Genetic Polymorphism of 16 Autosomal STRs Loci of the PowerPlex ESX 17 System in a Population Sample from Bangladesh
Sharif Akhteruzzaman1*, Mahamud Hasan2, Abu Sufian2, Pilu Momtaz2, Ashish Kumar Mazumder2, Jabedul Alam Khandaker2 and Amena Binte Husain2
1Department of Genetic Engineering and Biotechnology, University of Dhaka, Bangladesh
2National Forensic DNA Profiling Laboratory, Dhaka Medical College Campus, Bangladesh
Submission: July 29, 2018;Published: August 10, 2018
*Corresponding author: Sharif Akhteruzzaman, Department of Genetic Engineering and Biotechnology, University of Dhaka, Dhaka-1000, Bangladesh, Tel: +880-1714 111 267; Email: sazaman@du.ac.bd
How to cite this article: Sharif Akhteruzzaman, Mahamud Hasan, Abu Sufian, Pilu Momtaz, Ashish Kumar Mazumder, Jabedul Alam Khandaker, Amena Binte Husain. Genetic Polymorphism of 16 Autosomal Strs Loci ofthe PowerPlex Esx 17 Ayatemin A Population Sample from Bangladesh. J Forensic Sci & Criminal Inves 2018; 10(2): 555784. DOI:10.19080/JFSCI.2018.10.555784.
Abstract
Allele frequencies of 16 autosomal short tandem repeats loci, D18S51, D21S11, TH01, D3S1358 , D16S539, D2S1338, D1S1656, D10S1248, FGA, D8S1179, VWA, D22S1045, SE33, D19S433, D12S391 and D2S441 were investigated in 95 unrelated Bangladeshi individuals using PowerPlex ESX 17 System Kit. Forensic efficiency parameters, the probability of match, power of discrimination, polymorphism information content, power of exclusion of paternity, observed heterozygosity and expected heterozygosity were calculated for the loci. The allele frequencies distribution and forensic statistical parameters indicated the usefulness of these loci in forensic investigation and personal identification in the Bangladeshi population. The combined probability of match (PM) and combined power of exclusion (PE) was 1.31×10-20 and 0.999736 for the loci.The PowerPlex ESX 17 System though was developed and recommended by ENFSI for DNA profile sharing across Europe, the system was also proved to be useful for forensic caseworks in Bangladeshi population.
Keywords: Allele Frequency; Power of Discrimination; Polymorphism; Heterozygosity; Forensic Investigation
Introduction
The PowerPlex® ESX 17 System loci primarily developed for use in European population. Forensic DNA assessment based on microsatellite sequences or short tandem repeats (STRs) has emerged as a powerful tool for human identification in criminal investigations, resolving parentage dispute and population genetic studies in recent days. Bangladesh is a South Asian country, bordered on the north, east and west side with India and, a short land and water frontier with Myanmar in the southeast. About 98% of the population in Bangladesh derives from Bengali ethno-liguistic group. The rest 2% are mainly etnic minorities living in the Chittagong Hill Tracts, Sylhet, Mymenshing and North Bengal regions of the country[1,2]. In this study we evaluated the effectiveness of thePowerPlex ESX 17 System loci primarily developed for use in European population.
Method
Whole blood or buccal cells were collected from randomly selected 95 individuals from Bangladeshi population . Samples were taken with informed consent from all individuals following procedures that are in accordance with the Helsinki Declaration of 1964, revised in 1983 [3]. Genomic DNA was extracted using the Chelex-100 method described by PS Walsh [4]. Extracted DNA was quantified by NanoDrop-1000 (Nano Drop Technologies, Inc, Washington DE 19810, USA).Approximately 1 to 2 ng of genomic DNA was used for PCR amplification procedure for each sample. The PCR amplification reaction was carried out by using VeritiThermal Cycler (Applied Biosystems, USA). The thermal cycling conditions were employed according to the manufacturer’s instructions (Promega Corporation, USA). The PCR amplified products were separated and typed by capillary electrophoresis on ABI 3500 Genetic Analyzer (Applied Biosystems, USA) using POP-4 polymer and Data Collection Software v1.0. Peak sizing and genotype assignments were done by Gene Mapper ID-X v1.2.Allele frequencies and statistical parameters of forensic efficiency were calculated by using PowerStat Microsoft Excel Workbook version 1.2 [5]. The Hardy-Weinberg equilibrium and forensic efficiency parameters were calculated using Arlequin Software version 3.5 [6].
Results and Discussion
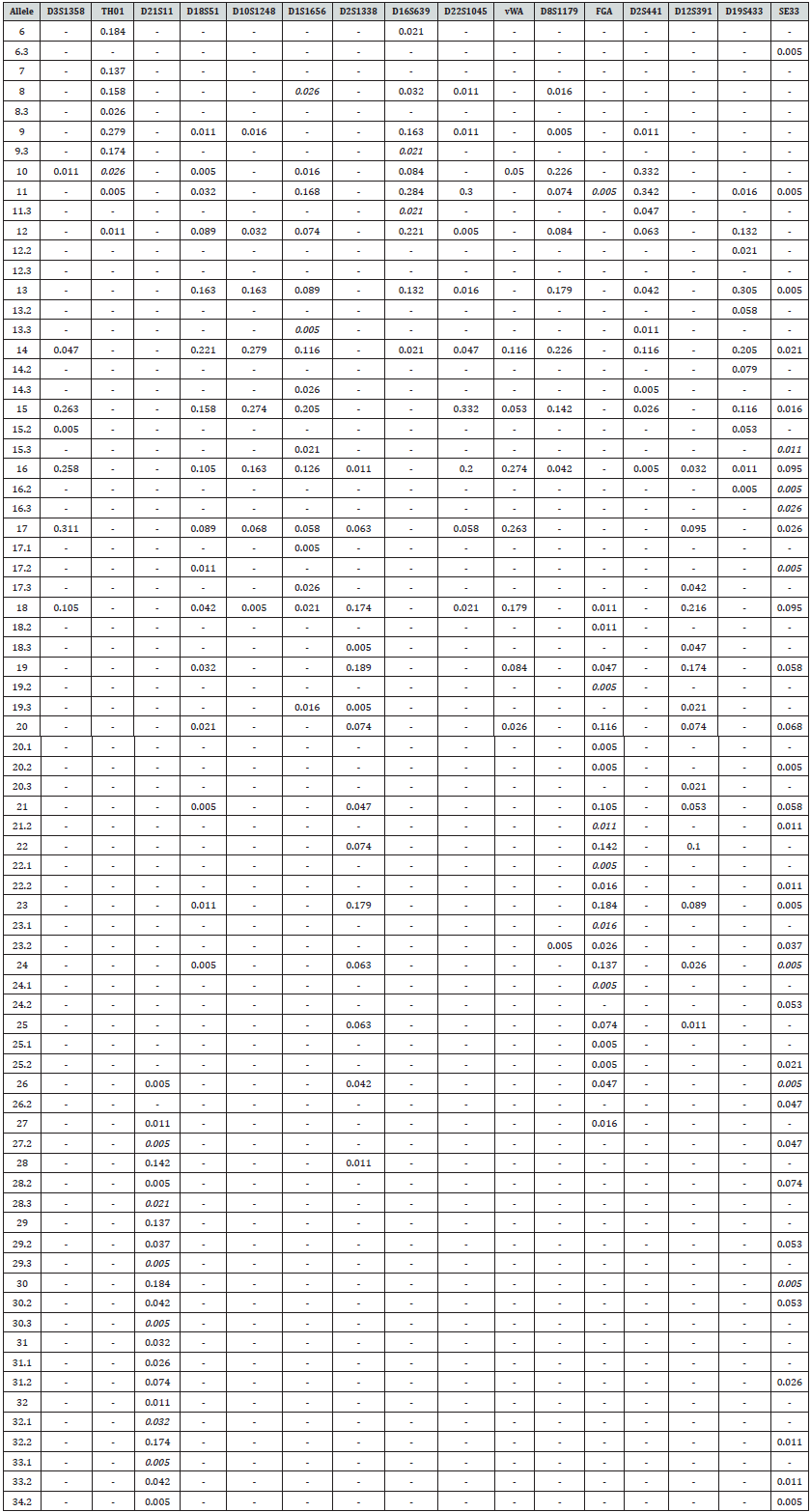
Allele frequency and forensic efficiency parameters for 16 STR loci are presented in Tables 1&2respectively. A total of 75 alleles found at the 16 loci studied. For each locus, at least seven alleles were observed (Table 1). SE33 was the highest polymorphic locus bearing 33 different alleles and 71 genotypes compared to the other loci. The combined frequencies were used to calculate forensic efficiency parameters. Eight out of the 16 STR loci studied were in Hardy–Weinberg equilibrium (using a 5% significance level) the eight remaining loci, D18S51, D1S1656, D16S639, D22S1045, FGA, D12S391 and SE33, showed a slight deviation from expectations (0.002<P<0.05). The PIC value for all the STR loci were highly informative (PIC>0.7). The most informative locus among the 16 STR loci was SE33 (PIC = 0.94), while the least informative were D3S1358, D22S1045 and D2S441 (PIC=0.71).A large number of microvarient and rare alleles (35 alleles) were observed and are shown in italics format (Table 1).The allele 29.3 at D21S11, allele 8 at D1S1656 and allele 19.3 at D12S391 were previously reported for Bangladeshi population [7,8]. and reported only once in STRBase [9]. Allele 11 was found to be the most frequent allele at D2S441 locus followed by allele 10 at D2S441 and allele 15 at D22S1045 loci, respectively. Out of the sixteen loci studied, D2S1338 and SE33 were the most informative locus based on observed heterozygosity (0.895 and 0.884 respectively). The combined probability of match (PM), combined power of discrimination and combined power of exclusion for the studied loci were 1.31×10-20, 1and 0.999736, respectively.
Table 1: Allele frequencies of 16 autosomal STR loci of PowerPlex® ESX 17 System in a population sample from Bangladesh (n = 95).
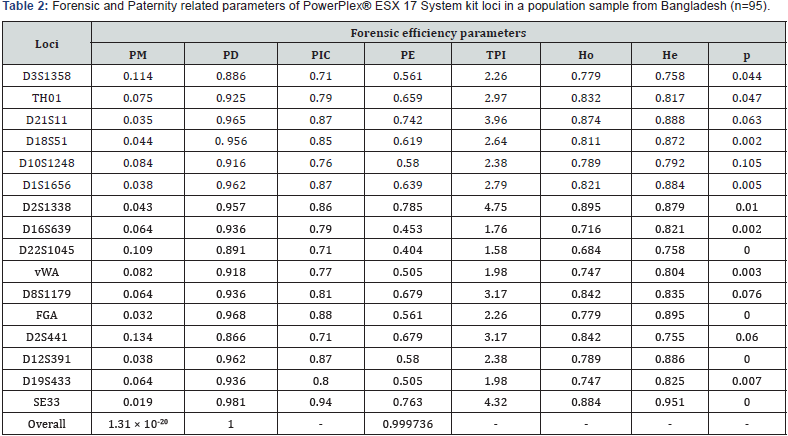
Forensic and Paternity related parameters of PowerPlex® ESX 17 System kit loci in a population sample frfrom Bangladesh (n=95). PM, probability of match; Ex1, Expressed as 1 in ...; PD, power of discrimination; PIC, polymorphism information content; PE, power of exclusion of paternity; TPI, typical paternity index; Ho, observed heterozygosity; He, expected heterozygosity; p, Hardy- Weinberg equilibrium exact test.
Conclusion
Genotyping of the 16 autosomal STR loci including SE33 locus may provide a useful adjunct to the previously studied autosomal STR loci evaluated for this population in the past.
Acknowledgement
This study was supported by the Multi-Sectoral Program on Violence Against Women (MSPVAW), Ministry of Women and Children Affairs, Government of the People’s Republic of Bangladesh.
References
- Abdullah N (2011) Survey on Socio-Demographic and Health Status of Tribal Community of Bangladesh: Santals. Dissertation, East West University, Bangladesh.
- Hannan MA (2015) Human rights of the aboriginals in the context of Bangladesh. OIDA International Journal of Sustainable Development 8(2): 45-68.
- Carlson RV, Boyd KM, Webb DJ (2004) The revision of the Declaration of Helsinki: past, present and future. Br J Clin Pharmacol 57(6): 695- 713.
- Walsh PS, Metzger DA, Higuchi R (1991) Chelex 100 as a medium for simple extraction of DNA for PCR- based typing from forensic material. Biotechniques 10(4): 506-513.
- Tereba A (1998) Tools for analysis of population statistics, Profiles DNA 2: 14-16.
- Excoffier L, Lischer HE (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows, Mol Ecol Resour 10(3): 564-567.
- Ferdous A, Ali ME, Alam S, Hossain T, Hany U, et al. (2006) Genetic data on 10 autosomal STR loci in the Bangladeshi population. Legal Medicine 8 (2006): 297-299.
- Rahmana AS, Das SA, Hasan MM, Akhteruzzamana S (2014) Evaluation of five new European Standard Set (ESS) STR loci in a Bangladeshi Population Sample. Mal J Forensic Sci 5(2):13-17.
- Short Tandem Repeat DNA Internet Database (NIST Standard Reference Database SRD 130).






























