratory Standard Institute, 940 West Valley Road, Suite 1400, Wayne, Pennysylvania 19087- 1898 USA 26(2).
- Archibald LK (2004) Gram-negative hospital acquired infections: A growing problem. Infection Control and Hospital Epidemiology 25(10): 809-811.
- Moccia G, Motta O, Pironti C, Proto A, Capunzo M, et al. (2020) An alternative approach for the decontamination of hospital settings. Journal of Infect Public Health 13(12): 2038-2044.
- Ejiogu CC, Okpara CB, Opara KD, Onyeocha IO, Azubuike C, et al. (2018) Effects of solid waste disposal on the physicochemical properties of borehole water along FUTO road, Eziobodo, Owerri-West, Nigeria. Current Journal of Applied Science and Technology 26(1): 1-7.
- Pechère JC, Hughes D, Kardas P, Cornaglia G (2007) Non-compliance with antibiotic therapy for acute community infections: A global survey. International Journal of Antimicrobial Agents 29(3): 245-253.
- Ariffin H, Navaratnam T, Kee KT, Balan G (2004) Antibiotic resistance patterns in nosocomial Gram-negative bacterial infections in units with heavy antibiotics usage. Journal of Tropical Pediatrics 54(1): 26-31.
- Donadio S, Maffioli S, Monciardini P, Sosio M, Jabes D (2010) Antibiotic discovery in the twenty-first century: current trends and future perspectives. Journal of Antibiotics (Tokyo) 63(8): 423-430.
- Fong K, Mu K, Rheaut JG, Goodridge L, Wang S (2020) Bacteriophage- Insensitive mutants of antimicrobial-resistant Salmonella enteric are altered in their Tetracycline resistance and virulence in Caco-2 intestinal cells. International Journal of Molecular Science 21(5): 1883.
- Emeh AA, Anyanwu GO, Onyeulor PN, Chimereze NC, Abba Father CAM (2020) Antimicrobial and synergistic potentials of Xylopia aethiopica (Uda) and Occimum gratissimum (Nchanwu) leaf extracts. International Journal of Innovative and Research Technology 5(4): 1179-1185.






























