regulaAnalysis of Codon Usage Bias in Atractylodes Chinensis (DC.) Koidz. Based on Transcriptome Data
Yang Lu1, Qingxiao Wang2 and Chunying Zhao1*
1Institute of Traditional Chinese Medicine, Chengde Medical University, Hebei
2Department of Biomedical Engineering, Chengde Medical University, Hebei
Submission:June 28, 2025;Published: July 08, 2025
*Corresponding author: Chunying Zhao, Institute of Traditional Chinese Medicine, Chengde Medical University, Hebei, 067000, China, Email: luyangmm@cdmu.edu.cn
How to cite this article: Yang Lu, Qingxiao Wang and Chunying Zhao. regulaAnalysis of Codon Usage Bias in Atractylodes Chinensis (DC.) Koidz. Based on Transcriptome Data. Juniper Online Journal of Public Health, 5(4). 555670.DOI: 10.19080/JOJHA.2025.05.555670.
Abstract
To investigate the codon bias characteristics and influencing factors of Atractylodes chinensis (DC.) Koidz. based on the transcriptome, a total of 40,309 coding sequences from the transcriptome data were selected as the research objects. Programs and software such as Codon 1.4.2, R language, and EMBOSS were used to study the codon usage bias of the A. chinensis transcriptome. The results showed that the GC content of codons in the CDS sequences of the A. chinensis transcriptome was concentrated between 35% and 50%, with an average GC content of 44.6% and a GC3 content of 43.26%. The results of neutral mapping, bias, and ENC-Plot analyses indicated that selective pressure was the main driving force for the formation of codon bias in the A. chinensis transcriptome sequences. Based on RSCU analysis, a total of 28 codons were high-frequency codons, most of which had A or U as the terminal base. Finally, GCA, GAA, GGA, and GUG were determined as the optimal codons.
Keywords:Atractylodes chinensis (DC.) Koidz; Transcriptome; Codon; Usage Bias; Selective pressure
Abbreviations:Met: Methionine, Trp: Tryptophan, CUB: Codon Usage Bias, CDS: Coding Sequence, ENC: Effective Number of Codons, CAI: Codon Adaptation Index, RSCU: Relative Synonymous Codon Usage, A: adenine, U: uracil, G: guanine, C: cytosine, T: thymine
Introduction
Atractylodes chinensis (DC.) Koidz. belongs to the Atractylodes genus within the Asteraceae family and is the original plant source of Atractylodis Rhizoma. As a commonly used bulk medicinal herb, it was first documented in the Shennong Ben Cao Jing [1]. Widely distributed in northern China, including Hebei, Inner Mongolia, and Liaoning provinces, Hebei serves as the primary production region and is recognized as one of the “Top Ten Hebei Medicines”. According to relevant studies [2], the core planting areas in Hebei are concentrated in the counties of Qinhuangdao and Chengde, where the herb yields the highest quality. The dried rhizome of A. chinensis, the main medicinal part, holds significant pharmaceutical value in traditional Chinese medicine, exhibiting effects of dispelling dampness, invigorating the spleen, relieving wind-cold, and improving eyesight [3,4]. Thriving in wild, cool, and humid environments, A. chinensis has seen increasing market demand due to its recognized potential in dietary therapy alongside social development and medical advancements. However, wild resources have become scarce due to long-term exploitation and ecological changes. Francis Crick’s central dogma illuminates the flow of genetic information from DNA to RNA to proteins, with codons play ing a pivotal role in this process [5,6]. As the bridge connecting DNA and proteins, codons are fundamental to gene transcription and translation [7,8]. In mRNA coding sequences, three nucleotides (a codon) specify a single amino acid [9].
Among the 20 basic amino acids, only methionine (Met) and tryptophan (Trp) are encoded by a single codon, while the remaining 18 amino acids are specified by 2 to 6 codons, illustrating the degeneracy of genetic codons [10]. Studies define multiple codons encoding the same amino acid as synonymous codons [11,12]. In organisms, the usage frequency of synonymous codons varies significantly, a phenomenon known as codon usage bias (CUB) [13,14]. The most frequently used codon among synonyms is termed the optimal codon [15], selected during gene expression to ensure encoding accuracy and efficiency. CUB is a universal phenomenon in nature, observed across diverse genomes. Due to codon wobble, bias typically manifests at the third codon position. Analyzing plant CUB helps characterize species-specific gene sequences and provides insights into evolutionary patterns and gene expression regulation [14]. Since the 1960s, CUB has drawn extensive attention as a key mechanism of gene expression regulation. Advances in bioinformatics have driven methodological innovations and systematic summaries of CUB across species, deepening our understanding of its biological significance from multiple perspectives. Current research on A. chinensis primarily focuses on pharmacology and active components, with limited studies on its codon usage. This study aims to reveal CUB patterns, explore influencing factors, and identify optimal codons in the A. chinensis transcriptome, providing a basis for species improvement and therapeutic applications.
Materials and Data
The test samples were collected from the Medicinal Botanical Garden of Chengde Medical University in Chengde City, Hebei Province. They were identified as Atractylodes chinensis (DC.) Koidz. by Professor Chunying Zhao from the Institute of Traditional Chinese Medicine, Chengde Medical University, and exhibited different plant phenotypes. Transcriptome sequencing was performed by Major bio (Shanghai, China). For the assembled Unigene sequences, coding sequence (CDS) analysis was conducted. Using Perl programs, 40,309 CDS sequences were screened with gene lengths exceeding 300 bp and redundant sequences removed. These sequences used ATG as the start codon and TAA, TAG, or TGA as the stop codon.
Methods
Base Content Analysis of Coding Sequences
CodonW1.4.2 software was used to analyze the effective number of codons (ENC), codon adaptation index (CAI), relative synonymous codon usage (RSCU), and optimal codon usage frequency in the CDS sequences of A. chinensis transcriptome. The GC content was determined using the cusp program of EMBOSS. Here, GC1, GC2, and GC3 denote the GC contents of the first, second, and third codon positions, respectively, while GC represents the mean of GC1, GC2, and GC3. Additionally, GC3s indicates the GC content at the third position of synonymous codons.
Neutrality Plot Analysis
Neutrality plot analysis was performed with GC3 as the abscissa and GC12 (the average of GC1 and GC2) as the ordinate, followed by linear regression analysis to examine the correlation between GC12 and GC3 for identifying the main factors influencing codon usage bias [16]. A regression coefficient close to 1 indicates a significant correlation between GC12 and GC3, suggesting mutational pressure dominance, whereas a coefficient close to 0 implies weak correlation and selective pressure dominance.
PR2-Plot Analysis (Parity Rule 2 Plot)
PR2-Plot analysis evaluated mutational equilibrium at the third codon position by plotting G3/(G3+C3) against A3/(A3+T3) with the central coordinate (0.5, 0.5). This analyzed the relationships among adenine (A), guanine (G), cytosine (C), and uracil (U) at the third position to infer factors affecting codon bias in A. chinensis [17]. When A=T and C=G, codon usage is unaffected by selection or mutation.
ENC-Plot Analysis
Using R language, a two-dimensional scatter plot was generated with GC3 values and effective number of codons (ENC) of A. chinensis CDS sequences as the x- and y-axes, respectively, along with a standard curve [18]. Genes close to the standard curve suggest mutational pressure-driven codon bias, while those far from it indicate regulation by natural selection. The standard curve formula is:
ENC = 2 + GC3 + 29/[GC3² + (1-GC3)²].
Codon Adaptation Index (CAI) Analysis
CAI, a key tool for evaluating gene expression efficiency, ranges from 0 to 1. Higher CAI values (approaching 1) indicate higher expression levels, whereas lower values reflect reduced expression [5].
Relative Synonymous Codon Usage (RSCU) Analysis
RSCU measures codon usage frequency by calculating the relative probability of a specific codon among synonymous codons encoding the same amino acid. RSCU values typically range from 0 to 2: RSCU > 1 indicates a high-frequency codon with strong bias; RSCU = 1 denotes no bias; RSCU < 1 signifies low frequency and weak bias [19].
Optimal Codon Analysis
CDS sequences were sorted by ENC values to establish highand low-expression gene libraries. RSCU values were calculated for each library, and codons with ΔRSCU (RSCU difference between libraries) > 0.08 were screened. Codons meeting both ΔRSCU > 0.08 and RSCU > 1 were identified as optimal codons in the A. chinensis transcriptome [20].
Data Processing
CodonW and EMBOSS were used to analyze transcriptome data for codon parameters. Neutrality plots, ENC-Plots, and PR2-bias Plots were generated using Excel and R language. Correlation analysis and heat maps of codon parameters were created with Origin Pro software.
Results
Analysis of Codon GC Content Composition
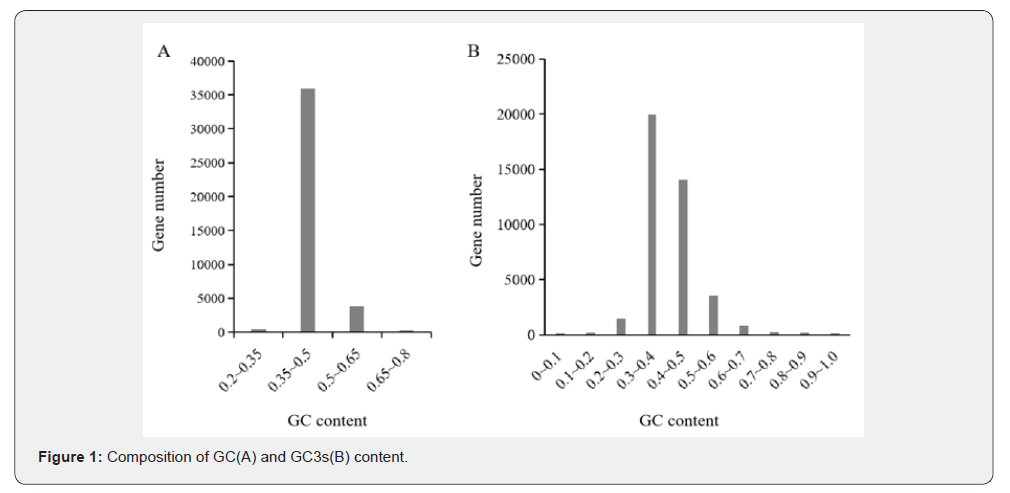
Using CodonW1.4.2 software, GC content analysis was performed on approximately 40, 309 CDS sequences from the Atractylodes chinensis transcriptome. As shown in Figure 1A, the total GC content was concentrated between 35% and 50%, with an average of 44.6%. The average GC3s content (GC content at the third codon position) was 43.26%, and most genes exhibited GC3s values primarily ranging from 30% to 50% (Figure 1B), indicating a preference for adenine (A) and uracil (U) at the third nucleotide position.
Neutrality Plot Analysis
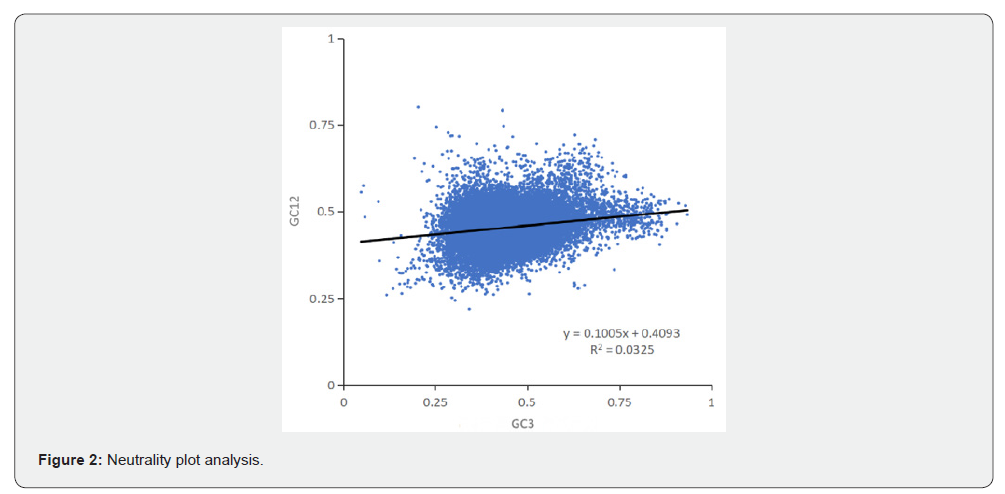
Neutrality plot analysis showed that GC12 (the average of GC contents at the first and second codon positions) had a relatively concentrated distribution, mainly clustering between 0.3 and 0.6, while GC3 (GC content at the third position) exhibited a more dispersed range of 0.25-0.75. The regression curve equation was y = 0.1005x + 0.4093, with a slope of 0.1005, R² = 0.0325, and a correlation coefficient r of 0.1803. The weak correlation among the first, second, and third codon position bases indicated that codon usage bias in A. chinensis was primarily regulated by selective pressure (Figure 2).
ENC-Plot Analysis
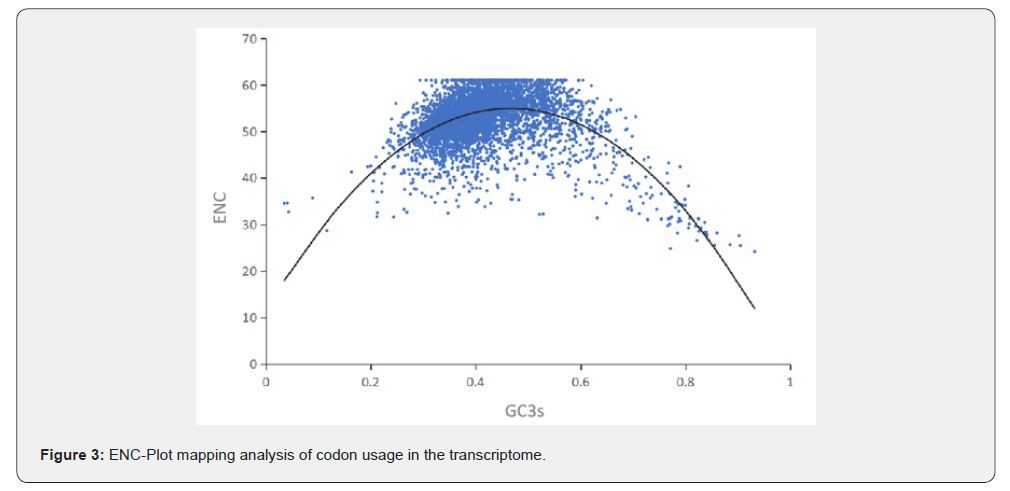
ENC-Plot analysis primarily serves to dissect the regulatory factors of codon usage bias, determining whether it is dominated by selective pressure or mutational pressure [21]. As shown by the distribution of data points for CDS sequences of Atractylodes chinensis transcriptome in Figure 3, most ENC values ranged from 35 to 61. Specifically, 367 sequences (0.9%) had ENC ≤ 35, indicating strong codon bias, while 1,839 sequences (4.6%) with ENC = 61 showed no codon bias. Additionally, the majority of genes were located below the standard curve, suggesting deviations between the actual ENC values and the ideal values, whereas a minority were distributed along or near the standard curve, indicating close proximity between the actual and calculated ideal ENC values. These results suggest that although mutational pressure plays a role in the codon bias of A. chinensis, selective pressure is the key factor in shaping its codon usage bias (Figure 3).
PR2-Bias Analysis
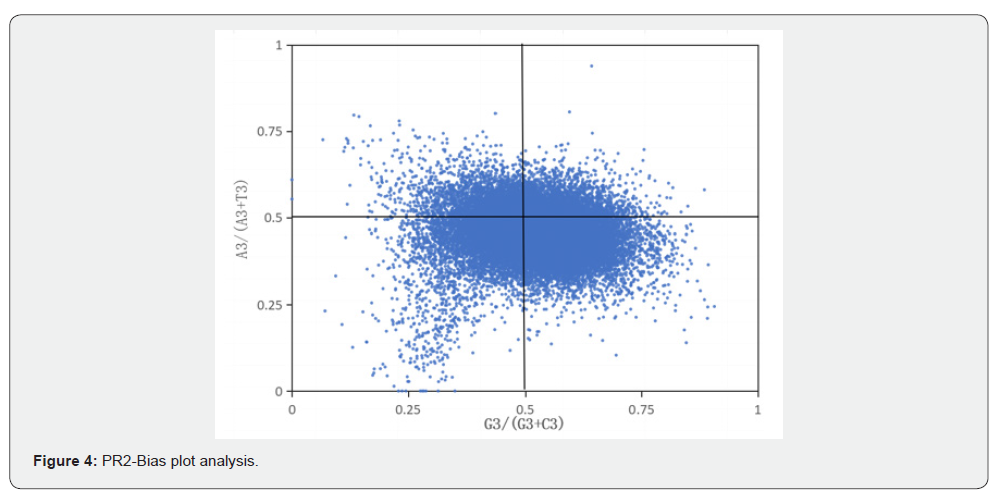
The PR2-Plot (Figure 4) revealed uneven distribution of genes across the four quadrants, with a general concentration in the lower two quadrants. Specifically, uracil (U) was used more frequently than adenine (A), and guanine (G) more than cytosine (C) at the third codon position. This demonstrated a preference in the usage of the third codon base, with relatively higher contents of thymine (T), cytosine (C), and guanine (G). These findings indicate that the codon usage bias in Atractylodes chinensis is regulated by both mutational factors and selective pressure.
Relative Codon Adaptation and Correlation Analysis of Transcriptome Parameters
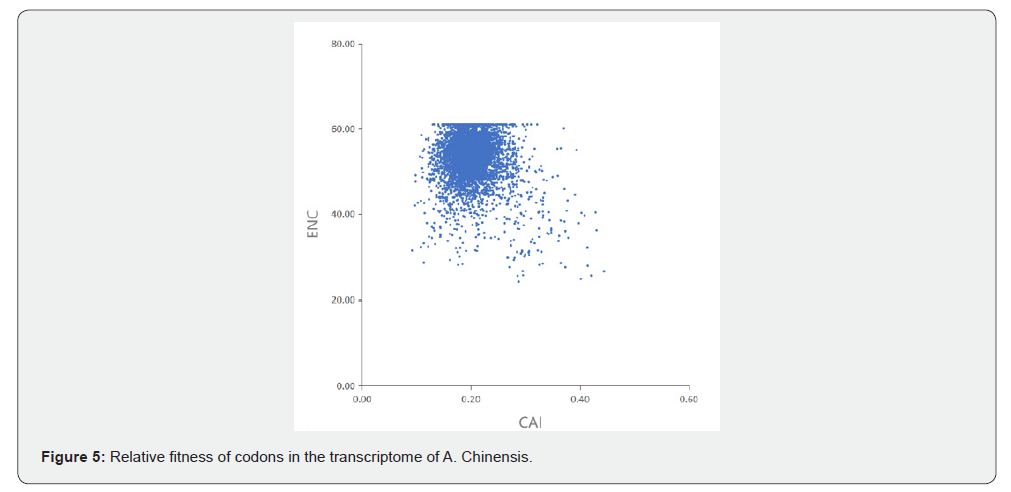
CAI (Codon Adaptation Index) is commonly used to evaluate gene expression levels [22]. In this study, CAI values ranged from 0.071 to 0.474, with the highest concentration around 0.2 (Figure 5), indicating low expression levels of Atractylodes chinensis transcriptome genes. This suggests that among the factors influencing codon bias in A. chinensis, natural selection exerts a stronger regulatory effect than mutational pressure. Correlation analysis of codon-related parameters in A. chinensis transcriptome sequences showed that GC1, GC2, and GC3 all exhibited significant correlations with the total GC content, but no significant correlations were observed among GC1, GC2, and GC3 themselves. Additionally, strong correlations were also observed between GC3, GC3s, GC and CAI, CBI (Codon Bias Index), FOP (Frequency of Optimal Codons). However, GRAVY (grand average of hydropathicity) showed no significant correlations with other parameters, even negative correlations in some cases, and ENC also showed no significant correlations with other parameters (Table 1).
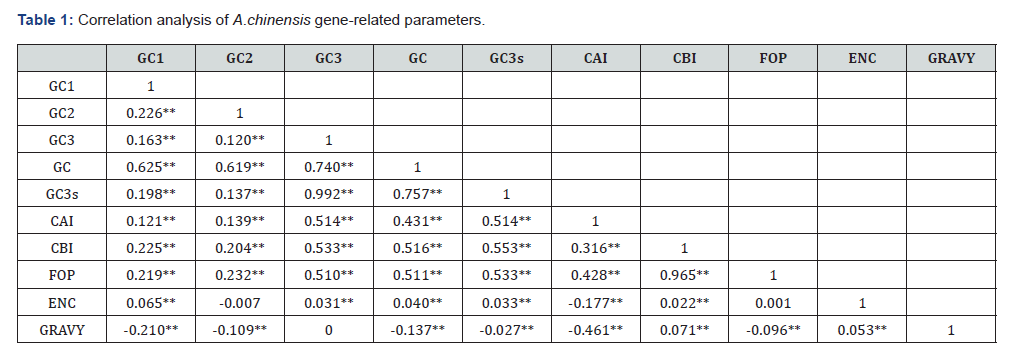
**P<0.01
Relative Synonymous Codon Usage (RSCU) and Optimal Codon Analysis
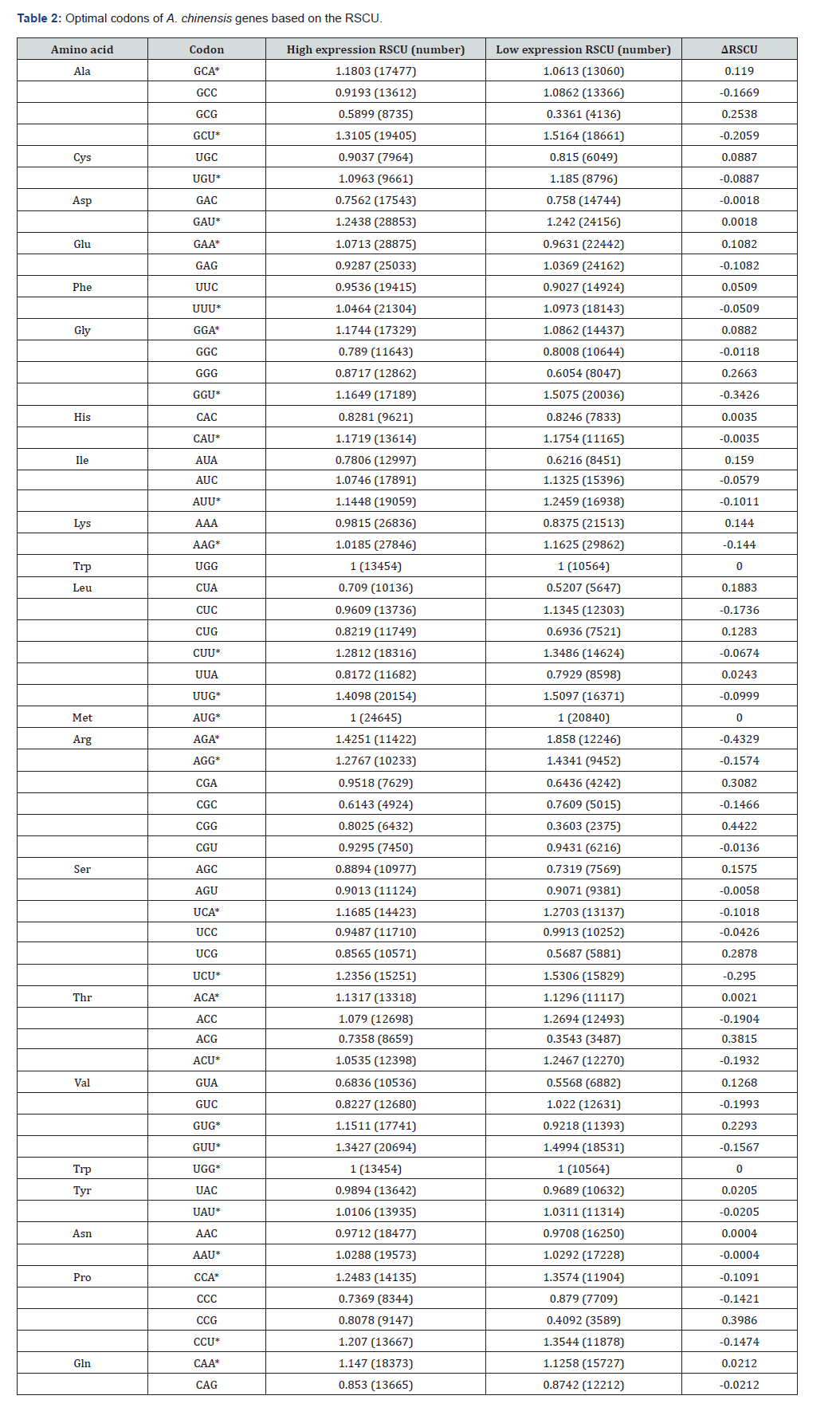
RSCU represents the ratio of the observed usage frequency of a specific codon to its expected frequency under random usage [23]. RSCU analysis of Atractylodes chinensis transcriptome sequences (excluding stop codons UAA, UAG, UGA) identified 28 high-frequency codons (RSCU > 1), including AUG, UGG, AAG, GUG, GAA, UAU, UUU, CAA, AAU, UGU, GGA, ACU, ACA, AUU, GGU, CAU, GCA, UCA, GAU, CCA, CCU, AGG, CUU, UCU, UUG, GCU, GUU, and AGA. Notably, most high-frequency codons ended with adenine (A) or uracil (U), indicating a terminal base preference for A/U. CDS sequences were sorted by ascending ENC values, and the top and bottom 10% were selected to construct high- and low-expression gene libraries. ΔRSCU (RSCU difference between libraries) was calculated, and codons with ΔRSCU > 0.08 were screened [24]. By integrating ΔRSCU > 0.08 with RSCU > 1 criteria, four optimal codons were identified: GCA, GAA, GGA, and GUG. This confirms a strong preference for adenine at the third codon position in A. chinensis (Table 2).
Discussion
Codon usage bias (CUB), a pivotal biological phenomenon, extends beyond fundamental science to exhibit profound applied implications. This study illuminates evolutionary trajectories, offering critical insights into biological evolution, genetic characteristics, and practical applications in germplasm improvement and therapeutic development. CUB emerges from organisms’ longterm interaction with the environment, governed by multiple regulatory factors-predominantly selective pressure and mutational bias [25-26]. However, debates persist in the field regarding the relative dominance of these forces in shaping codon bias.
Our analysis of 40, 309 coding sequences from the A.chinensis transcriptome revealed an average GC content of 44.6% (ranging 35%-50%) and a third-codon-position GC3s content of 43.26% (30%-50%), indicating weak codon bias with a pronounced preference for adenine (A) or uracil (U) at the third position. These findings align with studies on Ananas comosus [26], Camellia oleifera [27], and Canarium album [28], as the third codon baseless constrained by functional constraints-often serves as a key marker for CUB analysis [29]. Neutrality plot, ENC-Plot, and PR2- Plot analyses consistently validated selective pressure as the primary driver of CUB, echoing results from Dalbergia odorifera [30], Medicago sativa [31], and Sphaerophysa salsula [32].
The CAI values (0.071-0.474) reflected low gene expression levels in A. chinensis. RSCU analysis identified 28 high-frequency codons (RSCU > 1), of which 22 terminated with A/U and 6 with C/G. Notably, the optimal codons (GCA, GAA, GGA, GUG) also exhibited A/U endings, reinforcing the third-base preference. While CUB is influenced by multifaceted factors including selective pressure, mutational bias [33], gene expression levels [34], gene length [35], protein structure [36], tRNA abundance [37], and others [38- 39], our data unequivocally demonstrate that selective pressure constitutes the primary regulatory mechanism in A. chinensis. This study characterizes the weak CUB and low expression profiles in A. chinensis, identifying optimal codons and regulatory determinants. These insights deepen our understanding of genomic architecture and encoding mechanisms, providing a robust theoretical framework for future genetic research and biotechnological exploitation of this medicinal species.
Funding
This study was supported by the Chengde Science and Technology Project (202305B079) and the Natural Science Foundation of Hebei Province (H2022406053).
References
- Li H, Jin XH, Zhao BH (2019) Research Progress on Chemical Constituents and Pharmacological Activities of Atractylodes chinensis (DC.) Koidz. Agriculture of Jilin (3): 72-73.
- Liu XY, Li Y, Ji KK (2020) Genome-wide codon usage pattern analysis reveals the correlation between codon usage bias and gene expression in Cuscuta australis. Genomics 112(4): 2695-2702.
- Chinese Pharmacopoeia. Part 1. 2020: 168-169.
- Zhao C Y, Mao XX (2010) Research Progress on Chemical Components and Pharmacological Effects of Atractylodes chinensis (DC.) Koidz.[J]. J of Chengde Medical University 27(3): 309-311.
- Wang J, Wang TY, Wang LY (2019) Assembling and Analysis of the whole Chloroplast Genome Sequence of Elaeagnus angustifolia and Its Codon Usage Bias. Acta Botanica Boreali-Occidentalia Sinica 39(9): 1559-1572.
- Ji K, Song X, Chen G (2020) Codon Usage Profiling of Chloroplast Genome in Magnoliaceae. J of Agricultural Science and Technology 22(11): 52-62.
- Gao MQ, Zou JZ, Huo XW (2021) Analysis of Codon Usage patterns in Rheum officinale Based on Transcriptome Data. Chinese Traditional and Herbal Drugs 52(20): 6344-6349.
- Zheng YT (2020) Construction and Application of a Multifunctional Codon Analysis and Optimization Platform. Zhejiang University.
- Parvathy ST, Udayasuriyan V, and Bhadana V (2022) Codon usage bias, Molecular Biology Reports, 49(1): 539-565.
- Feng RY, Mei C, Wang HJ (2019) Analysis of Codon Usage Bias in Chloroplast Genome of Amaranthus hypochondriacs [J]. Chinese J of Grassland 41(04): 8-15.
- Ma ML, Zhang W, Meng HL (2021) Codon bias analysis of chloroplast genome in medicinal plants of Amomum Roxb. Chinese Traditional and Herbal Drugs 52(12): 3661-3670.
- Zhu H, Dai D, Wei Z (2024) Codon usage bias in chloroplast genomes of 17 Phoebe species. J of Southern Agriculture 55(12): 3646-3655.
- Gao SY, Li YY, Yang ZQ (2023) Codon usage bias analysis of the chloroplast genome of Bothriochloa ischaemum. Acta Prataculturae Sinica 32(07): 85-95.
- Li XH, Yang SC, Xin YX (2021) Analysis of the Codon Usage Bias of Chloroplast Genome in Erigeron brevis apus (Vant.) Hand-Mazz [J]. J of Yunnan Agricultural University (Natural Science) 36(03): 384-392.
- Rong Z, Wang J, Pei L (2025) Analysis on codon usage bias of chloroplast genomes in medicinal plants from genus Scutellarin. Chinese Traditional and Herbal Drugs 56(1): 269-281.
- Dan W, Jin Y, Tang Z (2020) Nucleotide composition and synonymous codon usage of open reading frames in Norovirus GII. 4 variants. J Biomol Struct Dyn 38(16): 4764-4773.
- Hu XY, Xu YQ, Han YZ (2019) Codon usage bias analysis of the chloroplast genome of Ziziphus jujuba var. spinosa. J of Forest and Environment 39(6): 621-628.
- Yuan X, Liu Y, Kang H (2021) Analysis of Codon Usage Bias in Chloroplast Genome of Malania oleifera. J of Southwest Forestry University(Natural Sciences) 41(3): 15-22.
- Mensah RA, Sun XL, Cheng CZ (2019) Analysis of codon usage pattern of banana basic secretory protease gene[J]. Plant Diseases and Pests 10(1): 1-49.
- Xiang H, Zhang RZ, Butler RR (2015) Comparative analysis of codon usage bias patterns in microsporidian genomes[J]. PLoS One 10(6): e0129223.
- Wang Y and Yang M (2021) Analysis of the Codon Usage Bias in the Chloroplast Genome of Allium mongolicum Regel. Molecular Plant Breeding 19(4): 1084-1092.
- Peixoto L, Zavala A, Romero H, Musto H (2003) The strength of translational selection for codon usage varies in three relicons of Sino rhizobium meliloti. Gene 320: 109-116.
- Mao L, Huang Q, Long L (2022) Comparative Analysis of Codon Usage Bias in Chloroplast Genomes of Seven Nymphaea Species. J of Northwest Forestry University 37(2): 98-107.
- Tgang DF, Wei F, Cai ZQ (2021) Analysis of codon usage bias and evolution in the chloroplast genome of Mesona chinensis Benth. Development Genes and Evolution 231(1-2): 1-9.
- Liu H, Lu Y, Lan B, Xu J (2020) Codon usage by chloroplast gene is bias in Hemiptelea davidii. J of Genetics 99: 8.
- Yang XY, Cai YB, Tan QL (2022) Analysis of Codon Usage Bias in the Chloroplast Genome of Ananas comosus. Chinese J of Tropical Crops 43(3): 439-446.
- Hao B, Xia Y, Ye H (2022) Analysis on codon usage bias of the chloroplast genome of Camellia osmantha. Journal of Central South University of Forestry & Technology 42(9): 178-186.
- Lai RL, Feng X, Chen J (2019) Codon Usage Bias and Its Influencing Factors in Transcriptome of Canarium album. J of Nuclear Agricultural Sciences 33(01): 31-38.
- Hu XY, Xu YQ, Han YZ (2019) Codon usage bias analysis of the chloroplast genome of Ziziphus jujuba var. spinosa. J of Forest and Environment 39(6): 621-628.
- Yuan XL, Li YQ, Zhang JF (2021) Analysis of Codon Usage Bias in the Chloroplast Genome of Dalbergia odorifera. Guihaia 41(04): 622-630.
- Yu F, Han M (2021) Analysis of codon usage bias in the chloroplast genome of alfalfa (Medicago sativa).Guihaia 41(12): 2069-2076.
- Liang XL, Guo S (2022) Analysis of Codon Usage Bias in Chloroplast Genome of Sphaerophysa salsula [J]. J of Northwest Forestry University 37(02): 121-126.
- Camiolo S, Melito S, Porceddu A (2015) New insights into the interplay between codon bias determinants in plants. DNA Res 22(6): 461-470.
- Paul P, Malakar AK, Chakraborty S (2018) Codon usage and amino acid usage influence genes expression level. Genetica 146(1): 53-63.
- Wada K, Wada Y, Ishibashi F (1990) Codon usage tabulated from the GenBank genetic sequence data. Nucleic Acids Res 18(Suppl): 2367-2411.
- Duret L, Mouchiroud D (1999) Expression pattern and, surprisingly, gene length shape codon usage in Caenorhabditis, Drosophila and Arabidopsis. Proc Natl Acad Sci USA 96(8): 4482-4487.
- Ikemura T (1985) Codon usage and t RNA content in unicellular and multicellular organisms. Mol Biol Evol 2(1): 13-34.
- Kimura M (1981) Possibility of extensive neutral evolution under stabilizing selection with special reference to nonrandom usage of synonymous codons. Proceedings of the National Academy of Sciences of the USA 78(9): 5773-5777.
- Lian CL, Yang H, Lan JX (2022) Comparative analysis of chloroplast genomes reveals phylogenetic relationships and intraspecific variation in the medicinal plant Isodon rubescens. PLoS ONE 17(4): e0266546.






























