- Review Article
- Abstract
- Introduction
- Molecular Markers
- Genotype identification and genetic variability
- Mapping quantitative trait loci
- Manipulating traits controlled by a few major loci
- Manipulating traits controlled by many loci
- Understanding the genetic architecture of quantitative traits
- Improving Efficiency of Breeding Programs
- Marker facilitated Study of complex populations
- Marker facilitated Study of Polyploids
- Revealing Evolutionary Relationships among Crop Species and Their Wild Relatives
- Cloning gene from map position
- Conclusion and Challenges for the Future
- References
Improvement of Crop Plants by Molecular Markers
Tibebu Belete*
*Department of Plant Sciences and Horticulture, Samara University, Ethiopia
Submission: May 17, 2018; Published: June 25, 2018
*Corresponding author: Tibebu Belete, College of Dry land Agriculture, Department of Plant Sciences and Horticulture, Samara University, P.O. Box 132, Samara, Ethiopia, Tel: +251911958252, E-mail: tibebubelet@su.edu.et
How to cite this article: Tibebu Belete. Improvement of Crop Plants by Molecular Markers. JOJ Hortic Arboric. 2018; 1(5): 555572
- Review Article
- Abstract
- Introduction
- Molecular Markers
- Genotype identification and genetic variability
- Mapping quantitative trait loci
- Manipulating traits controlled by a few major loci
- Manipulating traits controlled by many loci
- Understanding the genetic architecture of quantitative traits
- Improving Efficiency of Breeding Programs
- Marker facilitated Study of complex populations
- Marker facilitated Study of Polyploids
- Revealing Evolutionary Relationships among Crop Species and Their Wild Relatives
- Cloning gene from map position
- Conclusion and Challenges for the Future
- References
Abstract
Improvements in marker detection systems and in the techniques used to identify markers linked to useful traits, has enabled great advances to be made in recent years. While RFLP markers have been the basis for most works in crop plants, valuable markers have been generated from RAPDs and AFLPs. Simple Sequence Repeats (SSR) or microsatellite markers have been developed more recently for major crop plants and this marker system is predicted to lead to even more rapid advances in both marker development and implementation in breeding programs. Identification of markers linked to useful traits has been based on complete linkage maps and bulked segregant analysis. However, alternative methods, such as the construction of partial maps and combination of pedigree and marker information, have also proved useful in identifying marker/trait associations. The development of molecular techniques for genetic analysis has led to a great increase in our knowledge of cereal genetics and our understanding of the structure and behavior of plant genomes. These molecular techniques, in particular the use of molecular markers, have been used to monitor DNA sequence variation in and among the species and create new sources of genetic variation by introducing new and favorable traits from landraces and related species.
Keywords: Molecular marker; Genetic variability; QTL; Crop improvement
Abbreviations: RFLPs: Restriction Fragment Length Polymorphism; RAPDs: Random Amplified Polymorphic DNAs; STS: Sequence Tagged Sites; AFLPs: Amplified Fragment Length Polymorphism; SNPs: Single Nucleotide Polymorphisms
- Review Article
- Abstract
- Introduction
- Molecular Markers
- Genotype identification and genetic variability
- Mapping quantitative trait loci
- Manipulating traits controlled by a few major loci
- Manipulating traits controlled by many loci
- Understanding the genetic architecture of quantitative traits
- Improving Efficiency of Breeding Programs
- Marker facilitated Study of complex populations
- Marker facilitated Study of Polyploids
- Revealing Evolutionary Relationships among Crop Species and Their Wild Relatives
- Cloning gene from map position
- Conclusion and Challenges for the Future
- References
Introduction
Conventional plant breeding is time consuming and very dependent on environmental conditions. Breeding a new variety takes between eight and twelve years and even the release of an improved variety cannot be guaranteed. Hence, breeders are extremely interested in new technologies that could make this procedure more efficient. Molecular marker technologies offer such a possibility by adopting a wide range of novel approaches to improving strategies in plant breeding.
There have been several reports of the potential applications of molecular markers to plant improvement [1-3], but few reports have analyzed breeding programs in detail to identify applications having the greatest potential. Plant breeding in its simplest form consists of (a) generation of genetic variation through recombination and (b) selection to identify superior recombinants for advancement in the breeding program. All breeding methods share these two steps, but the protocols used within each step are complicated and differ with breeding method, breeding objective, and species [4].
Understanding the application of molecular markers to plant breeding requires a detailed understanding of plant-breeding methodology (resource allocation [economics], progeny testing, reproductive biology [cycle time]), quantitative genetics, and statistics. Knowledge of breeding methodology is required to understand how current cultivars and hybrids are developed. Knowledge of quantitative genetics and statistics is required to understand the theoretical basis of plant-breeding methodology; the design, implementation, and interpretation of plant breeding experiments; and the statistical power required to detect differences among cultivars and hybrids.
For the program to be successful, economics of cultivar development must be considered simultaneously with design of the breeding program. These areas represent a vast knowledge base that has expanded over the past 90 years. Many molecular marker experiments conducted today could not have been done without this knowledge base.
- Review Article
- Abstract
- Introduction
- Molecular Markers
- Genotype identification and genetic variability
- Mapping quantitative trait loci
- Manipulating traits controlled by a few major loci
- Manipulating traits controlled by many loci
- Understanding the genetic architecture of quantitative traits
- Improving Efficiency of Breeding Programs
- Marker facilitated Study of complex populations
- Marker facilitated Study of Polyploids
- Revealing Evolutionary Relationships among Crop Species and Their Wild Relatives
- Cloning gene from map position
- Conclusion and Challenges for the Future
- References
Molecular Markers
In recent years, different systems such as Restriction Fragment Length Polymorphism (RFLPs), Random Amplified Polymorphic DNAs (RAPDs), Sequence Tagged Sites (STS), Amplified Fragment Length Polymorphism (AFLPs), Simple Sequence Repeats (SSRs) or microsatellites, Single Nucleotide Polymorphisms (SNPs) and others have been developed and applied to a wide range of crop species. The relative advantages and disadvantages of these techniques are summarized in Table 1.
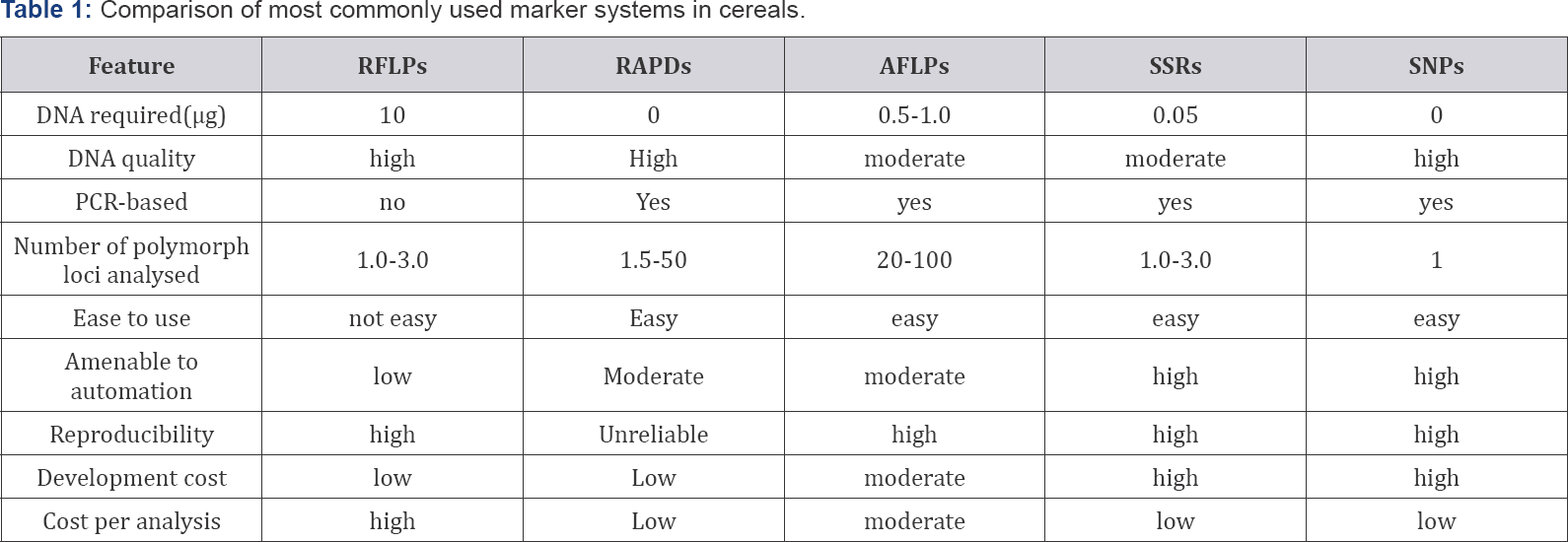
The information suggests the RFLP, SSR and AFLP markers are the most effective in detecting polymorphism. However, given the large amount of DNA required for RFLP detection and the difficulties in automating RFLP analysis, AFLPs and SSRs are currently the most popular markers in cereals. An increasing amount of sequence information and the determination of the gene function in cereals will lead in the near future to the preferred use of new marker types, such as SNPs. Application of these markers for genetic studies of cereals have been so much diverse [3]. Main uses include:
Assessment of genetic variability and characterization of germ plasm;
a) Identification and fingerprinting of genotypes;
b) Estimation of genetic distances between population,inbreeds and breeding material;
c) Detection of monogenic and quantitative trait loci (QTL);
d) Marker- assisted selection;
e) Identification of sequences useful of candidate genes,etc.
- Review Article
- Abstract
- Introduction
- Molecular Markers
- Genotype identification and genetic variability
- Mapping quantitative trait loci
- Manipulating traits controlled by a few major loci
- Manipulating traits controlled by many loci
- Understanding the genetic architecture of quantitative traits
- Improving Efficiency of Breeding Programs
- Marker facilitated Study of complex populations
- Marker facilitated Study of Polyploids
- Revealing Evolutionary Relationships among Crop Species and Their Wild Relatives
- Cloning gene from map position
- Conclusion and Challenges for the Future
- References
Genotype identification and genetic variability
It is vital for plant breeding programmes to have sufficient diversity available to allow for the production of new varieties that are aimed towards the improvement of crop productivity and able to withstand damage from biotic and abiotic factors. Assessment of genetic diversity is important in plant breeding if there is to be improvement by selection. For assessment of genetic diversity, molecular markers have been generally superior to morphological, pedigree, heterosis, and biochemical data (isozymes and chromatography) [5,6] Genetic diversity commonly is measured by genetic distance (GD) or genetic similarity (GS=1-GD), both of which imply that there are either differences or similarities at the genetic level and the former can be used as an estimation of expected genetic variances in different sets of segregating progenies derived from different crosses.
Melchinger [6] reviewed the application of molecular marker-based GD for assigning maize inbred lines to heterotic groups, determining the relation between inbred lines and hybrids, and predicting hybrid performance. Data showed that GS calculated from molecular marker data faithfully separated inbred lines into their heterotic groups. There also seems to be promise of assigning inbreeds of unknown pedigree to heterotic groups although a large number of markers (>100) and well- characterized reference populations may be needed to obtain an accurate assessment of GS. Strong correlations (0.61 to 0.95) between Malecot's coancestry coefficient (f) and GS for related (f>0) genotypes indicated that pedigree data provide reliable estimates of GS. Genetic similarity estimates based on molecular markers are expected to be superior to estimates of f because of unreliable or incomplete pedigree data and because of the assumptions required to calculate f. Molecular marker-based GD has some potential for predicting hybrid performance of related lines, but in typical hybrid-breeding programs, in which hybrids are produced from unrelated lines from different heterotic groups, molecular marker-based GD has been of no value in predicting hybrid performance. These results suggest the use of molecular marker-based GD for predicting hybrid performance in crops in which either hybrids are being explored, such as wheat (Triticum aestivum [Vill., Host] Mackey), and soybean (Glycine max (L.) Merr.), or hybrid breeding is being practiced, but distinct heterotic groups have not been developed.
Molecular marker-based GD also has potential for assessing changes in genetic diversity over time [7], protection of intellectual property rights [8], registration of germ plasm in countries having ratified the rules of the UPOV convention, and evaluation of new sources of germ plasm for their potential to increase genetic diversity [9].
Direct applications to plant breeding, however, have been limited so far to prediction of hybrid performance. But accurate prediction of hybrid performance does not seem likely unless gene action is primarily dominant or overdominant, complementary heterotic groups are established, trait heritability is high, at least 30 to 50% of the markers are linked to QTL, and no more than 20 to 30% of the markers are dispersed randomly [10]. Molecular markers, however, may be useful for early generation testing in hybrid-breeding programs. If individual markers or marker intervals associated with combining ability can be identified when a plant or progeny is crossed onto a given tester, then these markers could be used as a first screen to identify the top 50% of the progenies for field evaluation. Although this procedure would not decrease the time to cultivar development, it would decrease the amount of material tested or permit the evaluation of a wider range of germplasm for the same amount of field resources.
An equally important application of molecular marker-based GD may be in the selection of parents to cross in a breeding program. This application deserves serious attention because breeders currently rely primarily on pedigree and performance data for choosing parents in breeding programs. Using molecular markers to select parents has the potential to allow simultaneous maintenance of genetic diversity and performance. Dudley et al. [11] presented one application of molecular markers for choosing parents, and additional research is needed in this area. Using molecular markers to choose parents likely will require establishment of a relation between GD and genetic variation, and many of the same conditions necessary for predicting hybrid performance may be required for choosing parents. Using molecular markers as a diagnostic tool to survey new or exotic germplasm for novel genetic diversity also may be possible. It is unlikely, however, that this use will be possible with random genomic or cDNA clones because molecular marker-based genetic diversity will not guarantee genetic diversity for the traits of interest. Screening with probes of expressed genes with known function offers the greatest potential in this area.
DNA fingerprinting of crop species and cultivated varieties has a long scientific history. When DNA profiling technology first came in to use, restriction fragment length polymorphism (RFLP) were considered state-of-art. RFLP technology was followed random amplification of polymorphic DNA (RAPD), followed by amplified length polymorphism (AFLP) and most recently use microsatellite markers or single sequence repeat (SSR). Advantages of SSR markers are:
a) The method is relatively simple and can be automated;
b) Most ofthe markers are monolocus and show Mendelian inheritance;
c) SSR markers are high informative;
d) A high number of public SSR primer pairs are available;
e) Effective cost per genotype and primer (similar to that for RAPD).
- Review Article
- Abstract
- Introduction
- Molecular Markers
- Genotype identification and genetic variability
- Mapping quantitative trait loci
- Manipulating traits controlled by a few major loci
- Manipulating traits controlled by many loci
- Understanding the genetic architecture of quantitative traits
- Improving Efficiency of Breeding Programs
- Marker facilitated Study of complex populations
- Marker facilitated Study of Polyploids
- Revealing Evolutionary Relationships among Crop Species and Their Wild Relatives
- Cloning gene from map position
- Conclusion and Challenges for the Future
- References
Mapping quantitative trait loci
Using molecular markers, QTLs can be described by their chromosomal location, dosage effect, phenotypic effect(s), and sensitivity to environment. Such a description has long been possible for genes affecting simply inherited traits; however, only with the availability of high-density genetic maps has it become possible to obtain this information for individual QTLs.
- Review Article
- Abstract
- Introduction
- Molecular Markers
- Genotype identification and genetic variability
- Mapping quantitative trait loci
- Manipulating traits controlled by a few major loci
- Manipulating traits controlled by many loci
- Understanding the genetic architecture of quantitative traits
- Improving Efficiency of Breeding Programs
- Marker facilitated Study of complex populations
- Marker facilitated Study of Polyploids
- Revealing Evolutionary Relationships among Crop Species and Their Wild Relatives
- Cloning gene from map position
- Conclusion and Challenges for the Future
- References
Manipulating traits controlled by a few major loci
The manipulation of traits controlled by a few major loci (loci that can be studied using Mendelian genetics) may offer the greatest promise in the short term for application to plant breeding. The primary applications of this technique will be for traits controlled by a single gene (monogenic) or those controlled by at most two or three loci (oligogenic). The most successful applications will be in those species with well developed molecular marker maps. These applications will be immediately useful for "defensive breeding,” that is, when a desirable genotype is available but lacks resistance to important insects and diseases. Other applications, not fitting into the category of defensive breeding may include seed modifications controlled by a few genes, restorer genes for cytoplasmic male sterility, dwarfing genes for shorter plant height, and maturity genes for adaptability.
The first requirement of using molecular markers in this context is to develop a precise molecular marker linkage map and then to use these markers to map gene(s) controlling the trait of interest. Many methods for mapping genes of interest are available, including a variety of applications suitable to most species with polymorphic markers [12]. The final step is to use marker-facilitated selection to transfer the gene(s) to the genotypes desired. Two methods are available, both of which begin with inbred lines, backcross selection, and pedigree selection.
If the cost of molecular-marker technology is ignored, the primary factor affecting the design and the success of marker- facilitated selection is how tightly linked a single marker is to the gene or how tightly bracketed the gene is by two markers. The idea is to obtain a marker or a set of flanking markers linked tightly enough to the gene so that a recombination event does not occur between the marker and the gene during backcrossing or pedigree selection. Melchinger [12] presented extensive theoretical and numerical results for the backcross method concerning the optimal family size and the number of plants per family that must be genotyped with molecular markers. The results are complicated and will have to be assessed on a case by case basis. The economics of marker assisted backcrossing will be a function of the cost of marker assays, the cost of direct screening, and the value of accelerating the backcrossing program. Results regarding sample sizes required for pedigree selection are unavailable, although marker-gene linkage is the primary consideration [11].
There are many practical applications of molecular markers to traits controlled by few loci [12]. Most of the applications involve situations in which either screening for the trait is difficult or scoring of the trait occurs late in plant development. These applications may include pests for which natural inoculum is unreliable or artificial inoculation procedures are undeveloped or unreliable. Examples include nematodes or Aspergillus, both of which have broad host ranges and unreliable natural and artificial inoculation. Diseases in which resistance is influenced strongly by the environment also would be good candidates for marker facilitated selection. Unfortunately, the very situations favoring marker-facilitated selection also make it difficult to map the resistance genes precisely. Marker-facilitated selection has been advantageous for backcrossing recessive genes, when progeny tests are needed after every backcross generation to identify heterozygotes or when resistance can be determined only after flowering. Markers in these situations could greatly reduce workload and backcrossing time. Other examples include pyramiding resistance genes, developing multilines in which many race-specific resistance genes are involved that are sometimes difficult to distinguish, and selecting for resistance to exotic or quarantined pathogens.
One of the primary advantages of marker-facilitated backcrossing has been in increasing the speed of recovery of the recurrent parent genome [13]. In addition to having a tight marker-gene linkage, one or more neutral polymorphic markers will be required per chromosome arm. The idea is to screen for plants having the resistance gene and to identify those plants with the greatest proportion of markers homozygous for the recurrent parent. A possible limitation of this procedure is that there may be an increase in the number of plants needing to be assayed with markers. The procedure could be applied, however, to subsequent backcross generations to ensure the recovery of unlinked segments of recurrent parental genome.
- Review Article
- Abstract
- Introduction
- Molecular Markers
- Genotype identification and genetic variability
- Mapping quantitative trait loci
- Manipulating traits controlled by a few major loci
- Manipulating traits controlled by many loci
- Understanding the genetic architecture of quantitative traits
- Improving Efficiency of Breeding Programs
- Marker facilitated Study of complex populations
- Marker facilitated Study of Polyploids
- Revealing Evolutionary Relationships among Crop Species and Their Wild Relatives
- Cloning gene from map position
- Conclusion and Challenges for the Future
- References
Manipulating traits controlled by many loci
The molecular marker manipulation of traits controlled by many loci (from a plant breeding perspective many is generally greater than five) is of great interest to plant breeders and represents one of the fields greatest challenges. Plant breeders concentrate effort on breeding for quantitative traits, and breeding for qualitative traits is generally a trivial, albeit timeconsuming, process. The matter is further complicated because breeders usually evaluate simultaneously in many populations four or more complexly inherited traits. Obviously, any technology enhancing the breeders' ability to select for these traits would be adopted rapidly.
The molecular marker manipulation of qualitative traits is feasible precisely because so much is known about the biology of these traits. The number of loci is known; there are no questions about what a major locus is in fact, many of these loci have been cloned; the gene action is known precisely; epistatic interactions, if any, are relatively easy to decipher; and interactions with environment are easy to determine. In short, the amount that can be known about qualitative traits is limited only by one's desire to know. As pointed out earlier, for quantitative traits, the answers to these questions are based only upon averages over an unknown number of loci. At the outset, the manipulation of quantitative traits by molecular markers has a serious disadvantage.
Even with these limitations, there still may be many applications of marker-QTL associations. But very little theoretical or applied research has been published in this area. The most-cited application has been marker-assisted selection (MAS) although the context of application often has been ignored. In plant breeding, there are two distinct but related applications: germplasm enhancement (recurrent selection) and cultivar or hybrid development. These two applications are separated because recurrent selection usually is applied to random mating populations possibly at or near linkage equilibrium, whereas cultivar or hybrid development typically begins with populations derived by crossing elite inbred lines at or near maximum linkage disequilibrium. Clearly, two different approaches are needed for these breeding schemes.
Lande & Thompson [14] and Lande [15] investigated the efficiency of MAS for both individual and mass selection in random-mating populations. There are three approaches to applying MAS to plant breeding: (1) selection on markers alone with no measurement of phenotype; (2) simultaneous selection on markers and phenotype; and (3) two-stage selection, the first stage involving use of markers to select among seedlings and the second involving phenotypic selection among surviving adults. On individuals, the efficiency of MAS relative to that of phenotypic selection of the same intensity is (p/h2)1/2, where p is the proportion of the additive genetic variance accounted for by markers, and h2 is the heritability. Selection on markers alone will be more efficient than phenotypic selection only when the proportion of the genetic variance explained by markers exceeds the heritability of the trait. Therefore, selection on markers alone will be most useful for traits with low heritabilities when large proportions of their variability have been explained by markers. Lande & Thompson [14] concluded that molecular marker loci can be used to enhance the efficiency of artificial selection for quantitative traits. The potential efficiency of MAS depends upon the heritability of the trait, the proportion of the genetic variance explained by the markers, and the selection method. A major practical problem in using MAS is that recombination will reduce linkage disequilibrium between the markers and QTL, thus diminishing selection effectiveness. The successful application of MAS will require very tight linkages between markers and QTL.
- Review Article
- Abstract
- Introduction
- Molecular Markers
- Genotype identification and genetic variability
- Mapping quantitative trait loci
- Manipulating traits controlled by a few major loci
- Manipulating traits controlled by many loci
- Understanding the genetic architecture of quantitative traits
- Improving Efficiency of Breeding Programs
- Marker facilitated Study of complex populations
- Marker facilitated Study of Polyploids
- Revealing Evolutionary Relationships among Crop Species and Their Wild Relatives
- Cloning gene from map position
- Conclusion and Challenges for the Future
- References
Understanding the genetic architecture of quantitative traits
Although there have been few direct applications of molecular markers in plant breeding, published research has begun to provide information on the genetic architecture of quantitative traits. Molecular marker-mapping data from several species now suggest that genetic variation for quantitative traits is controlled by a few loci with large effects and many loci with progressively smaller effects [15]. Beavis et al. [16] found 14 genomic regions associated with plant height in four F2 maize populations, but few of these regions were in common across populations. All 14 regions, however, were associated with known qualitative genetic loci. Thus, circumstantial evidence supports that QTL have alleles with a range of effects, alleles with large effects causing recognition of the locus as qualitative, and alleles with small effects causing recognition of the locus as quantitative.
The results from QTL-mapping studies regarding gene action and epistasis are, so far, difficult to interpret. Stuber et al. [17] reported that, with one exception, all the QTL mapped for grain yield in maize showed heterozygote superiority. Either over dominance or pseudo-over dominance, therefore, likely was important in the mapping population. These results cannot separate the two causes of heterozygote superiority, primarily because the number of QTL residing in a marker interval cannot be resolved. Although not mentioned by the authors, heterozygote superiority also could result from overestimation of effects as pointed out by Lande and Thompson [14] and Lande [15]. Stuber et al. [17] found little evidence for epistasis in maize and tomato (Lycopesicon esculentum [E]), respectively. Lateef [18] concluded that molecular marker-mapping studies conducted to date may identify preferentially QTL that function independently of unlinked genetic factors. Those researchers suggested that the role of epistasis in quantitative inheritance needs to be studied in larger populations, with more closely spaced markers, and/ or with specially constructed genetic stocks carrying particular QTL.
Stuber et al. [17] found limited evidence for G x E interaction even though a standard analysis of the data revealed significant G x E interaction for many of the traits. Thus it may be possible to detect QTL with large effects in relatively few environments. Lateef [18] reported that individual QTL showed a range of sensitivities to environments, some QTL being detected in all environments and some being detected in only one environment.
- Review Article
- Abstract
- Introduction
- Molecular Markers
- Genotype identification and genetic variability
- Mapping quantitative trait loci
- Manipulating traits controlled by a few major loci
- Manipulating traits controlled by many loci
- Understanding the genetic architecture of quantitative traits
- Improving Efficiency of Breeding Programs
- Marker facilitated Study of complex populations
- Marker facilitated Study of Polyploids
- Revealing Evolutionary Relationships among Crop Species and Their Wild Relatives
- Cloning gene from map position
- Conclusion and Challenges for the Future
- References
Improving Efficiency of Breeding Programs
Plant breeders have developed many sophisticated strategies for accomplishing particular breeding objectives in different species, and under different situations [19]. However, these strategies are similar to one another in that breeding progress is largely determined by three factors:
a) Identifying genetic variation among different individuals, cultivars, races, or species which show different attributes, or which show a common attribute but due to different genes.
b) Making crosses and obtaining recombination between genotypes with different attributes, creating genotypes with new sets of attributes (or superior levels of a particular attribute).
c) Accurate selection of these occasional genotypes with new sets of attributes, from large populations grown in variable environments and including many inferior genotypes.
Each of these areas can benefit from use of genetic markers, especially DNA markers. Genetic markers represent genetic variation, permitting one to estimate relatedness between different genotypes, and consequently to predict which matings might produce new and superior gene combinations. Further by having markers for genes of interest, one can readily detect recombination between these genes, and perform accurate selection for genetically superior individuals, from among the masses of candidates including many pretenders who were favored by environment, rather than genetics.
A large number of cereal studies have used markers as a tool to identify major genes, QTLs, or to introduce new characters in elite germplasm. In wheat, for example, molecular markers have been identified that are associated with around 40 traits of economic importance. Knowing the location of these genes/ traits and specific alleles offers the possibility to apply marker- assisted selection (MAS) in cereals, because one of the main objectives of plant breeding is the introgression of one or more favorable genes from a donor parent in to the background of an elite variety. Marker-assisted selection allows plant selection at the juvenile stage from an early generation. For simply inherited traits, conventional PCR, which requires a small amount of DNA, is becoming very useful for screening large populations of segregating progenies. Unfavorable alleles can be eliminated or greatly reduced during the early stages of plant development through MAS, focusing the selection in the field on reduced numbers of mature plants [2].
- Review Article
- Abstract
- Introduction
- Molecular Markers
- Genotype identification and genetic variability
- Mapping quantitative trait loci
- Manipulating traits controlled by a few major loci
- Manipulating traits controlled by many loci
- Understanding the genetic architecture of quantitative traits
- Improving Efficiency of Breeding Programs
- Marker facilitated Study of complex populations
- Marker facilitated Study of Polyploids
- Revealing Evolutionary Relationships among Crop Species and Their Wild Relatives
- Cloning gene from map position
- Conclusion and Challenges for the Future
- References
Marker facilitated Study of complex populations
Most natural populations of plants, as well as many experimental populations, are genetically quite different from the classical linkage mapping populations. While linkage mapping populations are commonly derived from two generation crosses between two parents, many natural and experimental populations are derived from multi-generation matings between assortments of different parents, resulting in a massive reshuffling of genes. Individuals in such populations carry a complex mosaic of genes, derived from a number of different founders of the population. Gene frequencies in the populations as a whole may be modified by natural or artificial selection, or by genetic drift (e.g. chance) in small populations.
Study of such complex populations has yielded much information on both adaptation by natural selection [20], and on the genetic change as a result of artificial selection [17].These two modes of selection are quite different, since natural selection tends to favour an intermediate phenotype, which is adequately suited to a number of diverse environments [21], while artificial selection tends to favour an extreme phenotype, which gives superior performance in a rigidly defined environment.
Classical quantitative genetics was a starting point for investigating genetic change under long-term selection regimes; however, simply measuring the phenotype of an individual provided insufficient information about the underlying genetic factors [20].
Given a complex population with superior average expression of a trait, the breeder might wish to (1) maintain or improve the expression of the trait of interest, while maintaining desirable levels of other traits; and (2) maintaining sufficient genetic diversity that rare desirable alleles influencing the trait(s) of interest are not lost before their frequency can be increased by selection. Genetic markers might be especially valuable in accomplishing this second objective; for example, one might select a fraction of the population based on favorable phenotype (perhaps the several traits -one might readily employ index selection), the apply genetic markers to this fraction and keep a subset which represent much of the allelic diversity within the population.
Most natural populations and many synthetic populations are genetically complex. Genetic markers have provided a glimpse at the nature of changes which underlie the continuous process of phenotypic selection, both natural and artificial. Strategies for extracting a minimum of desirable phenotypic variation from such populations remain an important area of breeding research. An integrated approach merging classical phenotypic selection with molecular marker based analysis of, may aid in extracting valuable genes from heterogeneous populations. Again, strategies for studying heterogeneous populations would be of special value in species of large stature or long generation time, where one might gain access to heterogeneous natural or cultivated populations, rather than waiting for two generations of crossing to produce ideal mapping populations.
- Review Article
- Abstract
- Introduction
- Molecular Markers
- Genotype identification and genetic variability
- Mapping quantitative trait loci
- Manipulating traits controlled by a few major loci
- Manipulating traits controlled by many loci
- Understanding the genetic architecture of quantitative traits
- Improving Efficiency of Breeding Programs
- Marker facilitated Study of complex populations
- Marker facilitated Study of Polyploids
- Revealing Evolutionary Relationships among Crop Species and Their Wild Relatives
- Cloning gene from map position
- Conclusion and Challenges for the Future
- References
Marker facilitated Study of Polyploids
Unlike the diploid species, several agriculturally important species are polyploid, containing more than two copies of each chromosome. Some polyploids, called allopolyploids, are similar to diploids in segregation and recombination. Allopolyploids contain several sets of chromosomes which are derived from a common ancestor, but have diverged enough to be readily distinguishable and thus, usually pair with only one partner during sexual reproduction. Examples of allopolyploids include wheat (6x), Oat (4x). Other polyploids, called autopolyploid, are very different from diploids in segregation and recombination. Auto polyploids contain several sets of chromosomes which are not readily distinguishable and thus, a particular chromosome may pair with different partners in different regions.
Genetics and gene mapping in auto polyploids is complicated by the presence of several copies of each chromosome. Rather than being limited to having 0,1, or 2 copies of an ancestral gene like diploids, polyploids can have from 0 to x copies ( x being the number of copies of each chromosome).Hence, study of gene dosage effects, and also of epistasis, is much more complicated than in diploids. Further, study of recombination is quite complicated in auto polyploids, because of multiple opportunities for chromosome pairing. Finally, one seldom has enough genetic markers to distinguish each of (for example) four homologous chromosomes; more commonly, one chromosome can be distinguished from the other three. Consequently, only a fraction of the total number of recombination events can be studied using genetic markers; thus, one must study more individuals to compensate for the loss of information.
One method of linkage analysis in polyploids [22] involves the identification of bands that represent a single dose restriction fragment (SDRF).In this approach, each restriction fragment is analyzed for its presence of absence in the progeny If the fragment is represented by a single dose, it will segregate 1:1 (presence: absence) in the gametes or 3:1 in the selfed progeny, regardless of whether the species is allopolyploid or autopolyploid. Single-dose restriction fragment markers in auto ployploids represent the genotype of one parent or the other, and thus not discriminate between individuals which are heterozygous for those which are homozygous for the parental fragment (unlike co dominant markers used in the study of diploids). Estimation of recombination distance between two SRDF markers is similar to that for diploids, except that the case of repulsion phase linkage (presence of a marker at one locus linked to absence of a marker at a nearby locus) is very difficult to detect using SRDFs.
Population or family sizes required for detecting SRDFs and their linkage have been estimated by Wu et al. [22]. If one considers a cross between two individuals, one of which has a particular band that is lacking in the other, the first step is to test the hypothesis that the band represents a SRDF (not a multiple- dose band).
In view of the complicated genetics associated with auto ployploids, it comes as little surprise that the genetic maps of these species are as yet less detailed than those for many diploids and allopolyploids. At the same time, the inherent complexity of polyploid genetics may add even greater value to the simplicity achieved by identifying markers for a trait. Markers might permit the breeder to identify the individuals carrying different desirable genes, and him/her to select transgressive progeny from crosses between such individuals. Breeding objectives which have classically been cumbersome, relying heavily up on phenotypic selection among random-mated individuals of unknown genotype, might be accomplished more rapidly and efficiently with the information added by DNA markers.
- Review Article
- Abstract
- Introduction
- Molecular Markers
- Genotype identification and genetic variability
- Mapping quantitative trait loci
- Manipulating traits controlled by a few major loci
- Manipulating traits controlled by many loci
- Understanding the genetic architecture of quantitative traits
- Improving Efficiency of Breeding Programs
- Marker facilitated Study of complex populations
- Marker facilitated Study of Polyploids
- Revealing Evolutionary Relationships among Crop Species and Their Wild Relatives
- Cloning gene from map position
- Conclusion and Challenges for the Future
- References
Revealing Evolutionary Relationships among Crop Species and Their Wild Relatives
An important use of molecular markers has been to attempt to discern evolutionary relationships, within and between species, genera, or larger taxonomic groupings. Such studies involve studying similarities and differences among taxa, using numerous genetic markers. Although phylogenetic trees have previously been established for many species on the basis of visible and isozyme markers and chromosome homology, e.g. tomato [23] wheat [24], DNA markers have recently added to the breadth of phylogenetic information available for a number of species. Such studies are important in classifying newly discovered germplasm, and in establishing possible sources from which valuable traits might be readily being transferred to crop species. This later objective also relies heavily upon studies of cross ability among related species [25].
In a few cases, it has been possible to reveal the consequences of evolutionary divergence on chromosome organization in crop species. These consequences of evolutionary divergence are of academic interest, and also have much practical value. By defining the sites of chromosomal rearrangement, one also defines intervening regions in which genes are arranged similarly in different organisms. To a lesser degree, comparative may also be possible in regard to QTLs. It was suggested that QTL mapping information from one pedigree might somewhat predictive of QTL locations in other pedigrees, or in this particular case, other species [3].
- Review Article
- Abstract
- Introduction
- Molecular Markers
- Genotype identification and genetic variability
- Mapping quantitative trait loci
- Manipulating traits controlled by a few major loci
- Manipulating traits controlled by many loci
- Understanding the genetic architecture of quantitative traits
- Improving Efficiency of Breeding Programs
- Marker facilitated Study of complex populations
- Marker facilitated Study of Polyploids
- Revealing Evolutionary Relationships among Crop Species and Their Wild Relatives
- Cloning gene from map position
- Conclusion and Challenges for the Future
- References
Cloning gene from map position
The sophisticated molecular tools and techniques available for studying individual genes are of much potential use to the crop breeder. Some of this potential can be realized by marker -facilitated selection that is, using linked DNA markers to select for nearby genes which are difficult to manipulate by other means. However, new strategies for crop improvement might result, if one could isolate genes relevant to particular components of agricultural productivity. Because the biochemical function of such genes is usually unknown, the elegant strategies which exist for cloning genes of known function are of little help to the breeder. Further, because measures of agricultural productivity are influenced by many genes, together with environment, one can readily identify individuals in which only one of the many genes has been turned off, by insertion of an easily identified DNA element [26]. One piece of information the breeder can readily obtain is the chromosomal location, or map position of genes affecting agricultural productivity.
- Review Article
- Abstract
- Introduction
- Molecular Markers
- Genotype identification and genetic variability
- Mapping quantitative trait loci
- Manipulating traits controlled by a few major loci
- Manipulating traits controlled by many loci
- Understanding the genetic architecture of quantitative traits
- Improving Efficiency of Breeding Programs
- Marker facilitated Study of complex populations
- Marker facilitated Study of Polyploids
- Revealing Evolutionary Relationships among Crop Species and Their Wild Relatives
- Cloning gene from map position
- Conclusion and Challenges for the Future
- References
Conclusion and Challenges for the Future
Plant breeding is widely recognized as making substantial contribution to long term improvement of agricultural productivity. Genetic markers are considered to be a potentially valuable tool in accomplishing many plant breeding objectives. Marker-facilitated breeding is already showing its value in specialized breeding experiments, such as, introgression of traits from wild species, and may stimulate interest in this important(but often difficult) undertaking. The broader impacts of genetic markers on the overall process of crop improvement will likely to be influenced by many factors, such as:
A. The genetic basis of variation among elite varieties. A superior variety is often distinguished from its lesser brethren by tiny differences in phenotype. Such small differences might prove difficult to map, in that an enormous number of individuals would have to be studied. Further, these differences could be partly due to epistasis, which has proven elusive in mapping experiments. Carefully designed mapping experiments using a small number of elite crosses might offer valuable information on the genetic basis of variation among elite breeding lines.
B. The availability of diverse germplasm with known properties. Wild species and feral accessions of crop species hold much of the existing genetic variation in plant species. In some species, extensive germplasm collections have been accumulated, while other species have been less thoroughly collected. Further, much of the collected germplasm have been characterized only minimally, if at all. It was believed that DNA markers are an especially valuable tool for introgression of desirable traits from exotic germplasm in to elite cultivars. However, the germplasm must be available, and its attributes must be known. Thus, collection and characterization of exotic germplasm continues to be priority, and is made ever more urgent as human activities rapidly alter the global environment.
C. Identifying biophysical measures of agricultural productivity. Classical plant breeding methods rely heavily upon end point measurements of agricultural productivity, which are influenced by different parameters and consequently different genes in different environments. If more specific measures of agricultural productivity can be identified for example, physical or chemical properties of the plant which relate directly to productivity under a particular environmental stress, it will be much more feasible to identify the underlying genes by mapping.
D. The ability to identify polymorphic genetic markers in cultivated germplasm. While cross pollinated species such as, maize generally show an abundance of genetic variation, it has proven difficult to find polymorphism among cultivars of many self-pollinated species such as, wheat soybean, and tomato, impeding genetic mapping experiments. New classes of highly polymorphic genetic markers, techniques for detecting tiny genetic differences, and techniques for rapidly screening many candidate markers for polymorphism, may help in this regard.
E. Development of tools or techniques for high resolution analysis of large genomes. Much progress in plant breeding might be made by having markers to more easily select for traits of interest. However, this marker-facilitated selection represents only one facets of how molecular biology might contribute to plant breeding. Cloning of individual genes responsible for variation in agriculturally important traits would undoubtedly contribute to basic biology, and might also create new opportunities for improving agricultural productivity. Tools for physical analysis of large genomes, such as YACs, along with improved strategies for interfacing genetic and physical maps, will be a high priority.
Simplification of DNA marker technology Current procedures for using DNA markers in genetic mapping require considerable laboratory infrastructure, beyond the present capabilities of many breeding stations. Techniques are needed which require a minimum sophisticated equipment, and are easily scaled up to handle large numbers of individuals. New developments such as nonradioactive detection of DNA probes, allele specific DNA probes not dependent on restriction enzymes or elctrophoretic separations, and finally, amplification and direct visualization of particular DNA sequences are all promising steps toward simpler molecular marker technology.
- Molecular Markers
- Genotype identification and genetic variability
- Mapping quantitative trait loci
- Manipulating traits controlled by a few major loci
- Manipulating traits controlled by many loci
- Understanding the genetic architecture of quantitative traits
- Improving Efficiency of Breeding Programs
- Marker facilitated Study of complex populations
- Marker facilitated Study of Polyploids
- Revealing Evolutionary Relationships among Crop Species and Their Wild Relatives
- Cloning gene from map position
- Conclusion and Challenges for the Future
- References
References
- Mondini L, Noorani A, Pagnotta MA (2009) Assessing plant genetic diversity by molecular tools. Diversity 1(1): 19-35.
- Jiang GL (2013) Molecular markers and marker-assisted breeding in plants. In: Andersen SB, editor. Plant breeding from laboratories to fields. Rijeka: In Tech Pp. 45-83.
- Nadeem MA, Nawaz MA, Shahid MQ, Dogan Y, Comertpay G, et al. (2018) DNA molecular markers in plant breeding: current status and recent advancements in genomic selection and genome editing, Biotechnology & Biotechnological Equipment 32(2): 261-285.
- Jiang GL (2013) Molecular Markers and Marker-Assisted Breeding in Plants, Plant Breeding from Laboratories to Fields Sven Bode Andersen, Intech Open.
- Joshi M, Deshpande JD (2011) Polymerase chain reaction: methods, principles and application. Int J Biomed Res 2(1): 81-97.
- Melchinger AE (1993) Use of RFLP markers for analyses of genetic relationships among breeding materials and prediction of hybrid performance. Proc. of the First Int. Crop Science Congress.
- Smith OS, Smith JSC, Bowen SL, Tenborg RA, Walls SJ (1990) Similarities among a group of elite maize inbreds as measured by pedigree, F1 grain yield, grain yield heterosis, and RFLP's. Theoretical and Applied Genetics. 80(6): 833-840.
- Wang Y, Ghouri F, Shahid MQ, Naeem M, Baloch FS (2017) The genetic diversity and population structure of wild soybean evaluated by chloroplast and nuclear gene sequences. Biochem Syst Ecol 71: 170178.
- Smith OS, Smit JSC (1992) Measurement of genetic diversity among maize hybrids; a comparison of isozymic, RFLP, pedigree, and heterosis data. Maydica 37: 53-60.
- Bernardo R (1992) Relationship between single-cross performance and molecular marker hetero zygosity. Theoretical and Applied Genetics 83(5): 628-634.
- Dudley JW, Saghai Maroof MA, Rufener GK (1992) Molecular marker information and selection of parents in corn breeding programs. Crop Science 32: 301-304.
- Melchinger AE (2006) Use of Molecular Markers in Breeding for Oligogenic Disease Resistance. Plant Breeding 104(1): 1- 19.
- Tanksle SD, Rick CM (1980) Isozymic gene linkage map of the tomato: applications in genetics and breeding. Theoretical and Applied Genetics 58(2): 161-170.
- Lande R, Thompson R (1990) Efficiency of marker-assisted selection in the improvement of quantitative traits. Genetics 124: 743-756.
- Lande R (1992) Marker-assisted selection in relation to traditional methods of plant breeding. Pp. 437-451. In: H. R. Stalker and J. P. Murphy (eds.) Plant breeding in the 1990s. CAB International, Wallingford, UK.
- Beavis WD, Grant D, Albertsen M, Fincher R (1991) Quantitative trait loci for plant height in four maize populations and their associations with qualitative genetic loci. Theoretical and Applied Genetics 83(2): 141-145.
- Stuber CW, Moll RH, Goodman MM, Schaffer HE, Weir BS (1980) Allozyme frequency changes associated with selection for increased grain yield in maize (Zea mays L.). Genetics 95(1): 225-236.
- Lateef DD (2015) DNA Marker Technologies in Plants and Applications for Crop Improvements. Journal of Biosciences and Medicines 3(5): 7-18.
- Khalf-Allah AM, Abdel-Razzak HS, Feleafel MN, Harfoush EA (2014) Efficiency of Selection Breeding Programs to Improve Yield and Root Quality of Turnip. International Journal of Vegetable Science 20(4): 354-365.
- Mastretta-Yanes A, Acevedo Gasman F, Burgeff C, Cano Ramirez M, et al. (2018) An Initiative for the Study and Use of Genetic Diversity of Domesticated Plants and Their Wild Relatives. Front. Plant Sci 9: 209.
- Lande R (2017) The maintenance of genetic variability by mutation in a polygenic character with linked loci. Genet Res. 89(5-6): 373-87.
- Wu KK, Burnquist W, Sorrells ME, Tanksley SD, Tew TL, et al. (1991) The detection and estimation of linkage in polyploids using single dose restriction fragments 83(3): 294-300.
- Rick CM (1979) Bio systematic studies in Lycopersicon and closely related species of Solanum. In” The biology of solanaceae” (Hawkes JC, Lester RN, Skeldings AD, eds). Academic press, New York.
- Riley R (1965) Cytogenetics and evolution of wheat. In "Essays on crop plant evolution” (Hutchinson JB Ed.). Cambridge University Press, Cambridge, England.
- Dou Q, Wang C, Lei Y, Yu F, Li Y, et al. (2013) Genome analysis of seven species of Kengyilia (Triticeae: Poaceae) with FISH and GISH. Genome 56(11): 641-649.
- Feldmann KA, Marks MD, Christianos ML, Quatrano RS (1989) A dwarf mutant of Arabdopsis generated by T-DNA insertion mutagenesis. Science 243(4896): 1351-1354.






























