Forensic Tools and Techniques of Absolute Human Identification: Physical and Molecular Approach
SS Daga1, Umema Ahmed1 and RK Kumawat2*
1Vivekananda Global University, Jaipur, Rajasthan
2DNA Division, State Forensic Science Laboratory Rajasthan, Jaipur
Submission:July 05, 2021;Published:August 02, 2021
*Corresponding author:RK Kumawat, DNA Division, State Forensic Science Laboratory Rajasthan, Jaipur
How to cite this article:SS Daga, Umema A, RK Kumawat. Forensic Tools and Techniques of Absolute Human Identification: Physical and Molecular Approach. J Forensic Sci & Criminal Inves. 2021; 15(3): 555913 DOI:10.19080/JFSCI.2021.15.555913.
Abstract
Absolute human identification is the goal in most of the crimes or civil cases like murder, sexual assault, kidnapping, disputed paternity, establish kinship relatedness, emigration cases, robbery, identification of unknown and many more. In these cases, main aim to identify accused person, linking of victim and/or suspect with crime. Therefore, forensic tools and techniques of absolute human identification are the revolutionary role in crime investigation. There are two most commonly human identification approaches at physical and molecular level employed in crime investigation. The physical approach which is fingerprinting that is found on hands of individuals. Fingerprints of an individual are persisting unchanged throughout the life, and this makes the most infallible means of identification. The molecular approach is DNA fingerprinting, has become the most powerful technique for judiciary system to aid in the conviction of guilty as well as exoneration of the innocent. A DNA profile can tell the scientist if the DNA belongs to a man or woman, and if the sample being tested belongs to which individual. A DNA profile of an individual arises from father and mother both. There is 23 chromosomes occurs from father and 23 chromosomes occurs from mother which forms a child having 23 pair of chromosomes. Here we present potential value of both the approach for crime investigation and comparative study between these approaches.
Keywords: Human identification; Fingerprinting; DNA; Murder; Crime investigation
Introduction
Forensic science is a multi-disciplinary science which deals with application of different methods and techniques to aid criminal justice delivery system [1]. It is well known that forensic science, when properly applied and presented, can be critical to the outcome of a prosecution and exoneration. Due to speed of technological advancement and increasing globalization, crime is continuously evolving and adapting. New crime trends have emerged, with people committing crimes in cyberspace, crime against women and children, and abusing new psychoactive substances and drugs [2]. However, the technological advancement for detection of crime did not happen at the same pace at which crime has emerged. Therefore, new technology advancement is required to meet the ever-increasing challenges for solving crimes. Unless need based research is conducted; the forensic problems may not find solution and will persists. Absolute human identification is the goal in most of the crimes or civil cases like murder, sexual assault, kidnapping, disputed paternity, establish kinship relatedness, emigration cases, robbery, identification of unknown and many more [3,4]. In these cases, main aim to identify accused person, linking of victim and/or suspect with crime. Therefore, forensic tools and techniques of absolute human identification are the revolutionary role in crime investigation. There are two most commonly human identification approaches at physical [5] and molecular level employed in crime investigation [6].
Physical approach for absolute human identification
Fingerprints are the distinctive ridges appearing as corrugated lines on the tips of fingers and thumbs. The corrugation results due to rising of a portion of the upper layer of fingertip skin slightly above the normal level. Since the upper layer of skin is called epidermis, the finger ridges are also referred to as epidermal ridges. The depression between two ridges is called a furrow or a valley. The ridges and valleys form a complex, curved pattern on the fingertips. The pattern on each finger of a person is so unique that it is not repeated on another finger of the same person or on the fingers of any other person. This makes fingerprints the most infallible means of identification.
Fingerprint science works on the following basic principles:
i. No two persons and no two fingers of the same person have identical ridge design on the fingertips. Fingerprints are unique – more unique than the DNA, the genetic material in the human cells. Although identical twins have same DNA sequencing, they do not have identical fingerprints.
ii. The fingerprints remain unchanged throughout life. The ridge pattern begins to take shape during fetal stage and does not alter during a person’s lifespan. It is only after death that decomposition sets in, and the finger ridges are destroyed. The prints do not show any variation. In the growing age, the fingerprint pattern expands; because of aging, it may shrink. However, the basic design remains the same.
iii. Fingerprints can be classified for record keeping. When a person commits a crime and is arrested, he is fingerprinted by the police. The fingerprint record is then passed on to the nearest fingerprint bureau. There are about twenty-five state level fingerprint bureaus in India. Their functioning is coordinated by Central Fingerprint Bureau, Ministry of Home Affairs. Each bureau maintains a catalogue of fingerprints.
Fingerprint Patterns
In 1888, a British anthropologist by the name of Francis Galton established the first classification of fingerprints to hasten the retrieval process. In 1896, an English Police Official stationed in India, Sir Edward Richard Henry, revised the Galtonian system and devised a classification system based on the different patterns in the fingerprints of various individuals. The system was mainly intended for use in the identification and tracking of criminals, and its groupings are still the foundation of the fingerprint classification and storage that is employed today. Henry’s system is based on four distinct groups of patterns, with each group possessing the same basic characteristics and resemblances. Within each major group there exist sub-groups containing similar differences among patterns in that group. Henry ‘s four types of pattern groupings (arch, loop, whorl, composite) (Figure 1) and their interpretations are as follows:
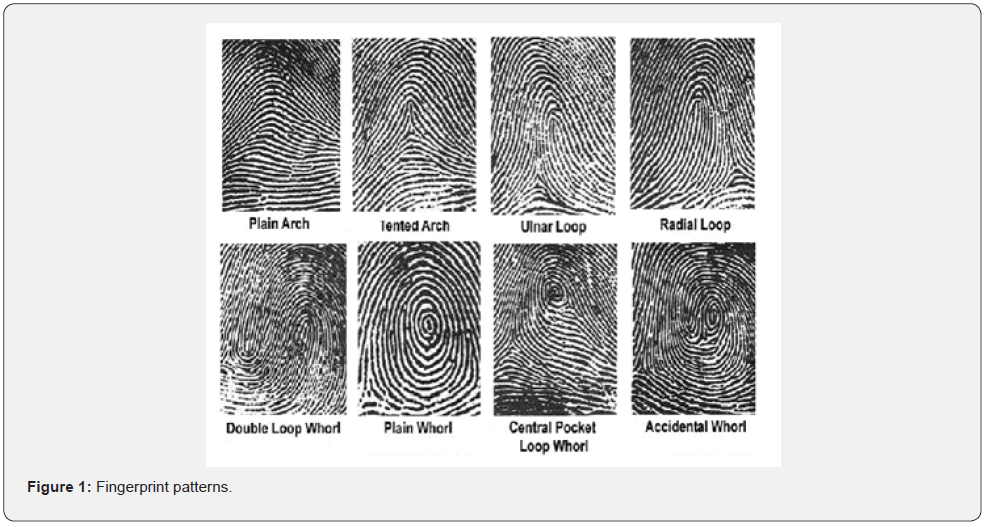

Arch
In arches, the ridges of the finger run continuous from one side of the finger to the other with no recurving. There are two sub-groups that further define the arch pattern:
Plain Arch: This pattern has a consistency of flow to it. It starts on one side of the finger, and then the ridge cascades upward slightly, almost resembling a wave out on the ocean. The plain arch then continues its journey along the finger to the other side. The plain arch is the simplest of the fingerprint patterns to discern.
Tented Arch: this pattern is like the plain arch in that it starts on one side of the finger and flows out in a similar pattern to the other side. However, the difference in the tented arch lies in the ridges in the center, which are not continuous as in the case of the plain arch. The ridges, which adjoin each other in the center, converge and thrust upward, giving the impression of a pitched tent.
Loop
In loops, the ridges make a backward turn but do not twist. This backward turn, or loop, is differentiated by how the loop flows on the hand and not how it flows on the card on which the imprint is taken. The imprint on the fingerprint card is like the reverse image we see when we look in the mirror at ourselves. There are two sub-groups that Henry identified in this category:
Radial Loop: These are looping that flow toward the radius bone of the hand or, in other words, when the downward slope of the loop is from the direction of the little finger toward the thumb of the hand.
Ulnar Loop: These are looping that flow toward the ulna bone of the hand or, in other words, when the downward slope of the loop is from the direction of the thumb toward the little finger of the hand.
Whorls
In whorls, there are patterns in which there are two or more deltas (first ridge nearest the divergence point of two type lines) and there exists a recurve preceding each delta. There are four sub-groups of whorls:
Plain Whorl: in these whorls, the ridges make a turn of one complete circuit and, therefore, are circular or spiral in shape. The plain whorl is the simplest form of whorl and the most common. There are at least two deltas and a ridge whose circuit may be spiral, oval, or circular in shape.
Central Pocket: in these whorls, one or more of the simple recurves of the plain whorl recurves a second time.
Double Loop: in these whorls, there are two separate loop formations. In each of these formations, there are two entirely separate and distinct sets of shoulders and deltas.
Accidental Whorl: in these whorls, the composition of the pattern is derived from two distinct types of patterns with at least two deltas. Whorls which contain ridges matching the characteristics of a particular whorl sub-grouping are classified as accidental whorls.
Composites
In composites, there are patterns found in fingerprints which are combinations of arch, loop and whorl. Henry subdivided the composites into four sub-groups:
Central Pocket Loop: these loops recurve a second time forming a pocket within the loop.
Twinned Loop: also referred to as the Double Loop, these loops consist of two separate loop formations.
Lateral Pockets Loop: These loops are like the Twinned Loop except that their ridges bend sharply down on one side before recurving, forming a pocket. The F.B.I. finds it too difficult to locate these two loops and classifies both kinds of as Double Loops.
Accidental Loops: these loops are a combination of any two types of patterns with the exception on the plain arch that basically has no pattern.
Today, different governing bodies may classify these patterns somewhat differently than Henry did more than a century ago. However, the original system still represents a body of work that has held up under critical review for well over a hundred years. On an average, there are about 80 ridge characters in a fingerprint. According to the Indian law, the location of eight characteristics is enough to individualize a person. This means that if the placement of eight characteristics in a fingerprint lifted from a crime scene, matches with the inked fingerprint of a person, then the latter is the accused. Thus, even if one-tenth of total characters on a fingertip can be visualized, the identity of the person can still be established.
Molecular approach for absolute human identification
DNA fingerprinting is the way to identify a certain person [1]. It has been around for at least twenty-five years and has two other names, Genetic fingerprinting, and DNA profiling [7]. Some different uses of DNA fingerprinting are, diagnosis of inherited disorders, developing cures for inherited disorders, biological evidence, and personal identification. The main basis of DNA technology is based on:
a) The genome of everyone is unique (except for twins) and is inherited from parents (both mother and father).
b) No two persons have the same DNA profile.
c) Probe subsets of genetic variation to differentiate between individuals (statistical probabilities of a random match are used).
d) DNA typing must be performed efficiently and reproducibly (information must hold up in court).
Markers are specific positions in the genome that are studied to find the genetic variation among individuals. Since the inception of forensic DNA technology, various markers have been used for individualization in the forensic investigation as well as genealogical studies. There are different techniques to DNA fingerprinting; two of which well-known DNA techniques are RFLP and STR as follows.
Restriction Fragment Length Polymorphism (RFLP)
Restriction enzymes have a characteristic property of cutting the DNA at specific base sequences, the restriction sites, and form the DNA fragments of variable length [8]. Agarose gel electrophoresis is used to separate the variable-length DNA fragments followed by southern blotting. The fragments get hybridized to short radio labelled probes and can be easily detected by radiography as bands. The loss or absence of such a few restriction sites forms the basis of individuality in humans. Thus, a unique RFLP signature for an individual can be generated. Forensic experts match such signatures originating from different sources with the one lifted from the crime scene [8].
For so long, the RFLP technology proved to be a reliable tool in DNA analysis, but it had few limitations which lead the forensic community to further exploring better markers to resolve the issues. The limitations posed were the need for a large quantity of non-degraded DNA, cost-effectiveness, the involvement of huge time length and labour, UV radiations, radioactive probes, and the requirement of much statistical work for analysis. These limitations were then overcome by PCR based markers [9-11].
Micro-Satellites or Short Tandem Repeats (STRs)
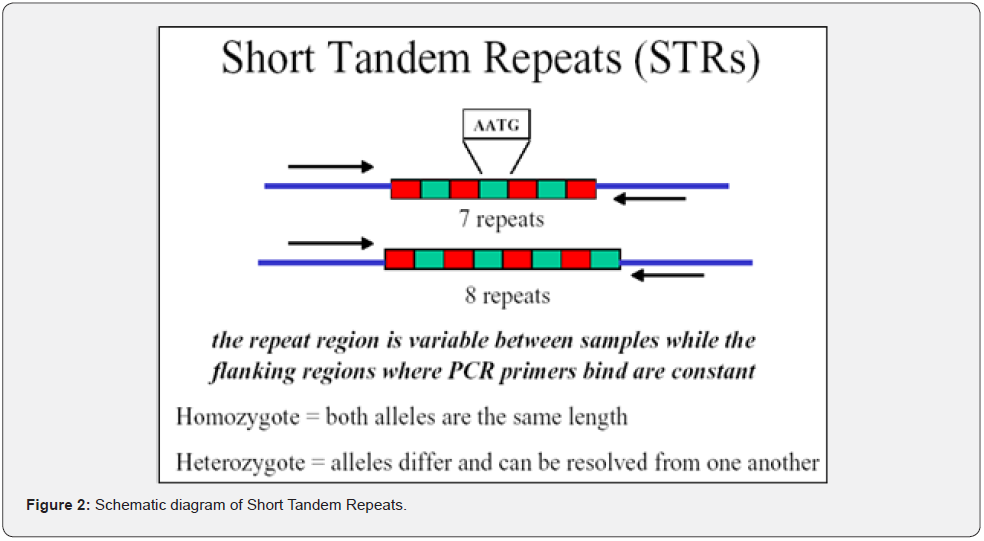
Short Tandem Repeats (STRs) are the specific regions of DNA, having 2-7 base pairs length repeating units of nucleotides (Figure 2) [12,13]. Because of the polymorphic and relatively small size of STR alleles (generally 100–350 nucleotides), amplification by polymerase chain reaction (PCR) is relatively easy, with the additional advantage of making the typing of highly degraded forensic samples possible [14]. The STRs of DNA is so highly polymorphic that is used as a potential tool for various genetic studies as well as in the forensic application for Justice. STRs are being used in population studies to generate allele frequencies for many different nationalities. This statement is since specific alleles will be consistent within a population or nationality. Hence, the study of STRs in any given population is a potential source of information about both population histories and evolutionary processes.
Properties of STR Markers
1) There are varied properties of STR loci that decide their suitability to several genetic applications.
2) The variations in the repeating units to make them highly efficient tools for human identification.
3) STRs show wide and uniform distribution throughout the genome.
4) They have compatibility for multiplex amplification because of their short sequence length.
5) The STRs are most useful for various molecular applications because it is easy to amplify them through Polymerase Chain Reaction (PCR).
6) They have high levels of relatively stable polymorphism.
7) STR alleles have lower mutation rates, which makes the assessment of STR allele transfer from parents to the child inferable, stable, and predictable.
8) The STR markers are the first choice for forensic applications due to their smaller size which also makes them suitable for the amplification of even highly degraded samples which is another requirement for forensic purposes.
Types of STR Markers
Different variants of STR markers are being used in the forensic investigation for various purposes. These variants are as follows:
Sex Determining Markers
The Amelogenin is the most common STR marker, which is used for sex determination in forensic investigation. This marker is found in Y and X chromosomes. From the initiation of STR based multiplexing to till date this marker is part of all the multiplex systems commercially available for forensic DNA casework analysis [15]. The gene Amelogenin produces several mRNAs that codes to generate a group of proteins. These proteins are responsible for tooth enamel formation. In humans, this gene is found in X as well as in Y chromosomes at p22.1-22.3 region and Yp11.2 region respectively. These genes are subsequently called as AMELX and AMELY gene. The mutation in the gene affects the enamel production (Amelogenesis imperfect). The Amelogenin protein is synthesized maximally by AMELX. The two genes are homologous (89%) and have highly conserved sequences between 2 - 6 exons. Thus, homogeneity and the presence of distinguishing features serve as reliable sex-determining markers. The AMELX gene has a 6bp deletion. Since females possess only X chromosome only a single fragment is observed whereas in males one fragments each of X and Y is observed due to this deletion. This forms the basis of determination and differentiation between male and female originated samples. The Amelogenin being a reliable marker for sex determination also have few drawbacks. Issues related to false-positive results due to allele dropout, Amel deletion, effects of PCR inhibitors, and failure to process degraded samples are being faced by experts daily [16-19].
Y- Chromosome STR markers
Human males exclusively have a haploid Y chromosome. This Y chromosome is extremely helpful in solving forensic cases for resolving the issues in male lineage and finding low male DNA in a mix of high female in sexual assault. The specific regions of the Y chromosome can be looked for the features which can differentiate them with female counterparts. Such a set of microsatellites found on the Y chromosome are collectively called as Y-STRs. Since these are male-specific, they help in the analysis of the mixed samples originating from both males and females, as in the case of rape, sexual assault, or otherwise in the case of a mass disaster. The Y STR typing in the above said cases can reveal the male component.
Autosomal STR Markers
Autosomal STR markers are the gold standard, highly stable, and widely used in forensic DNA application, genealogical studies, and medical research. Autosomal STR marker shows wide variation due to mutations, independent assortment, and recombination. Initially, four STR loci i.e., TH01, FES/FPS, VWA, and F13A1 were developed for forensic casework and termed as quadruples and known as a first-generation multiplex kit. The CTT multiplex kit was the first commercially available kit having TPOX, CSF1PO, and TH01 loci based on silver staining. It was extensively used in the United States. The US Federal Bureau of Investigation (FBI) soon formed an Index system called Combined DNA Index System more commonly referred to as CODIS which included the use of 13 STR loci for forensic investigation purposes. Lately to increase the usefulness of DNA fingerprinting technique seven ethnic group-specific markers were also added to the existing CODIS list.
The CODIS core loci and after extensive inter-laboratory exercise newly added STR locus to form CODIS 20 [20]. Most of the STR loci included in the CODIS system have tetra nucleotide repeats except D22S1045 which has a tri nucleotide repeat.
References
- D Primorac, MS Schanfield, D Primorac (2000) Application of forensic DNA testing in the legal system. Croat Med J 41 (2000) 32-46.
- AM Cutter (2006) To clear or to convict? The role of genomics in criminal justice, Genomics, Soc Policy 2: 1-15.
- PJ Newall, ML Richard, E Kafarowski, WJ Donnelly, GE Meloche, et al. (1996) Homicide case report: successful amplification and STR typing of bloodstains subjected to fingerprint treatment by cyanoacrylate fuming. Can Soc Forensic Sci J 29: 1-5.
- T Magalhães, RJ Dinis Oliveira, B Silva, F Corte Real, et al. (2015) Biological evidence management for DNA analysis in cases of sexual assault. Sci World J
- R Saferstein (2007) Criminalistics: An introduction to forensic science, Pearson Prentice Hall Upper Saddle River, NJ.
- JM Butler (2011) Forensic DNA typing: biology & technology behind STR markers, Academic Press.
- JM Butler (2011) Advanced topics in forensic DNA typing: methodology, Academic press.
- P Gill, AJ Jeffreys, DJ Werrett (1985) Forensic application of DNA fingerprints. Nature 318: 577.
- D Primorac, MS Schanfield, D Primorac (2000) Application of forensic DNA testing in the legal system. Croat Med J 41 32-46.
- L Roewer (2013) DNA fingerprinting in forensics: past, present, future. Investig Genet 4: 22.
- A Grover, PC Sharma (2016) Development and use of molecular markers: past and present. Crit Rev Biotechnol 36: 290-302.
- K Kasai, Y Nakamura, R White (1990) Amplification of a variable number of tandem repeats (VNTR) locus (pMCT118) by the polymerase chain reaction (PCR) and its application to forensic science. J Forensic Sci 35: 1196-1200.
- JM Butler (2005) Forensic DNA typing: biology, technology, and genetics of STR markers, Elsevier.
- J Koreth, JJ O LEARY, J OD McGEE (1996) Microsatellites and PCR genomic analysis. J Pathol 178: 239-248.
- JM Butler (2015) The future of forensic DNA analysis. Philos Trans R Soc B Biol Sci 370: 2014252.
- K Thangaraj, AGReddy, L Singh (2002) Is the amelogenin gene reliable for gender identification in forensic casework and prenatal diagnosis. Int J Legal Med 116: 121-123.
- RK Kumawat, P Shrivastava, D Shrivastava, GK Mathur, S Dixit (2020) Genomic blueprint of population of Rajasthan based on autosomal STR markers. Ann Hum Biol 1-6.
- RK Kumawat, P Shrivastava, D Shrivastava, GK Mathur, HR Dash, et al. (2020) Peopling of Rajasthan, India: Evaluating the gene flow from east and west. Gene Reports 100990.
- RK Kumawat, P Shrivastava, D Shrivastava, GK Mathur (2020) Molecular diversity of 23 Y-STR genetic markers in the population of Rajasthan, India, Meta Gene100694.
- DR Hares (2015) Selection and implementation of expanded CODIS core loci in the United States. Forensic Sci Int Genet 17: 33-34.






























