DIP-Linked DNA Polymorphism-A Novel Type of DNA Marker to Detect Mixture DNA
Jinding Liu and Gengqian Zhang*
School of Forensic Medicine, Shanxi Medical University, Jinzhong, Shanxi, P.R.China
Submission:October 3, 2019; Published: October 14, 2019
*Corresponding author:Gengqian Zhang, School of Forensic Medicine, Shanxi Medical University, Jinzhong, Shanxi, P.R.China
How to cite this article:Jinding Liu, Gengqian Zhang. DIP-Linked DNA Polymorphism-A Novel Type of DNA Marker to Detect Mixture DNA. J Forensic Sci & Criminal Inves. 2019; 12(5): 555850. DOI: 10.19080/JFSCI.2019.12.555850.
Abstract
The typing of mixture is a challenge in forensic and clinical medicine, especially for those unbalanced-and-degraded mixtures (UDM). To determine the minor contributor of the mixture is crucial to find the criminal in forensics. Traditional forensic DNA typing, such as autosomal STRs and Next Generation Sequencing have a limited sensitivity to find the minor contributor. Here, we make a mini review for the advance of new markers and their applications in forensic and clinical medicine. We also summarized and compared the merits and shortcomings of the traditional methods and new markers.
Keywords: Forensic genetics; Mixture; DNA markers
Abbreviations: UDM: Unbalanced and Degraded Mixture; LR: Likelihood; FACS: Fluorescence-Assisted Cell Sorting; LCM: Laser Capture Microdissection; CE: Capillary Electrophoresis; NGS: Next Generation Sequencing; DIP: Deletion/Insertion Polymorphism; UM: Unbalanced Mixture
Introduction
Detection of mixture stains has been one of the forensic difficulties for decades. Most techniques showed the mixture’s phenotype with the pattern of admix of alleles or phenotypes. Generally, the detected phenotype may be varied due to the constituent proportion. The phenotype of mixture stains was often used for exclusion of a suspect, rather than identify one. Provided the advance DNA genetic markers and fluorescent-labeling techniques make a huge step to extend the markers’ polymorphism and detection sensitivity, the principle to identify or exclude the suspect from stain evidence is not improved much. For example, current technology, such as STR-PCR followed by capillary electrophoresis showed the collective genotype of the mixture. Forensic experts had to differentiate the contributors based the probability of the putative allele combinations to produce a likelihood (LR) value to assess the possibility of the suspect involving the case [1,2]. Many software programs have been developed to meet the huge calculation task [3,4]. In recent years, many novel have been developed DNA markers based on the haplotypes of the contributors of the mixture. The novel markers usually have a higher sensitivity and specificity (based on the allele-specific amplification) compared with traditional STR examination. This review mainly focuses on the adventure of new markers and their (putative) applications in the forensic medicine.
The Strategies to Separate Contributor from Mixture Stains
As mentioned above, the conventional method to treat mixture stains is to take them as a whole. Moreover, researcher developed many strategies to separate the contributors of the mixture on different levels so as to get the unambiguous results of each contributor.
Separate the contributors on the cell level
To separate contributors with different cell types using antibodies- or fluorescence-assisted cell sorting to select the wanted cell type (FACS) [5,6]. The DEP Array™ system uses a single-use micro-fluidic cartridge that contains an array of electrodes that can be individually controlled, allowing cell going to different path [7]. A breakthrough of this method is to use the laser capture microdissection (LCM) system to select the target cells. This method needs unique equipment and has the possibility of collecting impure fractions [7,8]. All the above methods employed immune- or physical-separation to acquire the target cells before performing DNA analysis. However, such methods are tedious to separate the cells. Moreover, the limitation of such method is that it is only applicable to mixture consisting with different cell types. For those with the identical cell type, this method will fail to separate them. And unwanted peaks and allele drop-out/drop-in were reported using few target cells when using above techniques [7]. Another strategy widely used in forensic investigation is to separate the sperm cells from the vaginal fluid using the sperm properties of resisting digestion with Protein K. This method used different buffer to dissolve epithelial cells first and then digest the spermatozoa [9,10]. Such a strategy is very successful for identifying mixture consisting of spermatozoa and vaginal epithelial cells.
To Separate the contributors using male-specific genetic markers
One typical method is the application of Y-chromosome markers to identify the male criminal who might contribute to the semen/vaginal fluid mixture stain happened in a sexual assault case [11]. Most of the STRs and SNPs on the Y-chromosome are male-specific, and this is especially helpful for the mixtures consisted with semen and female resource cells. The Y-STRs have an extremely high sensitivity for successful detection of the minor component of the male/female mixture [11]. Y-chromosome is passed on in a paternal mode of inheritance, providing limited discriminatory power and is impossible to pin to specific individual or identify his criminal merely by the Y-chromosome STRs. However, it is great tool to help find the male perpetrator through the pedigree searching of Y-STRs [12].
Cloning method
Report a cloning method for mt DNA typing in a male-female mixture in a rape case [13]. The contributor’s DNA will be distributed to each clone after molecule ligation with the vector
Treat the integrated information of the mixture
This is the most widely used method to analysis mixtures. In many cases, the condition to separate the contributors does not exist, such as the requirement of consisting of different cell types, sperm-female cells. So, the mixtures have to be treated as a single-resource stain, especially when one does not know whether the stain is a mixture or not. All the conventional genetic markers are applicable to such a strategy. Such as ABO blood type, short tandem repeats (STRs), SNPs. Given the polymorphism and application feasibility, the strategy of STR typing with capillary electrophoresis (CE) is still the predominant technique to type mixture. The merit of this method is to get the combined information of all the contributors of the stain without limitation of cell types only if the component of the contributor is high enough (usually >10-20% in STR analysis). Besides the request of the amount of the component, another difficult is to calculate the likelihood value (LR) to assess how much possibility of the suspect is involved in. Such a calculation task becomes more and more difficult, even impossible along with the increase of mixture contributors [3]. Usually, the stain DNA would be amplified before CE or sequencing. The PCR used the primers undistinguished binding to each contributor’s DNA. Thus, the minor contributor would be masked by the major contributor unless it more than 10%-20%.
The NGS Technology
Next Generation Sequencing (NGS) reads and records sequences of clones of each template molecule. However, the primer-for-all-molecule strategy is used in NGS. The undistinguished binding primers would also have a cover effect for minor component DNA. Compared with STR-CE strategy, NGS is reported having similar or more sensitive detection ability for minor component from mixture [14,15]. As an advantage technique platform, most markers of could be performed NGS in a compatible run length, such as STR, SNP, In Del, microhaplotype and markers in linkage. All the markers should have a improved performance in mixture detection when they run on NGS than CE. However, this technique is costly and not widely equipped in regular forensic laboratories.
Microhaplotype (Micro haps)
Kidd et al. defined a microhaplotype to be a locus with two or more single nucleotide polymorphisms (SNPs) that occur within a short segment of DNA, usually 200bp, that can be covered by a single sequence run and collectively define a multi-allelic locus [16]. Micro hap markers have the inherent merits of SNP, which possess the lower mutation rates, pedigree properties and extensively distribution on human chromosome, moreover, it has higher polymorphic than independent SNPs [17]. The micro haps have a higher discrimination power and sensitivity when it combined with NGS. Choosing of appropriated micro haps will provide the perpetrator’s information of ancestral, eye color etc, other than the merely polymorphism information. The great potential of optimally selected micro haps in forensic casework encouraged the search for novel loci that were better suited for forensic applications.
InDel and DIP-linked DNA polymorphism and their implications
To get the minor contributor from the mixture, allelespecific primers were designed mainly based on the insertion/ deletion polymorphism (InDel, also termed as deletion/ insertion polymorphism, DIP). The Indel sequence is present or not in the human genomics, termed as insertion or deletion, separately. This strategy used allele-specific primer to avoid the masking effect of the major DNA, showing a 1:1000 sensitivity for selecting minor DNA.
InDel marker
InDel is a kind of promising marker to differentiate donor and recipient’s DNA after the patient received a bone marrow transplant surgery and the quantity of donor’s DNA could be determined using Taqman technology by establishing a standard curve [18]. We prove that InDel is enough to identify the mixture using quantitative PCR (qPCR) technique with Sybr green dye [19]. The difference of Ct value (D-value) could be used to test whether the stain is a mixture or not without establishing standard curve. The InDel should be cautiously selected since the length of InDel sequence affects the Tm and the D-value [19]. The sensitivity for detecting minor DNA is between 1:50 to 1:1000 or more. InDel could be used to exclude a suspect by the different allele combination patterns, however, the shortage of polymorphism reduced its power in validating a suspect.
DIP-STR
Several sets of DIP-STR markers have been proposed in recent years [20-23]. DIP-STRs are novel genetic markers composed of a DIP tightly adjacent to an STR polymorphism, detected by a specially designed pair of PCR primers (L-long allele for insertion and S-short allele for deletion). These compound markers are able to target a minor contributor in the presence of a 1000-fold excess of background DNA from an unbalanced DNA mixture. Bayesian networks was used for evaluating DIP-STR profiling. Moreover, DIP-STR was applied to UDM from touch DNA samples and fetal DNA from maternal plasma, showing the superior sensitivity than other methods [20,24]. The extremely high sensitivity for the minor contributor of the mixture is endorsed by the selectively binding property of DIP-matching PCR primers, i.e. one primer matching to the insertion, the other the deletion sequence.
DIP-SNP
A set of DIP-SNP markers has been first proposed in 2017 by our team [25]. In 2019, we proposed a new set of DIP-SNP markers, which were used with unbalanced and degraded DNA mixture [26]. A DIP-SNP marker is a tightly linked compound marker composed by DIP (deletion/insertion) and SNP with a total length of 200bp. We designed this method and find DIPSNP markers because most DIP-STRs are too long to be used for degraded samples. DIP-SNP showed a superior capability for degraded DNA. Since the DIP-SNP markers have an extensively distribution on human genome, more DIP-SNP markers should be used to identify DUM, which is an important issue in forensic medicine. The DIP sequence length is critical for the specificity and sensitivity and preferred sequence length should be 2-10bp. Shorter than such length would failed to guarantee the specificity, on the other hand, longer sequence would be difficult to make a multi-plex PCR since the variable sequence would change Tm too much [25].
SNP-STR
This kind of marker was developed by Zhang Lin et al and the SNP was used to design specific primers and the linked STR to improve the discrimination power [27-29]. The SNP is used as discriminative marker using specific PCR primers to differentiate the linked STR allele. They stated a higher sensitivity for unbalanced mixture. All the markers used allele-specific PCR primers to select the minor contributor of the mixture, such as InDel, DIP-STR, DIP-SNP and SNP-STR markers, are emerging as a new kind of DNA markers to discriminate minor DNA using their representative allele, showing a superior sensitivity for unbalanced DNA mixture than the non-selecting binding primer strategies.
Discussion
Two difficulties of mixture analysis have to be mentioned here. The first is that how to type the degraded and unbalanced mixture (DUM). The second is that the identification of mixture containing many contributors. For STR analysis, 3 methods have been suggested for inferring the number of contributors, NOCIt, maximum likelihood estimator, and maximum allele count [30]. Y-STRs analysis can be used to estimate the number of contributors in a mixture stain which are not restricted by the content of the female. It must be noted that the mixture deconvolution becomes extremely difficult when the number of contributor increase using co-amplified strategy, such as STR, SNP, micro haps. For example, the knob of the hotel door may contain epithelial cells derived from tens of people with varying proportions, i.e. the touch DNA. It is impossible to separate each contributor using antibody-staining or antibodydriving methods to separate each contributor since they are the same cell type. Suppose that one STR with limited alleles, such as 12 alleles in certain population, the swab of the doorknob may contain most of the alleles, even all the alleles, and such situation may be happening for other STRs. How to find the target contributor, or putative crime? It is impossible to conduct the co-amplified strategy, regardless the following step is NGS or CE. The DIP-linked DNA marker have the potential to resolve such cases, because they divide each contributor based on the whether the varied linked-allele present or not.
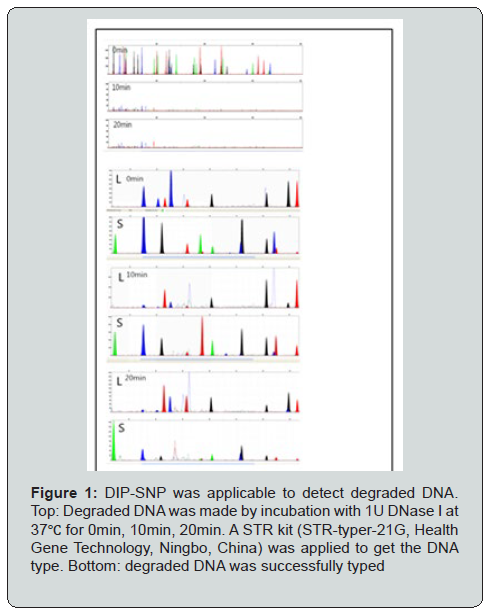
The mixture that contains minor contributor less than 10% is termed unbalanced mixture (UM) [31]. Degraded and unbalanced mixture (DUM) occur in forensic medicine and clinical examination. Stains usually experience a period of ex vivo exposure and degradation in different environment before taking DNA test. In clinical practice, mixture samples might be shaped in the blood of a pregnant woman with fetal degraded DNA etc. To resolve the DUM, the shorter DIP-linked DNA markers, such as DIP-SNPs are proffered. We proved that the DIP-SNPs are capable of detecting the degraded DNA when the STR kit fails to type (Figure 1).
Conclusion
In summary, people strive to resolve mixture typing, especially for the mixture containing multi-donors or UDM. The DIP-linked DNA polymorphism seems a promising tool to resolve this problem. However, there is a long way to make it suitable to resolve all the issue.
Acknowledgemnet
This work was supported by Key Research and Development (R&D) Projects of Shanxi Province (201803D31069) and Shanxi Scholarship Council of China (2016-055).
References
- Gill P, Brenner CH, Buckleton JS, Carracedo A, Krawczak M (2006) DNA commission of the International Society of Forensic Genetics: Recommendations on the interpretation of mixtures. Forensic Sci In 160(2-3): 90-101.
- Bieber FR, Buckleton JS, Budowle B, Butler JM, Coble MD (2016) Evaluation of forensic DNA mixture evidence: protocol for evaluation, interpretation, and statistical calculations using the combined probability of inclusion. BMC Genet 17(1): 125.
- Barrio PA, Crespillo M, Luque JA, Aler M, Baeza Richer C, et al. (2018) GHEP-ISFG collaborative exercise on mixture profiles (GHEP-MIX06). Reporting conclusions: Results and evaluation. Forensic Sci Int Genet 35: 156-163.
- Bright JA, Taylor D, McGovern C, Cooper S, Russell L, et al. (2016) Developmental validation of STR mix, expert software for the interpretation of forensic DNA profiles. Forensic Sci Int Genet 23: 226-239.
- Verdon TJ, Mitchell RJ, Chen W, Xiao K, van Oorschot RA (2015) FACS separation of non-compromised forensically relevant biological mixtures. Forensic Sci Int Genet 14: 194-200.
- Xu Y, Xie J, Chen R, Cao Y, Ping Y, et al. (2016) Fluorescence- and magnetic-activated cell sorting strategies to separate spermatozoa involving plural contributors from biological mixtures for human identification. Sci Rep 6: 36515.
- Williamson VR, Laris TM, Romano R, Marciano MA (2018) Enhanced DNA mixture deconvolution of sexual offense samples using the DEPArray system. Forensic Sci Int Genet 34: 265-276.
- Vandewoestyne M, Van Hoofstat D, Van Nieuwerburgh F, Deforce D (2009) Automatic detection of spermatozoa for laser capture microdissection. Int J Legal Med 123(2): 169-175.
- Horsman KM, Barker SL, Ferrance JP, Forrest KA, Koen KA, et al. (2005) Separation of sperm and epithelial cells in a microfabricated device: potential application to forensic analysis of sexual assault evidence. Anal Chem 77(3): 742-749.
- Budowle B, Baechtel FS, Comey CT, Giusti AM, Klevan L (1995) Simple protocols for typing forensic biological evidence: chemiluminescent detection for human DNA quantitation and restriction fragment length polymorphism (RFLP) analyses and manual typing of polymerase chain reaction (PCR) amplified polymorphisms. Electrophoresis 16(9): 1559-1567.
- Roewer L (2009) Y chromosome STR typing in crime casework. Forensic Sci Med Pathol 5(2): 77-84.
- Kayser M (2017) Forensic use of Y-chromosome DNA: a general overview. Hum Genet 136(5): 621-635.
- Hatsch D, Amory S, Keyser C, Hienne R, Bertrand L (2007) A rape case solved by mitochondrial DNA mixture analysis. J Forensic Sci 52(4): 891-894.
- van der Gaag KJ, de Leeuw RH, Hoogenboom J, Patel J, Storts DR, et al. (2016) Massively parallel sequencing of short tandem repeatsPopulation data and mixture analysis results for the Power Seq system. Forensic Sci Int Genet 24: 86-96.
- Wang L, Chen M, Wu B, Liu YC, Zhang GF, et al. (2018) Massively Parallel Sequencing of Forensic STRs Using the Ion Chef and the Ion S5 XL Systems. J Forensic Sci 63(6): 1692-1703.
- Kidd KK, Speed WC (2015) Criteria for selecting microhaplotypes: mixture detection and deconvolution. Investig Genet 6: 1.
- Kidd KK, Speed WC, Pakstis AJ, Podini DS, Lagace R, et al. (2017) Evaluating 130 microhaplotypes across a global set of 83 populations. Forensic Sci Int Genet 29: 29-37.
- Alizadeh M, Bernard M, Danic B, Dauriac C, Birebent B, et al. (2002) Quantitative assessment of hematopoietic chimerism after bone marrow transplantation by real-time quantitative polymerase chain reaction. Blood 99(12): 4618-4625.
- Liu J, Wang J, Zhang X, Li Z, Yun K, et al. (2017) A mixture detection method based on separate amplification using primer specific alleles of INDELs-a study based on two person’s DNA mixture. J Forensic Leg Med 46: 30-36.
- Oldoni F, Castella V, Grosjean F, Hall D (2017) Sensitive DIP-STR markers for the analysis of unbalanced mixtures from “touch” DNA samples. Forensic Sci Int Genet 28: 111-117.
- Oldoni F, Castella V, Hall D (2015) A novel set of DIP-STR markers for improved analysis of challenging DNA mixtures. Forensic Sci Int Genet 19: 156-164.
- Cereda G, Biedermann A, Hall D, Taroni F (2014) Object-oriented Bayesian networks for evaluating DIP-STR profiling results from unbalanced DNA mixtures. Forensic Sci Int Genet 8(1): 159-169.
- Castella V, Gervaix J, Hall D (2013) DIP-STR: highly sensitive markers for the analysis of unbalanced genomic mixtures. Hum Mutat 34(4): 644-654.
- Moriot A, Hall D (2019) Analysis of fetal DNA in maternal plasma with markers designed for forensic DNA mixture resolution. Genet Med 21(3): 613-621.
- Liu Z, Liu J, Wang J, Chen D, Liu Z, et al. (2018) A set of 14 DIP-SNP markers to detect unbalanced DNA mixtures. BiochemBiophys Res Commun 497(2): 591-596.
- Liu J, Li W, Wang J, Chen D, Liu Z (2019) A new set of DIP-SNP markers for detection of unbalanced and degraded DNA mixtures. Electrophoresis 40(14): 1795-1804.
- Tan Y, Bai P, Wang L, Wang H, Tian H, et al. (2018) Two-person DNA mixture interpretation based on a novel set of SNP-STR markers. Forensic Sci Int Genet 37: 37-45.
- Wei T, Liao F, Wang Y, Pan C, Xiao C, et al. (2018) A novel multiplex assay of SNP-STR markers for forensic purpose. PLoS One 13(7): e200700.
- Gao JS, Ye Y, Hou YP (2017) Application of SNP-STR Composed by D18S51 and Three SNPs of Its Flanking Region in Paternity Testing]. Fa Yi Xue Za Zhi 33(6): 607-610.
- Alfonse LE, Tejada G, Swaminathan H, Lun DS, Grgicak CM (2017) Inferring the Number of Contributors to Complex DNA Mixtures Using Three Methods: Exploring the Limits of Low-Template DNA Interpretation. J Forensic Sci 62(2): 308-316.
- Cereda G, Biedermann A, Hall D, Taroni F (2014) An investigation of the potential of DIP-STR markers for DNA mixture analyses. Forensic Sci Int Genet 11: 229-240.






























