Evaluation of Touched Documents Papers for Successful STR Typing Using Powerplex® 16 HS Amplification Kit
Ahmed Ch Kassab*, Rahma RashdanAlmutiri, Ahmed Saad Gouhar, and Saranya Ramesh Babu
Department of Forensic Sciences, Naif Arab University for Security Sciences, Saudi Arabia
Submission:March 28, 2020; Published:April 15, 2020
*Corresponding author:Ahmed Ch Kassab, Chairman of Forensic Sciences Department, College of Criminal Justice, Naif Arab University for Security Sciences, Saudi Arabia
How to cite this article:Ahmed C K, Rahma R A, Ahmed S G, Saranya R B. Evaluation of Touched Documents Papers for Successful STR Typing Using Powerplex® 16 HS Amplification Kit. J Forensic Sci & Criminal Inves. 2020; 13(5): 555875. DOI: 10.19080/JFSCI.2020.13.555875.
Abstract
Touch DNA is a recent concept in DNA typing provides an alternative DNA sources for DNA typing in forensic application. Touched documents, has become an investigative tool in forensics. This study aimed to evaluate three types of touched document paper, i.e. A4 copy, Coated and Chart papers, as a source of DNA for STR genotyping using Power Plex® 16 HS Amplification Kit. Skin cells from the touched papers were collected by adhesive tape collection method. The DNA extraction from the tapes were extracted following Chelex® extraction protocol, and quantities were estimated using real time PCR and, amplification of the STR loci was done using Power Plex® 16 HS Amplification kit. The amounts of DNA collected from all touched papers were low and classify as low copy number (LCN) (<100 pg.), and in most of the cases it was not quantifiable. The success rate in obtaining STR profile for above 10 markers from a folded and sketched area ofA4 copy paper was 60% and 40% and from coated paper were 20% and 10% respectively. While for chart paper the success rate for folded and sketched area was 60% and null. There was no effect of gender on the quantity of DNA extracted from various types of papers used in this study. However, a sex-related difference was observed in the STR profiles from some types of papers. This study support in part the use of touched document as a source of DNA in means of Touch DNA used to identify or relate a person to touched document.
Keywords:Touch DNA; Forensic sciences; DNA typing; Skin surface cell; Tape lifting; Document papers
Introduction
Forensic sciences evaluate all the physical evidence lifted in the crime scene depending on Locards exchange principle that theorizes an exchange occurs between two objects when they come into contact with others [1]. DNA typing which was first described 1985 by English genetics named Alec Jeffrey’s, is one of these evidences which used for human identification or toproof the relation to crime scene [2]. It depends on all the biological evidence that offender left behind in the crime scene such as blood, saliva, and semen. These traces almost visible but not associated with a wide range of crimes. In a new millennium, Touch DNA concept becomes the focus of recent forensic investigation. It refers to DNA/ epithelial skin cells that is transferred to items through casual contact. It plays an important role in forensic DNA community and become a much requested and successful test for DNA laboratories to perform in the violentor cold cases crime scene. It introduced in 1997 by van Oorschot et al., where there was seemingly no blood stain or other body fluid present [3]. For the last 15-20 years, the touch DNA performed by most public crime laboratories utilizes STR methods. Different objects have been evaluated as source for Touch DNA such as fingerprint [4] Cartridge cases [5] and bedding [6] using several collection methods such as swabbing [7,8] adhesive tape lifting [9-11], ii. Different extraction methods [12-14] and iii. Different PCR amplification kit and parameters [15,16]. Questioned document examination has been used as part of forensic investigation for over 100 years, including extortion notes, threatening letters and white powder contamination for latent indentations and recently DNA analysis. The purpose of the examination is to obtain information about the origin, history, authenticity and authorship of a document for legal purpose [17,18]. In this study we will evaluate three different papers mainly used as document a substrate for DNA typing using adhesive tape, anon destructive method for DNA recovering, and Power Plex® 16 HS Amplification kit.
Materials and Methods
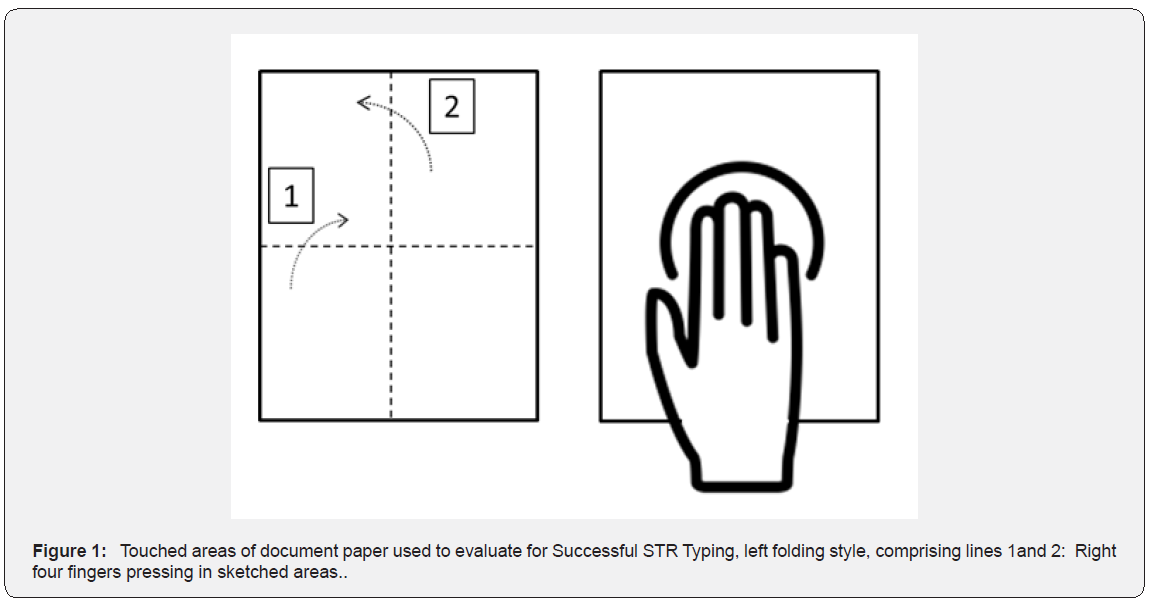
Three different paper supplies were purchased locally viz. standard 80 gsmA4 office/ copy paper (Roco, Indonesia), uncoated chart paper (Golden Coin, India)and coated paper (Feldmuehle, Germany) as it is the most commonly encountered paper type used for certificates printing and documents cover. All the tested paper types were decontaminated by exposure to UV light for 2 hrs. 10 healthy volunteers (5 male and 5 female) were recruited for the study. All the volunteers participating in this study were asked to refrain their hands from washing for at least 1 hour prior to their participation in experiments. Volunteers exposed their right hand above the boiling water bath for 10 seconds this is done to enable sufficient build –up of cells in their hands for the transfer and then probed their hand in the test paper for 2 mins; the site of the hand in the paper was sketched. Later the volunteers asked to fold the paper in predefined usual way and placed in an envelope as shown in Figure 1. Samples paper were prepared with the restriction to collect the sample of only one kind of paper for each day and stored at room temperature. Buccal swabs of the volunteers were collected as reference samples. Adhesive tape lifting technique was employed to collect the cells from the sketched area in the paper as well as along the fold sides on all the three types of paper. DNA was extracted from buccal swabs and Adhesive tape by Chelex® extraction protocol as described by [19] and quantified in 7500 Real-Time PCR System(PCR) using Quantifiler® Duo DNA Quantification Kit (Applied Biosystems- USA) to regulate the maximum input DNA for PCR amplification. Extracted DNA were amplified for 15 STR loci (D3S1358, THO1, D21S11, D18S51,Penta E, D5S818, D13S317,D7S820, D16S539, CSF1PO, Penta D,D8S1179, vWA, TPOX, FGA) in addition to the sex determining marker Amelogenin by multiplex assay using The PowerPlex®16HS Amplification kit (Promega Corporation, USA) in Veriti® 96-Well Thermal Cycler. Amplified STR alleles were size separated by capillary electrophoresis (CE) using 3130 Genetic Analyzer® (Life Technologies, USA) following the manufacturer recommended protocol. ILS 600 was used as an internal lane standard. Fragment size determination of the allele was analyzed using Gene MapperID-XSoftware Version 3.1 (Applied Biosystems, USA).Allele’s designation was based on comparison with the allelic ladder provided in the kit.
Results
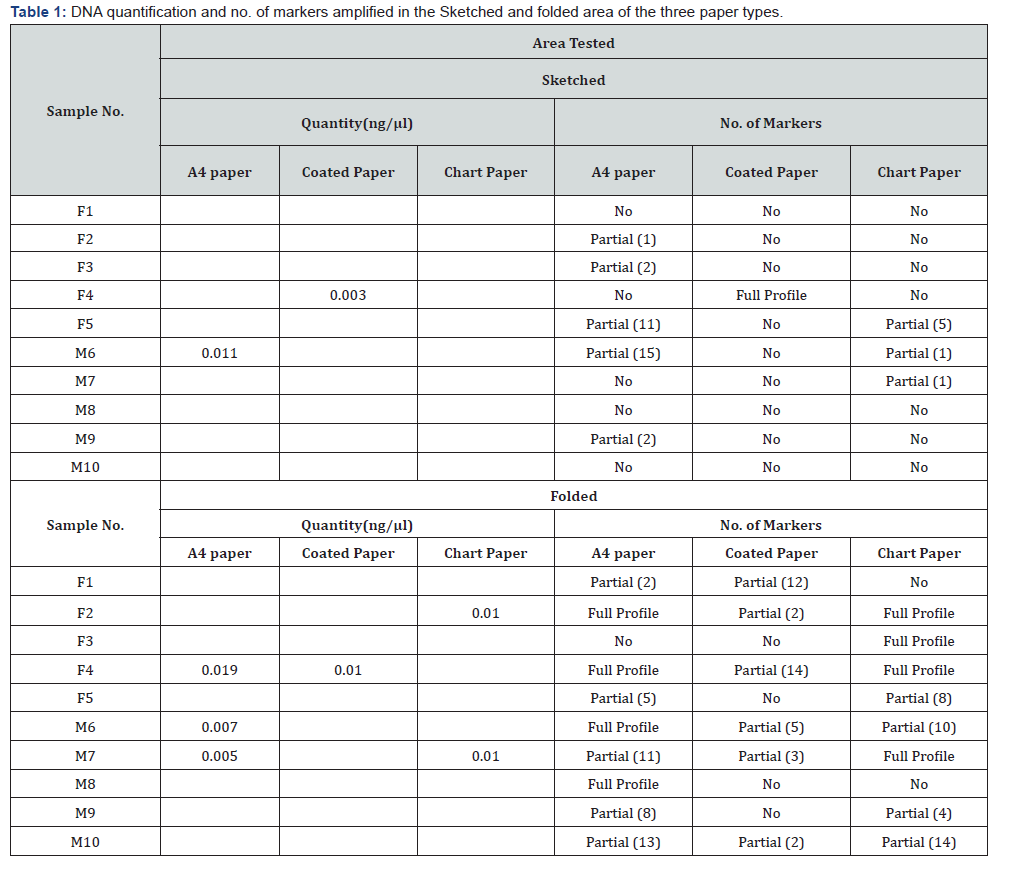
DNA quantitation using real time technique results provided the basics of understanding that only very few cells can be extracted from the touched samples. The concentration of the reference samples ranged between 2.63-0.37 ng/μl. By the way only 3 sample in total shows up for quantitation in different paper substrates. As expected, folded area in the paper accounts for higher quantitation than the sketched area in all the paper types. Of all, the fold area A4 paper brought about concentration 0.019,0.007,0.005 ng/μl for samples 4,6,7 respectively, whereas only sample 6 showed 0.011ng/μl in sketched area. In the case of coated paper, sample 4 displayed 0.03 and 0.01 ng/μl for sketched and folded area correspondingly and for chart paper sample 2 and 7 displayed 0.01ng/μl in the folded area is shown in Table 1. In all the samples, internal positive control’s cycle threshold was within the accepted range which inferred that the extracted DNA was void of PCR inhibitors. For PCR amplification 10 micro liters of the extracted DNA were placed into final reaction volume of 25μl and were amplified using standard parameters at 28 cycles. The genotyping of the samples with PowerPlex® 16HS Amplification kit was in concordant with real time concentration outcomes. About 50% Sketched samples in A4 paper showed no profile and 40% showed partial profile with allele drop out. Sample 6, in sketched area of A4 paper showed profile with allele dropout and mismatched allele at PentaE, D7S820, CSF1PO, and FGA when compared with reference sample. In contrast with folded area in A4 paper, full profile appeared at samples 2,4,6 and 8 whereas partial profile with allele dropout was observed at sample 1,4,5,7,9,10. Mismatched allele in samples compared to reference sample was seen at D3S1358 in sample 5, 7 and 10; TH01 at sample10; D21S11 at sample 7; D5S818 at sample 5 and 10; D16S539 at sample 7 and 10 and Penta D at sample 10.As in the instance of coated paper, except for sample 4 no profile was observed in all other samples of sketched area .Similar results were noted for sample3, 5,8 and 9 ;partial profile with allele dropout was observed for sample 1,2,4,6,7 and 10 in the folded area. Certainly no profile were detected in Chart paper of sketched area at sample 1,2,3,4,8,9 and 10; partial profile with allele dropout were also spotted at sample 5,6and 7. The folded area gave promising result with full profile for sample 2, 3 and 7; no profile were determined at sample 1 and 8.Partial profile with allele drop out were prominent at sample 4,5 6,9 and 10. Mismatched allele were observed at locus D13S317, D7S820 and vWA at sample 5 for both the area, likewise the sketched area of sample 4 showed mismatched alleles at locus D3S1358 and D7S820. These differences give the impression that folded area raged out the sketched area in the profiling pattern. Both A4 and chart paper in the folded area displayed almost 100% profiling in 4 and 3 samples correspondingly.
Discussion
In this study, we evaluate touch DNA as a tool in document investigation according to three categories, (i) The paper type, and (ii) DNA location, i.e. The touched side and the folded lines area and gender relatedness. Three types of press and writing papers substrate for touch DNA were used.PowerPlex® 16HS Amplification kit was used following conventional protocols for STR typing. A non-destructive method of sampling i.e. adhesive tape, has been selected to collect the skin cells from the substrates because its proved to be effective and more acceptable with the papers materials to avoid any damage may occur with the other collecting methods such as wet swabbing or wet/ dry swabbing. The amount of DNA recovered from almost all the experiments i. e folded line and touched area using 5% Chelex was low and varied between individuals and even in same individual. this support previous studies finding, that DNA deposition process effecting by many factors such as shedding status of the individuals [20], the nature of the touched surface, pressure applied on the surface during a contact [21], and the habits of thee person [22]. Therefore, using standard parameters at 28 cycles of amplification was not enough to amplify all the targeted loci. Increasing the cycle number to 34 cycles, to simulate LCN condition may give allelic data due to increasing of detection sensitivity. Comparing of DNA profiling using PowerPlex® 16HS Amplification kit, showed a noticeable that full profile were been able to acquire in the folded area.
Conclusion
Document examination is a routine work in forensic laboratory in many crimes involved, threading letters, counterfe it documents, and anonymous notes encountered. According on the results of this study, Touch DNA could be an effective tool in these crimes investigation to determine identity or the relation or contact with a concern document. The amount of DNA could be recovered from the touched document is low in general, but the folded line is could have more DNA than the other locations such as, surface area. Furthermore, selection the appropriate method for skin cells selection, and applying a low copy numbers of protocols are wanted for DNA typing instead of the standard protocols mainly used in forensic laboratories.
Acknowledgment
Authors would like to thank the participants in the study and Naif Arab University for security Science for their continuous support
References
- Locard E (1930) The analysis of dust traces. Am J Police Sci 1: 276.
- Butler, JM (2005) Forensic DNA typing: biology, technology, and genetics of STR markers. Academic Press.
- Van Oorschot RA, Jones M (1997) DNA fingerprinting from fingerprinting. Nat 387: 767.
- Sewll J, Quinones I, Multaney B, Curitis S, Seeboruth H, et al. (2008) Recovery of DNA and fingerprints from touched documented. Forensic Sci Int Genet 2(4): 281-285.
- Horsman Hall KM,Orihuela Y, Karczynski SL, Davis AL, Ban JD, et al. (2009) Development of STR profiles from firearms and fired cartridge cases. Forensic Sci Int Genet 3: 242-250.
- Petricevic SF, Bright JA, Cockerton SL (2006) DNA profiling of trace DNA recovered from bedding. Forensic Sci Int 159: 21-26.
- Sweet D, Lorente M, Lorente J, Valenzuela A, Westwood PD (1997) An improved method to recover saliva from human skin. J Forensic Sci 42: 320-322.
- Abaz J, Walsh SJ, Curran JM, Moss DS, Cullen J, et al. (2002) Comparison of the variables affecting the recovery of DNA from common drinking containers. Forensic Sci Int 126(3): 233-240.
- Joel J, Glanzmann B,Germann U, Cossu C (2015) DNA extraction of forensic adhesive tapes- A comparison of two different methods. Forensic Sci Inter: Genet Suppl Ser 5: 579-581.
- VerdonTJ, MitchellRJ, Van Oorschot RA (2014) Evaluation of tape lifting as a collection method for touch DNA. Forensic SciInter: Genet 8(1): 179-186.
- Barash M, Reshef A, Brauner P (2010) The use of adhesive tape for recovery of DNA from crime scene items. J Forensic Sci 55(4): 1058-1064.
- Stoop B, Defaux PM, Utz S, Zieger M (2017) Touch DNA sampling with SceneSafe Fast TM minitapes. Leg Med 29: 68-71.
- Alfadaly N, Kassab AC, Alrasheedy SF (2017) Evaluation of the techniques used in analysis of touch DNA collected from crime tools in Hail, Kingdom of Saudi Arabia. J Forensic Sci and CrimInvest 3(5): 1-6.
- Whitaker JP (2001) A comparison of the characteristics of profiles produced with the AMPFlSTR SGM plus multiplex system for both standard and low copy number(LCN)STR DNA analysis. Forensic Sci Inter 123(2-3): 215-223.
- Jeffreys A (1988) Amplification of human minisatellites by the polymerase chain reaction: Towards DNA fingerprinting of single cells. NuclAcid Res 16(23): 10953-10971.
- Tsai LC, Lee JC, Lin YM, Lai PY, Hsieh HM (2010) STR residues inside gloves. Forensic Sci J 9(1): 1-8.
- Ellen D (2006) Specific examination of documents: Methods and techniques Taylor and Francis group, LLV.
- Voyce J (2012) History of ESDA. Global Forensic Science Today. 30-31.
- Walsh PS, Metzger DA, Higuchi R (1999) ChelexTM 100 as a medium for the simple extraction of DNA for PCR-based typing from forensic material. Biotechniques 10(4): 506-513.
- Lowe A, Murray C, Whitaker J, Tully G, Gill P (2002) The propensity of individuals to deposit DNA and secondary transfer of low-level DNA from individuals to inert surfaces. Forensic Sci Int 129(1): 25-34.
- Tobias SH, Jacques GS, Morgan RM, Meakin GE (2017) The effect of pressure on DNA deposition by touch. Forensic Sci Int Genet 6: 12-e14.
- Phipps M, Petricevic S (2007) The tendency of individuals to transfer DNA to handled items. Forensic Sci Int 168(2-3): 162-168.






























