Role of DNA in Paternity Testing
Kareem Essam1*, Mona Hamza2 and Ayman Diab3
1B.Sc from the faculty biotechnology at MSA University, Egypt
2Consultant in forensic DNA, MD. Medical Microbiology and immunology, Kasr El Einy
3Ph.D. in Biotechnology from Cornell University, USA and Dean of the faculty of Biotechnology at MSA University, Egypt
Submission:June 21, 2020; Published:June 29, 2020
*Corresponding author:Kareem Essam, Researcher in the faculty of Biotechnology at MSA University, Egypt
How to cite this article:Kareem E, Mona H, Ayman D. Role of DNA in Paternity Testing. J Forensic Sci & Criminal Inves. 2020; 14(2): 555882. DOI: 10.19080/JFSCI.2020.14.555882.
Abstract
Egyptian society suffers from increasing in the number of divorce cases which leads to different problems. One of them is the children’s street that is a result of paternity issues. Paternity cases are caused due to huge problems between married couples because women may be entering a sexual partner in her life so the child will be not the son of her husband. This study discusses the techniques that are used in paternity cases in order to detect the biological father and prove which technique is more accurate compared to other techniques. Blood groups were the first approach that was used to detect whether the case is inclusion or exclusion. The most common and accurate technique is a short tandem repeat. This technique depends on 16 short tandem repeat loci founded on the chromosomes. It also depends on the length of each locus. This study discusses 6 samples concerned with 2 paternity cases while in one case concerned father, mother and two children.
Keywords:Paternity Test; Short tandem repeats; Deoxyribonucleic acid; Polymerase chain reaction; Genetic analyzer
Abbreviations: CPI: Combined Paternity Index; RI: Relationship Index; KI: Kinship Index; LR: Likelihood Ratio; PI: Paternity Index; CAPMAS: Central Agency for Public Mobilization and Statistics; STR: Short Tandem Repeat
Introduction
By the end of 2018, there were about 17,000 divorce cases in Egypt which marks a 13.4% rise in divorces. According to the Central Agency for Public Mobilization and Statistics (CAPMAS), the total number of divorces reached 199,867 in comparison to 180,244 in 2014. In Egypt, most women marry between the age of 20 and 25 while most men marry between the ages of 25 to 30. Men between 60 and 65 years and women over 65 years had the lowest marriage rate so that they had the lowest judicial problems between them like the paternity issues. Paternity issues always happen due to huge problems between married couples. It happens when married women enter a sexual partner in her life so the child will be not the son of her husband. There are also many cases that lead to paternity and also lead the father to deny his son or daughter and vice versa. The science has developed and played important role paternity problems by solving these problems through different techniques. The old one is ABO blood typing and the latest one is STR. (Aswatmasriya.com, 2017).The aim of this research is to identify the biological parents and to prove that the case is inclusion or exclusion by extracting the DNA sample from a blood sample or from a buccal swab in order to identify the 16 codes core STR loci with chromosomal positions. Egypt faces many paternity cases that cause several issues for children who grow up and didn’t know their biological parents. These cases have a bad effect on the children’s psychology that may lead them to fail in their future and it is also one of the main reasons for children’s streets. Some estimates say that two million children are living on Egypt’s streets. A quarter of street children are to be less than 12 years old, two-thirds between the age of 13 and 16, and just under 10 per believed cent over 17. They find themselves on the street for a variety of reasons, including family breakdowns due to divorce and remarriage. Some of these Egyptian children are victims for backward society and they are deprived of their rights in education, health, and social care and especially the right to family care so that the sciences present the STR polymorphism in paternity analysis to solve the paternity cases and return the right of these children in family care [1-5].
Modern DNA Typing Procedures
Nowadays, the most common technique that is used in paternity cases depends on the identification of 16 STR core loci with the chromosomal position. These loci are identified in each individual related to the paternity case. After identification, the DNA profile of the alleged father and mother is compared to the DNA profile of the child. The process steps of paternity testing include five steps. These five steps are illustrated in the sample collection, DNA Extraction, quantification, amplification, and STR analysis. In the sample collection step, the DNA is extracted from its biological source moreover it is measured in order to evaluate the quantity of DNA recovered in the quantitation step. In the amplification step, specific regions of the DNA are targeted and copied with polymerase chain reaction which is known as PCR. Finally, in the STR analysis step, the Commercial kits are commonly used to enable simultaneous PCR of 13 to 16 short tandem repeat (STR) markers. STR alleles are interpreted relative to PCR amplification artifacts following separation by size using capillary electrophoresis and data analysis software [6].
DNA Quantitation Using RT-PCR
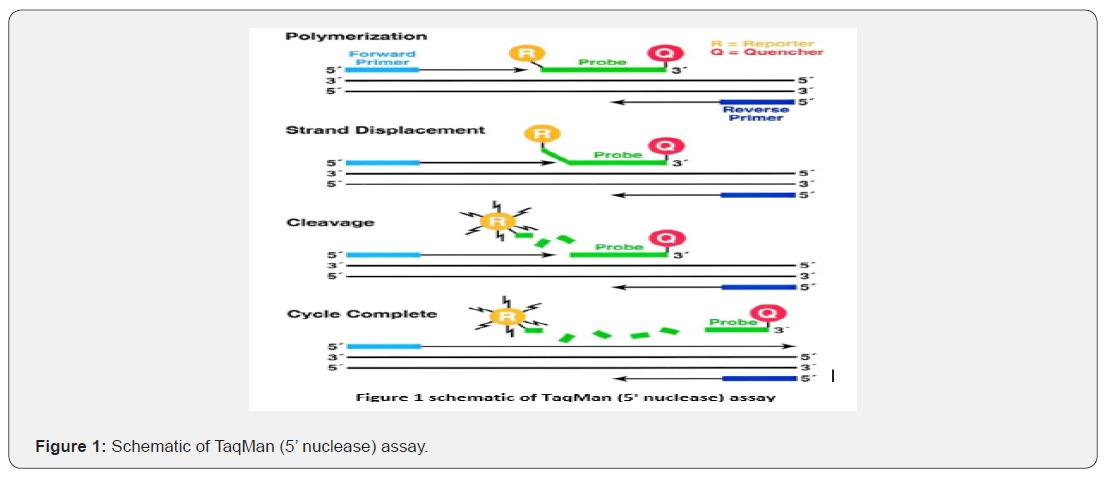
To ensure that DNA recovered from extraction is human rather than from another source such as bacteria, human-specific DNA quantitation is required. Only after DNA in a sample has been isolated can its quantity and quality be reliably assessed. Detection of the appropriate amount of DNA template to include in PCR amplification of short tandem repeat loci avoiding offscale data and associated artifacts are the main purpose of DNA quantitation in paternity casework [7-10]. PCR amplification of too much DNA results in overblown electropherograms that make interpretation of results more challenging and time consuming to review. Too little DNA can result in loss of alleles due to randomly amplification and failure to equally sample the STR alleles present in the sample. A number of DNA quantitation tests have been used over the years to estimate the amount of total DNA or human DNA present in a sample. Several DNA quantitation tests are used in many approaches such as yield gels, Pico Green, end-point PCR, real-time quantitative PCR, UV absorbance, and slot blot. UV absorbance is the most common technique to determine DNA yield and purity. It could be argued that fluorescence measurement is easier. Absorbance measurement is simple, moreover, it requires commonly available laboratory equipment. All that is needed for the absorbance method is a spectrophotometer equipped with a UV lamp, UV-transparent cuvettes depending on the instrument, and a solution of purified DNA. Absorbance readings are performed at 260nm (A260) where DNA absorbs light most strongly, and the number generated allows one to estimate the concentration of the solution. To ensure the numbers are useful, the A260 reading should be within the instrument’s linear range (generally 0.1-1.0). DNA concentration is estimated by measuring the absorbance at 260nm, adjusting the A26 measurement for turbidity (measured by absorbance at 320nm), multiplying by the dilution factor, and using the relationship that an A260 of 1.0 = 50μg/ml pure dsDNA. Real-time PCR is known as quantitative PCR because it analyzes the cycle to cycle change in fluorescence signal resulting from the amplification of a target sequence during PCR. This analysis is performed without opening the PCR tube and therefore can be referred as a homogeneous detection assay. There are two common approaches that are used in DNA quantitative either the fluorogenic 5’ nuclease assay known as TaqMan or intercalating dye such as SYBER Green. Quantifying the DNA in a sample is used to detect the amount of DNA for adequate amplification. The smallest volume required for the reaction ranges for, 0.5 to 1.0 ng Figure 1.
PCR Amplification
PCR is an enzymatic process. It is a specific region of DNA. It is replicated over and over again to yield many copies of a particular sequence. A copy of the target DNA sequence is generated for every molecule containing the target sequence during each cycle. The amplified product is defined by oligonucleotide primers. These oligonucleotide primers are complementary to the 39- ends of the sequence of interest. In the ideal reaction with 100% amplification efficiency, approximately a billion copies of the target region on the DNA template have been generated after 32 cycles. Lower quantities of PCR products are produced due to a reduction in amplification efficiency through PCR inhibition or poor primer annealing [11]. The polymerase chain reaction (PCR) technique has become the standard process for DNA paternity testing. PCR technology allows scientists to amplify a very small quantity of DNA in order to increase the amount of DNA up to billions of copies of the same DNA that are made for testing and analysis. The PCR machine also makes this process happen fairly quickly. Using PCR technology, DNA relationship tests can be performed much more easily and quickly. Buccal swab specimens are collected from each tested party in a non-invasive manner in a standard DNA paternity test today, to make it ideal even for newborn babies. As the quantity of DNA required in the PCR based test is so small, genetic scientists can even test Amniotic fluid that contains the embryo’s DNA to determine paternity before the child is born (Figure 2) [6]. Thermal cycling typically entails 3 different temperatures that are repeated over and over again 25 to 35 times. There are three phases according to three different temperatures. Denaturation occurs at 95 C to separate double-strand DNA. Annealing occurs at about 60 C so the primers can anneal with their complementary sequence in the target DNA sequence. The temperature rises to 72 C in the extension phase, Because Taq polymerase functions optimally at this temp. and begins polymerization, adding nucleotides at 3’end of each primer attached to a DNA strand. PCR amplification has several advantages for biological evidence. It can use very small amounts of DNA template as little as a single cell [12-16]. DNA degraded to fragments only a few hundred base pairs in length can serve as an effective template for amplification and also large numbers of copies of specific DNA sequences can be amplified simultaneously with multiplex PCR reactions. It doesn’t amplify contaminated DNA such as fungal and bacterial sources due to the presence of human-specific primers. Commercial kits are now available for easy PCR reaction setup and amplification. However, three potential pitfalls are considered disadvantages of PCR. The presence of PCR inhibitors in the extracted DNA may not amplify the target DNA template. The sequences changes in the primer- binding region of the genomic DNA template may fail the amplification. Contamination from other human DNA sources besides the forensic evidence at hand or previously amplified DNA samples is possible without careful laboratory technique and validated protocols. PCR amplification has several precautions to prevent dangerous something to happen.

In order to prevent cross-contamination during liquid transfers, Aerosol-resistant pipette tips should be used and changed on every new sample. Reactions may also be set up in a laminar flow hood to prevent contamination. Disposable gloves should be worn and changed frequently. Equipment, such as pipettes, and reagents for sitting up PCR should be kept separate from other laboratory supplies, especially those used for the analysis of PCR products and also Pre- and post-sample processing areas should be physically separated. Usually, a separate room or containment cabinet is used for setting up the PCR amplification reactions. Importance of STRs in forensic DNA typing. DNA markers for human identification purposes are essential because these DNA markers exhibit the highest possible variation or a number of less polymorphic markers that can be combined in order to obtain the ability to distinguish between samples STR Project beginning in April 1996 and concluding in November 1997 involved 22 DNA typing laboratories and the evaluation of just 16 candidate STR loci: D8S1179, D21S11, D7S820, CSF1PO, D3S1358, THO1, D13S317, D16S539, D2S1338, D19S433, VWA, TPOX, D18S51, Amel, D5S818 and FGA [6]. These STR loci are located on the chromosomes [17-21].
Paternity Index
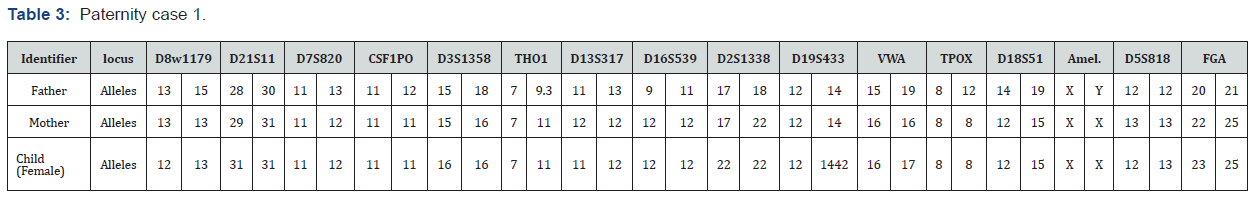
The paternity index (PI) compares between the possibilities that a genetic marker (allele) and that the alleged father (AF) passed to the child with the probability that a randomly selected unrelated man of similar ethnic background could pass the allele to the child. This is presented in the formula X/Y, where X is the chance that the alleged father could transmit the obligate allele and Y is the chance that some other man of the same race could behave transmitted the allele. X is assigned the value of 1 if the AF is homozygous for the allele of interest and 0.5 if the alleged father is heterozygous. Ratio of the two probabilities. This ratio discusses how much better the data (genotypes) fits with the hypothesis that the AF is the real father. The larger ratio is more evidence that this man is the real father. Two possible results are identified in this study. Inclusion, test man, or alleged father could be this child’s father. In the exclusion case, there is no way that the AF could be this child’s father when multiple genetic systems are tested, a PI is calculated for each system. The genetic system is inherited independently [22-25]. the product of system PIs is the combined paternity index (CPI) the combined paternity index is also determined by the multiplying the individual PIs for each locus tested. This value is referred to as system PI. Likelihood ratio (LR) is called the relationship index (RI) or kinship index (KI). The main functions of the likelihood ratio are to describes how strongly genotypes support one relationship versus the other relationship and also to express the likelihood of obtaining the DNA profiles under two mutually exclusive hypotheses Figure 3-5. Each independent locus tested produces its own relationship index, which can be multiplied by those of other independent loci to calculate a combined relationship index (CRI).
a. CRI > 1 supports the numerator (claimed relationship)
b. CRI < 1 supports the denominator (alternative relationship)
c. Larger CRI values provide more support for the claimed relationship


Materials and Methods
This study discusses two protocols in extracting DNA. Each one is used according to the sample collection and also according to the available kits that. These two protocols are extraction of DNA from fresh blood using thermos scientific kit and extraction of DNA from buccal swab spin. After extracting protocols, the DNA amplification process is done in order to identify the 16 codes core STR loci and the length of each locus is determined by electrophoresis step which is done by using the 3500 genetic analyzers to be able to compare the DNA profiling of each individual founded in the case and prove that whether the case is inclusion or exclusion. This methodology was applied to three paternity cases. Each case consists of three individual which means that the result of 3 paternity cases would be 9 DNA profiling [27-30].
Extraction of DNA from fresh blood using thermos scientific kit
200 μl of whole blood, 400 μl lysis solution, and 20 μl proteinase k were put in a 1.5 ml tube and then the tube was put in the vortex to mix well. The sample was incubated for 10 min at 56 °C in a shaking water bath. 200 μl of 96% ethanol was added and mixed well by vortexing spin. 820 μl of the lysate was transferred to a spin column and then the sample was centrifuged for 1 min at 8,000 rpm. A collecting tube containing the flowthrough solution was discarded and a new collecting 2 ml tube was placed. 500 μl of wash I was added centrifuged for 1 min at 10,000 rpm and the collecting tube containing the flow-through solution was discarded and placed a new collecting 2 ml tube. 500 μl of wash II was added and centrifuged for 3 min at max speed (14,000 rpm). The collecting tube was emptied and centrifuged at max speed for 1min. the collecting tube was discarded containing the flow-through solution and the spin column was transferred to a sterile 1.5 microcentrifuge tube. 170 μl of Elution buffer was added to the center of the spin column membrane to elute genomic DNA. The sample was incubated for 2 min at room temperature and centrifuged for 1 min at 10.000 rpm. The spin-column was discarded and finally, the purified DNA could be used immediately or stored at -20 °C [31-36].
Extraction of DNA from buccal swab spin
The buccal swab was placed in a 2 ml microcentrifuge tube. 400 μl of cotton and DACRON swab was added to the sample. 20 μl of QIAGEN protease stock solution and 400 μl cotton DACRON swab buffer AL were added to the sample and then mixed immediately by vertexing for 15 sec. The sample was incubated at 56 °C for 10 min.400 μl of cotton or DACRON swab was added to the sample, mixed again by vortexing, and then centrifuged briefly to remove drops from inside the lid. 700 μl of the mixture was applied to the spin column in a 2 ml collection tube without wetting the rim, the clap was cap and centrifuge at 6000 x g (8000 rpm) for 1 min. The spin column was placed in a clean 2 ml collection tube and the tube containing the filtrate was discarded. The previous step was repeated by applying up to 700 μl of the remaining mixture from step four to the spin column. The spincolumn was opened carefully and 500 μl buffer AW1 was added without wetting the rim. The cap was closed and centrifuged at 6000 x g (8000 rpm) for 1 min. the spin column was placed in a clean 2 ml collection tube. The spin-column was opened and 500 μl of buffer AW2 was added without witting the rim. The cap was closed and centrifuged at full speed (20,000 xg; 14,000 rpm) for 3 min. The spin column was placed in a new 2 ml collection tube and the collection tube with the filtrate was discarded and centrifuged at 20,000 xg (14,000 rpm) for 1 min. The spin column was placed in a clean 1.5 microcentrifuge tube and the collection tube containing the filtrate was discarded. The spin-column was opened, and 150 μl buffer AE was added. The sample was incubated at room temperature for 1 min and then centrifuged at 6000 xg (8000 rpm) for 1 min [37-40].
DNA amplification using Amp FISHER® Identifiler ® Plus Kit
10μl of Amp FISTR® Identifiler ® Plus Master mix was added to the tube and then 5 μl of Amp FISTR® Identifier ® Plus Primer Set was added per each reaction. The Am pFISTR® Identifiler ® Plus Kit Master Mix and the Amp FISTR® Identifiler ® Plus Kit Primer Set 11was thawed, then vortexed 3 seconds and centrifuged briefly before opening the tubes. The Amp FISTR® Identifier ® Plus Kit Master Mix and the Amp FISTR® Identifier ® Plus Kit Primer Set was thawed, then vortexed 3 seconds and centrifuged briefly before opening the tubes. The required volumes of components were pipetted into an appropriately sized polypropylene tube. The reaction mix was vortexed for 3 seconds and then centrifuged briefly. 15 μl of the reaction Mix was dispensed into each reaction well of a micro Amp® Optical 96- Well Reaction Plate or each Micro Amp® tube [41].
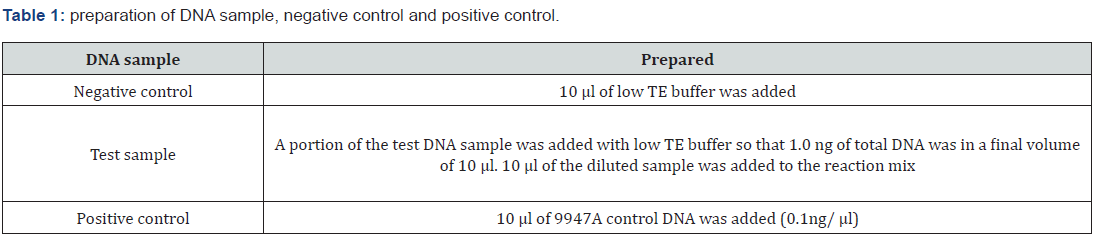
Prepared the DNA Samples
The final reaction column (sample or control plus reaction mix) is 25 μl. The MicroAmp® Optical 96-Well Reaction Plate with Micro Amp® Clear adhesive Film was sealed. The reaction mix was vortexed for 3 seconds, then centrifuge the tubes at 3000 rpm for about 20 seconds in a (Table 1-4) top centrifuge to remove bubbles. The samples were amplified in a Gene Amp® in PCR system 9700 with the gold-plated silver 96-well block. The plate was loaded into the thermal cycler and closed the heated cover and then Program was started to run One completion of the run, the amplified DNA was stored and protected from the light Figure 6.



Prepared samples for electrophoresis on the 3500 genetic analyzers
The volume of Hi-Di Formamide and GeneScan500Liz size standard was calculated to prepare the samples according to 0.3 μl Gene Scan 500 LIZ standard and 8.7 μl Hi-Di formamide per reaction. The required volumes of components were pipetted into an appropriated sized polypropylene tube. The tube was vortexed, then centrifuged briefly. 9 μl of the Hi-Di-Formamide and 1 μl of PCR product or allelic ladder were added into each well of a Micro Amp® Optical 96well Reaction plate. The reaction plate was sealed with appropriate septa, then checked the plate to ensure that the contents of each well are collected the bottom, and then. The reaction plat was heated in thermal cycler for 3 minutes at 95 °C. The plate was placed immediately on ice for 3 minutes. The plate was a prepared assembly on the autosampler. Electrophoresis was started to run. Analysis of injected samples was done according to gene mapper software analysis and electropherograms are plotted and compared [42-45].
Results
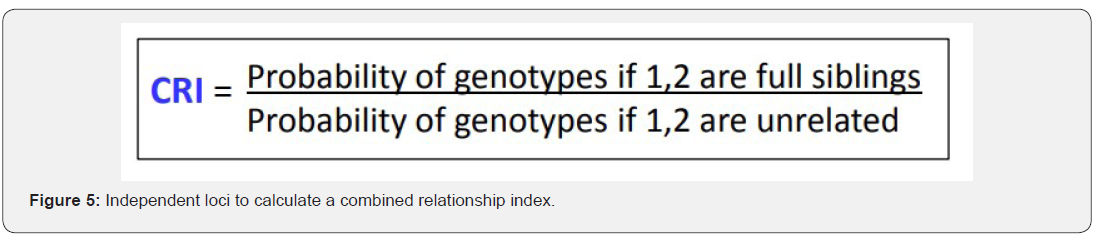
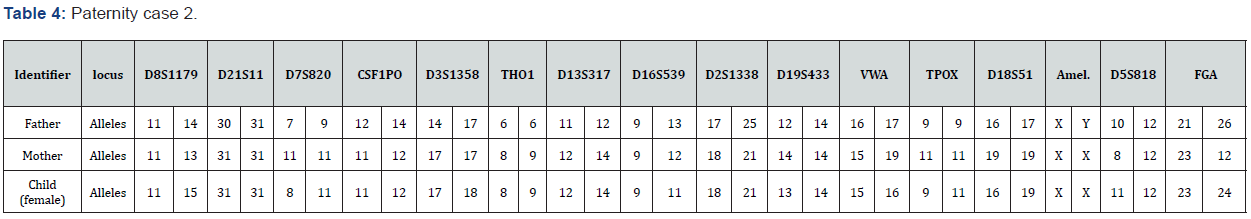
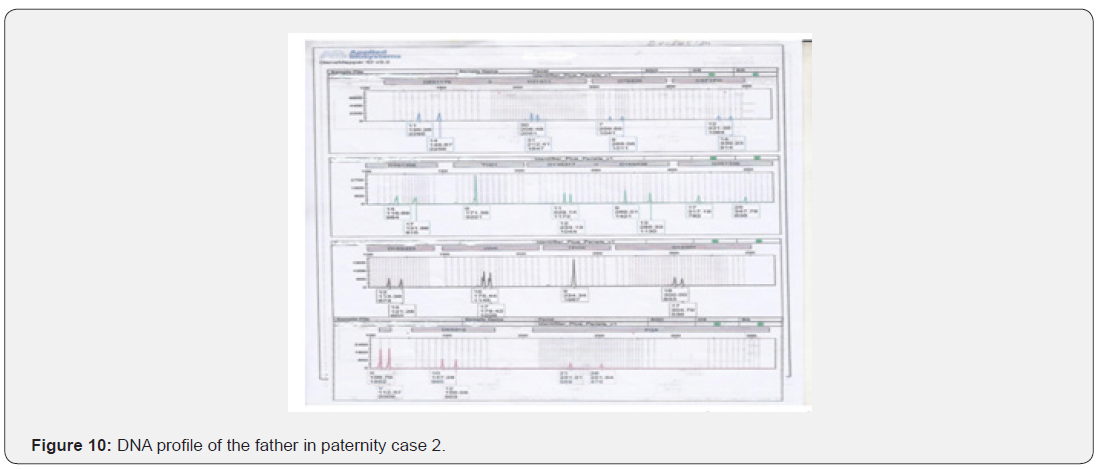
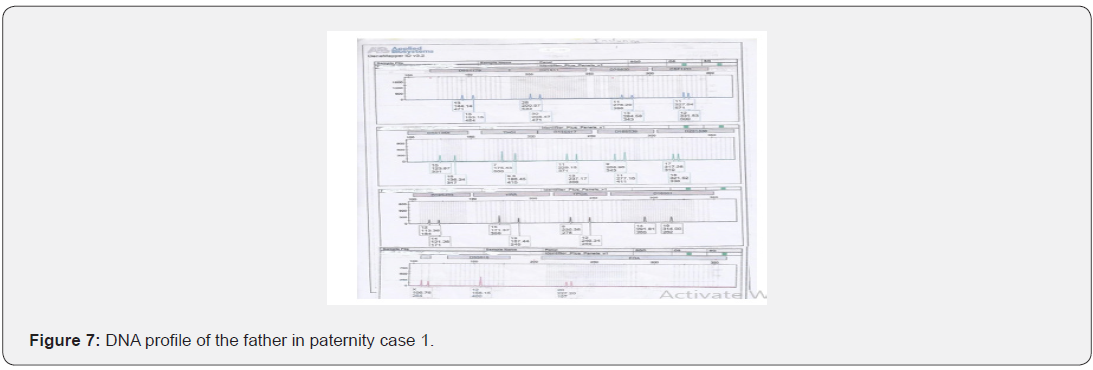
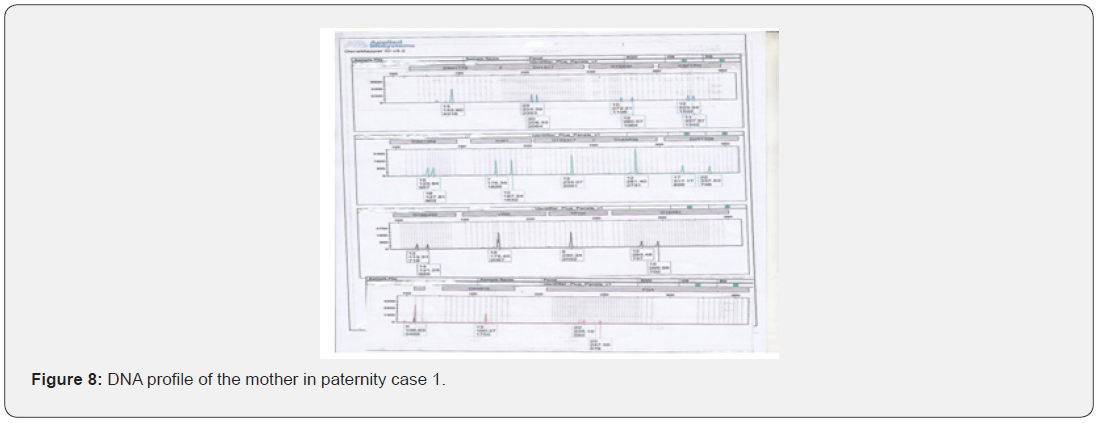
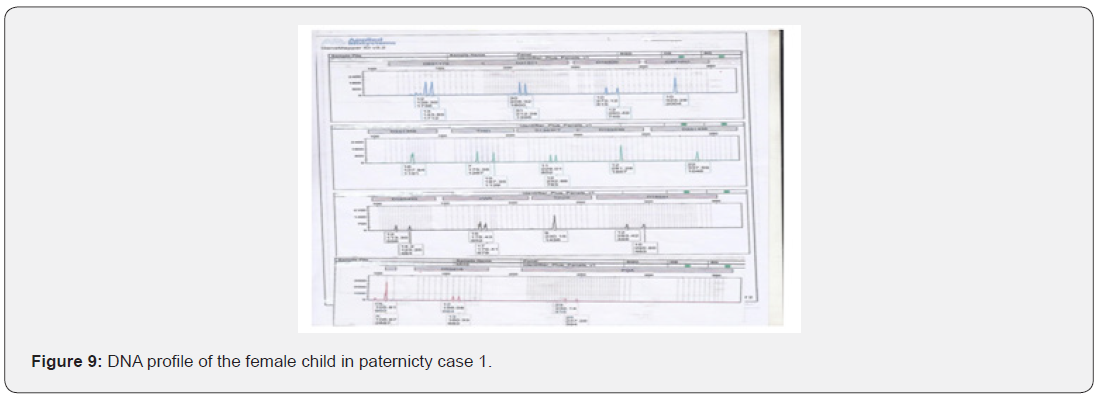
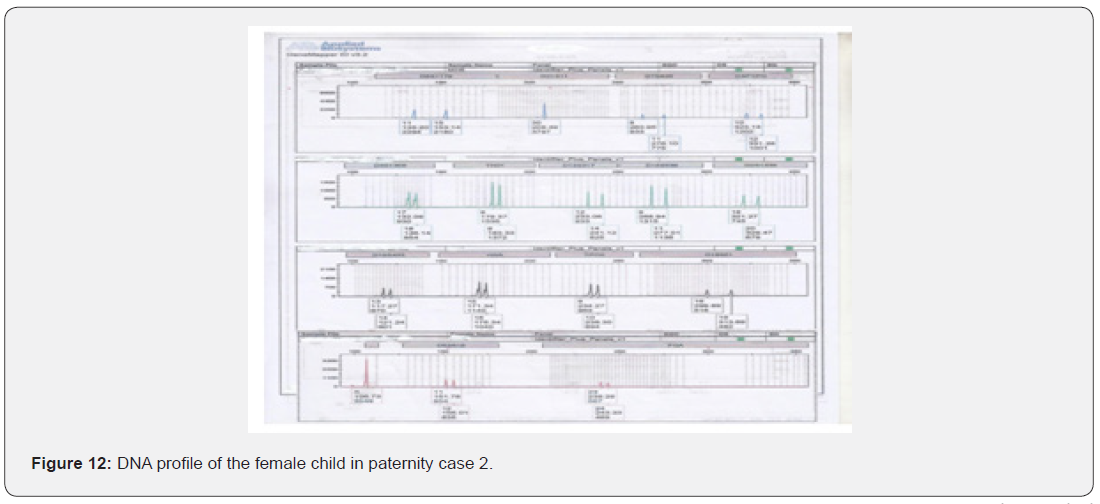
The total number of 6 samples concerned 2 paternity cases. Major cases included mother, father and child while in a single case At D8S1179, D7S820 and CSF1PO locus, the alleles of female child are 11: 15, 8:11and 10:12 respectively while the alleles on the father of the same locus are 11:14, 7:9 and 12:14 respectively and the mother 11:13, 10:11 and 11:12. This indicates that the alleles 15, 8 and 24 are not shared from this father Figure 7-13.


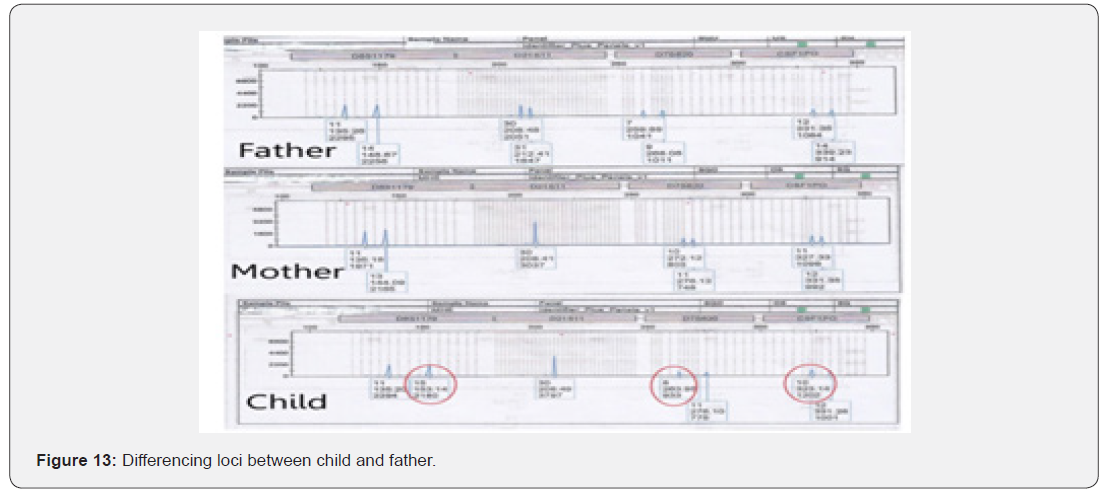
Discussion
In order to identify that the case is whether inclusion or exclusion, profiling DNA of father, mother and child have to be compared with each other according to the length of short tandem repeats of each locus. There are 16 candidate STR loci which are D8S1179, D21S11, D7S820, CSF1PO, D3S1358, THO1, D13S317, D16S539, D2S1338, D19S433, vWA, TPOX, D18S51, Amel, D5S818 and FGA. In inclusion cases, the child shares the length of each STR loci with his parents because each biological parent shares 23 chromosomes for their child. However, in exclusion cases, the child’s length of STR loci differs between the father and mother. In case 1 there is no difference between the length of short tandem repeats in the child comparing to the short tandem repeats in his mother and father because the child shares its loci with his father and mother while in case 2, At D8S1179, D7S820 and CSF1PO locus, the alleles of the female child are 11: 15, 8:11 and 10:12 respectively while the alleles on the father of the same locus are 11:14, 7:9 and 12:14 respectively and the mother 11:13, 10:11 and 11:12. This indicates that the alleles 15, 8, and 24 are not shared by this father. So DNA paternity testing is the more accurate than blood groups testing because If DNA patterns between mother, child, and the alleged father match on every DNA probe, the likelihood of paternity is 99.9 percent, while if the child and the alleged father do not match on two or more DNA probes, so the alleged father can be totally excluded. To handle DNA testing, either a blood or buccal swab, this provides a DNA sample for testing. Children can be tested at any age [46,47]. Paternity testing can even be done on an umbilical cord blood specimen at birth. The accuracy of testing performed on cheek cells utilizing the Buccal Swab is the same as an actual blood sample because DNA is the same in every cell of the human body.
References
- Bär, W (1994) DNA recommendations -1994 report concerning further recommendations of the DNA 160(2-3): 90-101.
- Bär, W (1997) DNA recommendations -further report of the DNA Commission of the ISFH regarding the use of short tandem repeat systems. International Journal of Legal Medicine 110: 175-176.
- Benecke M (1997) DNA typing in forensic medicine and in criminal investigationsa current survey 188: 181-188.
- Burger MF (2005) Buccal DNA samples for DNA typing: New collection and processing methods. BioTechniques 39: 257-261.
- Burgoyne LA (1996) Solid medium and method for DNA storage. US Patent 5(496): 562.
- Butler JM (2008) Addressing Y-chromosome short tandem repeat (Y-STR) allele nomenclature. Journal of Genetic Genealogy, 4(2): 125-148.
- Del Rio SA (1996) Reusing the same blood-stained punch for sequential DNA amplifications and typing. Biotechniques, 20, 970-974.
- Ellegren H (2004) Microsatellites: simple sequences with complex evolution. Nature Reviews Genetics 5: 435-445.
- Frankell M (1989) Ethics and the forensic sciences: Professional automony in the criminal justice system. Journal of Forensic Sciences 34(3): 763-771.
- Gemayel R (2010) Variable tandem repeats accelerate evolution of coding and regulatory sequences. Annual Review of Genetics 44: 445-477.
- (2013) Evaluation of the-uitability for purpose‖ of four filter papers as archival media. Analytical Chemistry 74: 1863-1869.
- Gill P (1994) Report of the European DNA profiling group (EDNAP)-towards standardisation of short tandem repeat (STR) loci. Forensic Science International 65: 51-59.
- Gill P (1997) Considerations from the European DNA profiling group (EDNAP) concerning STR nomenclature. Forensic Science International 87: 185-192.
- Gill P (1997) Development of guidelines to designate alleles using an STR multiplex system. Forensic Science International 89: 185-197.
- Gill P (2001) DNA Commission of the International Society of Forensic Genetics: Recommendations on forensic analysis using Y-chromosome STRs. Forensic Science International 124: 5-10.
- Gusmão L (2006) DNA Commission of the International Society of Forensic Genetics (ISFG): an update of the recommendations on the use of Y-STRs in forensic analysis. Forensic Science International 157: 187-197.
- Hollien H (1990) The expert witness: Ethics and responsibilities. Journal of Forensic Sciences 35: 1414-1423.
- Jin L (1994) The exact numbers of possible microsatellite motifs. American Journal of Human Genetics 55: 582-583.
- Kates J, Guttenplan H (1983) Ethical considerations in forensic science services. Journal of Forensic Sciences 28(4): 972-976.
- Kayser M (2004) A comprehensive survey of human Y-chromosomal microsatellites. American Journal of Human Genetics 74: 1183-1197.
- Lögdberg L, Reid ME, Lamont RE, Zelinski T (2005) Human blood group genes 2004: Chromosomal locations and cloning strategies. Transfus Med Rev 19: 45-57.
- Lögdberg L, Reid ME, Zelinski T (2011) Human blood group genes 2010: Chromosomal locations and cloning strategies revisited. Transfus Med Rev 25: 36-46.
- Lucas D (1989) The ethical responsibilities of the forensic scientist: Exploring the limits. Journal of Forensic Sciences 34(3): 719-729.
- Misner LM (2009) The correlation between skeletal weathering and DNA quality and quantity. Journal of Forensic Sciences 54: 822-828.
- Mulero JJ (2006) Letter to the Editor-Nomenclature and allele repeat structure update for the Y-STR locus GATA-H4. Journal of Forensic Sciences 51: 694.
- Paabo S (1990) DNA damage promotes jumping between templates during enzymatic amplification. Journal of Biological Chemistry 265: 4718-4721.
- Pfeifer GP (2005) Mutations induced by ultraviolet light. Mutation Research 571: 19-31.
- Pang BC, Cheung BK (2007) Double swab technique for collecting touched evidence. Legal Medicine (Tokyo) 9: 181-184.
- Rath G, Mitra R, Mishra N (2014) Blood groups systems. Indian Journal of Anaesthesia, 58(5): 524
- Ravanat JL (2001) Direct and indirect effects of UV radiation on DNA and its components. Journal of Photochemistry & Photobiology B 63: 88-102.
- Romero RL (1997) The applicability of formalin-fixed and formalin fixed paraffin embedded tissues in forensic DNA analysis. Journal of Forensic Sciences 42: 708-714.
- Shapiro R, Danzig M (1972) Acidic hydrolysis of deoxycytidine and deoxyuridine derivatives: the general mechanism of deoxyribonucleoside hydrolysis. Biochemistry 11: 23-29.
- Shikama K (1985) Effect of freezing and thawing on the stability of double helix of DNA. Nature 207: 529-530.
- Stray JE, Shewale JG (2010) Extraction of DNA from human remains. Forensic Science Review 22: 177-185.
- Sweet D (1997) An improved method to recover saliva from human skin: The double swab technique. Journal of Forensic Sciences 42: 320-322.
- Urquhart A (1994) Variation in short tandem repeat sequences a survey of twelve microsatellite loci for use as forensic identification markers. International Journal of Legal Medicine 107: 13-20.
- Von Dungern E, Hirschfeld L (1910) Über Vererbung gruppenspezifischer Strukturen des Blutes. Z Immunitätsforsch Exp Ther. 6: 284-293.
- Walsh PS (1991) Chelex 100 as a medium for simple extraction of DNA for PCR-based typing from forensic material. Bio Techniques 10: 506-513.
- Wassenegger M (2001) Advantages and Disadvantages of Using PCR Techniques to Characterize Transgenic Plants 17: 73-74.
- Westhoff CM (2004) The Rh blood group system in review: A new face for the next decade. Transfusion 44: 1663-1673.
- Yamamoto F, Clausen H, White T, Marken J, Hakamori S (1990) Molecular genetic basis of the histoblood group system. Nature 345: 229-233.
- Yamamoto F, Mc Neill PD, Hakamori S (1995) Genomic organization of the human histo-blood group ABO genes. Glycobiology 5: 51-58.
- Yoon JH (2000) The DNA damage spectrum produced by simulated sunlight. Journal of Molecular Biology 299: 681-693.
- Bugert P, Rink G, Kemp K, Klüter H (2012) Blood Group ABO Genotyping in Paternity Testing 39(3): 182-186.
- Butler JM (2005) Forensic DNA Typing: Biology, Technology, and Genetics of STR Markers. (2nd), Elsevier Academic Press, New York, USA.
- Dean L (2005) Blood Groups and Red Cell Antigens. (1st ), National Library of Medicine.
- Scaradavou A (2013) Transfusion Medicine and Hemostasis. (2nd ), Elsevier Science.






























