Faith and Forensic: A Case Study
Francesco Sessa1, Gabriela Perilli1, Amos Tambo2 Mohsin Roshan2, Luigi Cipolloni3, Christian Zammit2, Cristoforo Pomara1,2* and Monica Salerno1
1Department of Forensic Pathology, University of Foggia, Italy
2Department of Anatomy, University of Malta, Europe
3Department of Forensic Pathology, University of Roma “La Sapienza“, Italy
Submission: November 25, 2016; Published: January 03, 2017
*Corresponding author: Cristoforo Pomara, Department of Forensic Pathology, University of Foggia, Department of Anatomy, School of Medicine, University of Malta, D'Avanzo Hospital, 71122 Foggia, Italy, Europe, Tel: (39) 0881.736926; Email: cristoforo.ponara@unifg.it
How to cite this article: Francesco S, Gabriela P, Amos T, Mohsin R, Luigi C, et al. Faith and Forensic: A Case Study. J Forensic Sci & Criminal Inves. 2017; 1(3): 555563. DOI: 10.19080/JFSCI.2017.01.555563
Abstract
A multidisciplinary forensic approach is applied to the study of ancient human skeletal remains of an historical figure venerated as a Saint by the Catholic Church, Fortunato of Serracapriola (South of Italy). Despite the forensic multidisciplinary approach is recommended for the study of ancient human skeletal remains, to the best of our knowledge, it is rare that this approach can be used on historical figure venerated as a Saint by Catholic Church. A radiological investigation was carried out for preliminary information: sex, stature, age and/or pathological signs. Subsequently radiocarbon analysis for dating of skeletal remains was performed. Finally the biological profile (STRs and mtDNA) and statistical analysis of genetic data were done to confirm the sex and to establish the provenience. Although we were not able to matching our data, it was very important to collect as much as possible information from the sample, it was the only way to confirm the correspondence of the scientific data with Saint's life. This case highlights the complexity of study of remains that were skeletonized. Therefore a multidisciplinary investigation as well as paying careful attention in the collection and correct interpretation of findings is important. We found a strong scientific concordance between our findings and the époque of the saint's existence, sex, and ethnicity. Archaeological remains can provide concrete cases, making it possible to develop, refine or validate medico-legal techniques.
Keywords: Ancient DNA (aDNA); Skeletal remains; Multidisciplinary approach; STR panel; Mitochondrial DNA (mtDNA) analysis
Abbreviations: AMS: Accelerator Mass Spectrometry; aDNA: Ancient DNA; mtDNA: Mitochondrial DNA; QMN: Quasi-Median Network; UV: Ultraviolet; CEDAD: Center for Dating e Diagnostics; LT : Low Template
Introduction
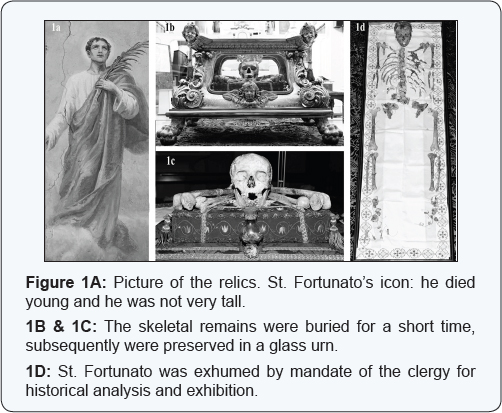
Little is known concerning the life of Fortunato of Serracapriola (a small town of countryside of Foggia- Puglia Region), an historical figure venerated as a Saint by the Catholic Church, particularly about the manner of his death. The first document on San Fortunato is an official letter written by Pope Innocent XI in 1687. With this letter the skeletal remains were donated to a local cardinal Carafa, after the removal from the catacombs ofthe family of Pope Pontianus in Rome, where initially conserved, and where these were identified by an inscription on the tombstone: "Fortunato Christi Martyris". Although no historical references are made to St. Fortunato's martyrdom, his death is dated back to the persecution of Christians under Maximinus Thrax, the Roman Emperor from 235 AD: in the historical reports, he was described, and so figurated, as a blond young boy not very tall (Figure 1A) [1]. In 1761 St. Fortunato was dubbed patron "minus principalis" of Serracapriola in 2010 (250 years later), and was exhumed (Figure 1B & 1C) by mandate of the clergy for historical analysis and exhibition. To the best of our knowledge, it is rare that a forensic multidisciplinary approach is request by the Catholic Church to study an historical figure venerated as a Saint. As previously described, radiological study was carried out with anthropological multi-slice computed tomography. This investigation was performed to carry out reconstructive identification of the skeletal remains, to seek evidence of taphonomic processes and to evaluate potential pathological changes. The radiological data were suggestive of a male, of height between 1.53m and 1.56m, age at death estimated between 20 and 40 years. No formal and specific signs of bone trauma such as hard bone callus were visible. No features indicating arthritis or bone infection were observed [2].
Following the radiological results, a specific permission was granted for perform radiocarbon and genetic analysis, beginning from a small fragment of bone (12g of femur). Radiocarbon dating was performed with accelerator mass spectrometry (AMS) with the objective to investigate if the bones date back to the 3rd century AD., so as to establish if the bones are compatible with St. Fortunato's historical data. AMS is the most sensitive method available for detecting and quantifying rare long lived isotopes with high precision. The most widely used isotope studied with AMS is 14C [3]. Genetic analysis was performed on the second bony fragment: bones, teeth and hair are the most widely used sources of samples for ancient DNA (aDNA) analysis [4]. Following DNA extraction, STR analysis was performed to determine the sex and to conjecture provenience with Random Probability. Finally was obtained the “mitochondrial DNA (mtDNA)'' profile, that it was analyzed with a valuable tool for graphical representation of mtDNA data, namely quasi-median network (QMN) for construction of reduced and filtered haplotypes. This tool is provided via the EMPOP database (website www.empop.org), a collaborative project for the provision of high-quality mtDNA population data for forensic purpose [5].
Case Report
DNA authenticity was guaranteed following criteria of previously described [6]. The bone sample was sub-divided into two pieces: one (7g) for radiocarbon dating and the other (5g) for DNA extraction. The outer surface of the bone sample used for genetic analysis was removed to approximately 2-3 mm depth with a sanding machine (Dremel) and fragmented into small pieces using a dental diamond disc. The bony fragments were exposed to ultraviolet (UV) light for 30 mins. The procedure was necessary as there was a risk of modern DNA contaminating the sample.
Radiocarbon dating
The surfaces of the bone fragment were cleansed as previously described. The 14C analysis was performed on CEDAD (Center for Dating e Diagnostics - Department of Innovation Engineering - University of Salento), following standard protocols for dating bone samples by collagen extraction, and AMS accelerator determination of the 14C content with the Uppsala 5MV pelletron system.
DNA extraction and Genetic analysis
For DNA extraction the bone fragment was triturated with a steel piston to obtain a fine powder. The samples were incubated on a rotor in EDTA solution for 24h, at 4°C in the dark. Following this step, the DNA was extracted using a QIAGEN® Investigator® extraction kit. Two extractions were performed from the bone for replication purposes. For each extraction, a negative control was used as one of the stringent quality assurance criteria in order to detect potential contamination. The AmpFLSTR® Identifiler® Plus PCR Amplification Kit was used for DNA amplification, the fragments were run on the Applied Biosystems® 3130 Genetic Analyzer, and the analysis was performed with GeneMapper® ID-X v1.4. Subsequently, the two hyper variable regions of the mtDNA (HVS1 and HVS2) were amplified using four pairs of overlapping primer sets for region, as previously described [5]. PCR products were handled by ABI Prism Big- Dye kit (Applied Biosystems, Foster City, CA), separated by ABI 3130 genetics analyzer (Applied Biosystems, Foster City, CA) and finally analyzed using the Sequence Navigator software sequencer (version 1.01; Applied Biosystems). Each sequence was confirmed with a new amplification. A mtDNA profile was obtained, in terms of Differences from Cambridge Reference Sequence [7], that we analyzed it in the EMPOP database (www. empop.org).
Results
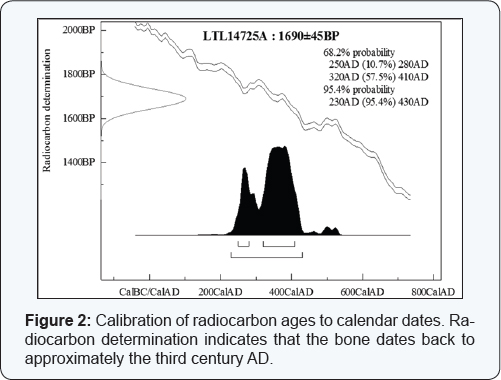
The radiocarbon dating revealed that the bone fragment dated back to 3rd century AD with a confidence limit of 95.5% (Figure 2). Although the skeletal remains were not preserved in an ideal manner during this period of time, the results show a high probability that they date back to the period in time in which St. Fortunato lived. The "genotyping” was particularly problematic because the DNA was isolated from an ancient specimen. When handling ancient bone samples one must consider two important aspects. Firstly, the structure of bones protects cells and DNA against exogenous and endogenous degradation factors, and mineral components such as hydroxyapatite stabilize the structure of the contained DNA molecules [4,8]. On the other hand, post-mortem phenomena can lead DNA damage, especially for hydrolysis and oxidation. Consequently, extracted aDNA is often very fragmented and is mostly low molecular weight. The length of DNA fragments ranges between 40-500bp; the average fragment size is 100bp [9,10]. Another problem when handling ancient specimen is the quantity of DNA that often is very low. Moreover DNA typing is accompanied by stochastic amplification effects, such as allele drop-outs and drop-ins; therefore, low template (LT) DNA profiles must be interpreted with care. The gold standard method for avoid these problems is "consesus method" that uses alleles confirmed in replicate analyze [11].
A low concentration of DNA was obtained from 0.05 g bone powder (≤ 100-200 pg /μL); however the standard method for STR amplification was utilized (28 cycles). All PCR amplification
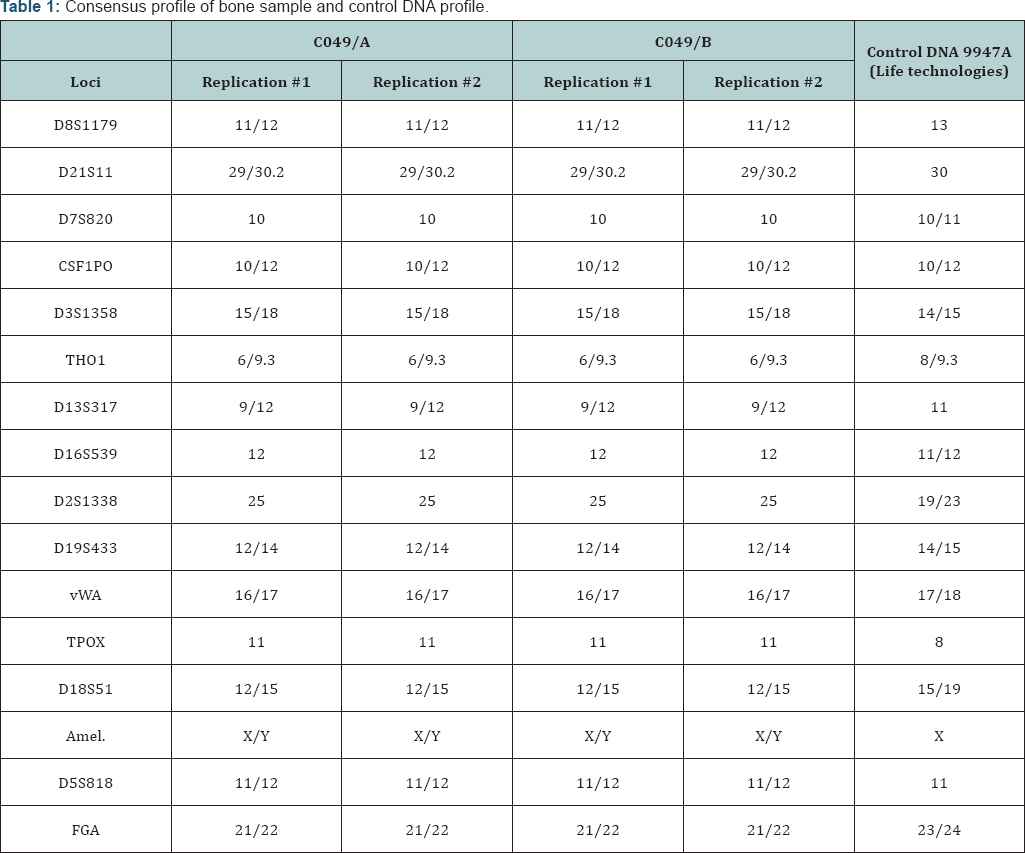
reactions were accompanied by negative controls. The negative controls showed no allele or locus drop-in, confirming the authenticity of the results. There was no indication of staff contamination when staff DNA profiles were compared to the results obtained from the bone sample. A positive control was utilized to verify the efficiency of the PCR (Control DNA 9947A, Life Technologies). Two replicates were produced independently for each aliquot of the bone sample: we obtained a complete DNA profile from bone sample. Consensus profile was summarized in (Table 1). Statistical analysis of STR data was performed: applying the combined matching probability for ethnicity and geographical region (SPSmart database). The combined matching probability reveals that the sample belongs to Caucasian male rather than a different ethnicity (American, Asian, Australian, and African). The statistical results are summarized in (Table 2). mtDNA analysis was conducted sequencing two hyper variable sections (HV1, HV2) of the mtDNA control region of bone sample. The data obtained was summarized in terms of Differences from Cambridge Reference Sequence.

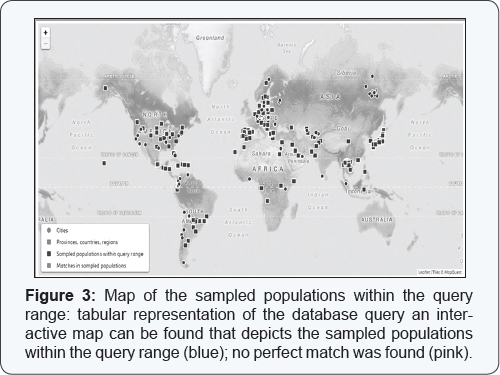

The following mtDNA haplotype was obtained: 16291A, 16326C, 58C, 64T, 146C, 152C, 263G, 315.1C; the data was subsequently analyzed in the EMPOP database (www.empop. org). The execution of a database query automatically directs the user to the results page: we obtained an interactive map that shows depicting the sampled populations within the query range in blue (Figure 3). No perfect match was found (pink). Haplogroup was estimated based on Phylotree and a curated database of full mtGenomes (currently approx. 20k haplotypes) using EMMA [12]. Haplotype names correspond to a haplogroup designation from Phylotree (e.g. H1b1+16362) and then represents a Phylotree branch [13]. mtDNA sequence queries often do not result in database matches, as in this case. Analysis of this haplogroup informative mtDNA polymorphisms indicates that the examined individual belong to haplogroup ROa'b (Rank 1), with no missing mutation, but with 3 private mutations (C16291A T146C T152C). Rank 2 indicated R0a. Rank 1 haplogroup estimates start with the haplotype with lowest cost to the query haplotype, rank 2 presents haplogroup estimates with the next lowest costs. Mouse-clicks on the haplogroup affiliations lead to the depiction of the geographical distribution of the haplogroup based on EMPOP mtDNA sequences (Figure 4) The haplogroup ROa'b is the most frequent of the Arabian Peninsula -specific haplogroups.
Discussion
The complete and multidisciplinary analyses of relics of Saints are rarely applied. In ancient skeletal remains, the quantitative and qualitative differences in results can be attributed to environmental factors or storage conditions. In addition, when dealing with religious relics, one is often limited to investigating a small sample. In this study, it was permitted only to use a small sample of femur that was further sub-divided into two pieces, one for radiocarbon dating and the other one for genetic analysis. Radiocarbon dating was utilized to determine the age of the bone; the results suggest that it originates from the 3rd century AD., the same era in which the martyr lived [1]. The genetic analysis was laborious: the degree of DNA preservation in this kind of sample can be primarily attributed to environmental factors and the type of materials that were in contact with the skeletal remains (such as dust, sand, stones, etc.) [14].
Further problems encountered in genetic analysis were quantity and quality of aDNA obtained: the low concentration for PCR purposes, the presence of PCR inhibitors as previously described [9,13] and DNA damage and fragmentation [15-17]. Early strategies to overcome these hindrances were to increase the volume of the Taq polymerase, in certain instances up to 10 U per PCR reaction [18,19]. A further strategy was to dilute the DNA extract, thereby diluting the co-extracted PCR inhibitors [20]. A wide range of techniques have been published to date, all of which aim to maximize DNA yields, while minimizing the co-extraction of PCR inhibitors [14,21]. Expensive commercial kits and specific equipment are often used in forensic and anthropology laboratories to achieve this. Satisfactory genetic profiles can be obtained when the DNA samples are well preserved, whereas in cases of poor preservation only partial profiles can be detected [22]. In addition, it is often difficult to obtain amplification products longer than 100bp from ancient specimens and bone materials that have been buried for a long time. In fact, new approaches have been implemented in this study such as NGS, shotgun sequencing, etc [23].
In this case we are able to obtain a complete STR panel and this could be achieved since the skeletal remains were buried for a short time and subsequently were preserved in a glass urn. Furthermore, a new and more user-friendly extraction method was utilized, which enabled recovery of longer and better preserved DNA fragments. Pre-treatment procedure as previously described was utilized prior to extraction, for removal of exogenous DNA. Moreover, treatment with EDTA allowed to obtain a complete demineralization of the bone sample. This method could be highly successful for the extraction and recovery of DNA profiles from degraded, old skeletal remains. Combined matching probability was utilized to conjecture the provenance of skeletal remains: in accord with radiological study, the bone sample appeared to belong from a European male. Finally a mitochondrial DNA analysis was conducted in order to obtain a haplotype in term of differences from the CR Sequences. The evolutionary rate of the control region is approximately ten times that of the gene-coding region, so variation is very much concentrated in HV1/HV2 [24]. These data shown as St. Fortunato's maternal ancestors may come from Arabian Peninsula. The Christian religion was born near this geographical region; following the death of Christ, his disciples preached the gospel to the world, especially in Europe. This is consistent with St. Fortunato's life, because the first preachers descend from this area.
This haplogroup is present in Spain, Greece, and Italy. In conclusion, this study has shown a multidisciplinary approach in identifying ancient human skeletal remains of a historical figure venerated as a Saint of the Catholic Church and analyzing the evidences or signs of traumatic death that can be related to the cause of the death in relation with the martyrdom practices. All the elements of identification, based on the initial postulate that the skeletal remains were potentially those of San Fortunato, were compatible with those of an Italian male, of height between 1.53 m and 1.56 m, with age at death estimated between 20 and 40 years. Despite the inability to perform a DNA matching test, there is a strong scientific concordance between these findings and the époque of the saint's existence, sex, and ethnicity. One cannot uncompromisingly attribute the skeletal remains to the saint, as is the case with the Catholic faith. However, the data are certainly compatible with the presumed era of St. Fortunato and despite the radiological analysis led to exclude any traumatic evidences at the bones level, there is high concordance between church interpretation and the other results. This case highlights the complexity of study of remains that were skeletonized. Therefore a multidisciplinary investigation as well as paying careful attention in the collection and correct interpretation of findings is important [25]. This study may offer a model that could be applied to poorly conserved organic remains both in historical studies and both in criminal cases with concealment of a corpse; furthermore, archaeological remains can provide concrete cases, making it possible to develop, refine or validate medico-legal techniques.
Acknowledgement
We thank Don Renato Orlando, parish priest of "St. Maria in Silvis e St. Mercurio" of Serracapriola (Italy), for financial contribution and for the opportunity to take part in this research.
References
- Adamo AS (1985) Fortunato Martire, Cenni Storici. Borrelli, Italy.
- Fabrice Dedouita, Giuseppe Guglielmi, Gabriela Perillif, Michelangelo Nasutod, Norbert Telmon, et al. (2014) Virtual anthropological study of the skeletal remains of St. Fortunato (Italy, third century AD) with multislice computed tomography. Journal of Forensic Radiology and Imaging 2(1): 9-16.
- Hellborg R, Skog G (2008) Accelerator mass spectrometry. Mass Spectrom Rev 27(5):398-427.
- Lindahl T (1993) Instability and decay of the primary structure of DNA. Nature 362(6422): 709-715.
- Gabriel MN, Huffine EF, Ryan JH, Holland MM, Parsons TJ (2001) Improved mtDNA sequence analysis of forensic remains using a "miniprimer set” amplification strategy. J Forensic Sci. 46(2): 247-253.
- Caramelli D, Vernesi C, Sanna S, Sampietro L, Lari M, et al. (2007) Barbujani G.Genetic variation in prehistoric Sardinia. Hum Genet 122(3-4): 327-36.
- Anderson S, Bankier AT, Barrell BG, de Bruijn MH, Coulson AR, et al. (1981) Sequence and organization of the human mitochondrial genome. Nature 290: 457-465.
- Tuross N (1994) The biochemistry of ancient DNA in bone. Experientia 50(6): 530-535.
- Pääbo S (1989) Ancient DNA: extraction, characterization, molecular cloning, and enzymatic amplification. Proc Natl Acad Sci U S A 86(6): 1939-1943.
- Haack K, Hummel S, Hummel B (2000) Ancient DNA fragments longer than 300 bp. Anthropol Anz 58(1): 51-56.
- Benschop CC, van der Beek CP, Meiland HC, van Gorp AG, Westen AA, et al. (2011) Low template STR typing: effect of replicate number and consensus method on genotyping reliability and DNA database search results. Forensic Sci Int Genet 5(4): 316-28.
- Röck AW, Dür A, van Oven M, Parson W (2013) Concept for estimating mitochondrial DNA haplogroups using a maximum likelihood approach (EMMA). Forensic Sci Int Genet 7(6): 601-609.
- Burger J, Hummel S, Hermann B, Henke W (1999) DNA preservation: a microsatellite-DNA study on ancient skeletal remains. Electrophoresis 20(8): 1722-1728.
- Rohland N, Hofreiter M (2007) Comparison and optimization of ancient DNA extraction. Biotechniques 42(3): 343-352.
- Hagelberg E, Clegg JB (1991) Isolation and characterization of DNA from archaeological bone. Proc Biol Sci 244(1309): 45-50.
- Handt O, Höss M, Krings M, Pääbo S (1994) Ancient DNA: methodological challenges. Experientia 50(6): 524-529.
- Parsons TJ, Weedn VW (1997) Preservation and recovery of DNA in postmortem specimens and trace samples. In: Marcella H Sorg, William D Haglund (Eds.), Forensic taphonomy: the postmortem fate of human remains. Boca Raton: CRC Press, USA, pp. 109-138.
- Lacan M, Keyser C, Ricaut FX, Brucato N, Duranthon F, et al. (2011) Ancient DNA reveals male diffusion through the Neolithic Mediterranean route. Proc Natl Acad Sci USA 108(24): 9788-9791.
- Knapp M, Lalueza-Fox C, Hofreiter M (2015) Re-inventing ancient human DNA. Investig Genet 6: 4.
- Nilsson M, Possnert G, Edlund H, Budowle B, Kjellström A, et al. (2010) Analysis of the putative remains of a European patron saint--St. Birgitta. PLoS One 5(2): e8986.
- Brown TA, Brown KA (1994) Ancient DNA: using molecular biology to explore the past. Bioessays 16(10): 719-726.
- Kemp BM, Smith DG (2005) Use of bleach to eliminate contaminating DNA from the surface of bones and teeth. Forensic Sci Int 154(1): 5361.
- Clio Der Sarkissian, Morten E Allentoft, Maria C Avila-Arcos, Ross Barnett, Paula F Campo, et al. (2015) Ancient genomics. Philosophical Transactions of the Royal Society Biological Sciences 370.
- Zimmermann B, Rock AW, Dur A, Parson W (2014) Improved visibility of character conflicts in quasi-median networks with the EMPOP NETWORK software. Croat Med J 55(2): 115-120.
- Pomara C, Gianpaolo DP, Monica S, Maglietta F, Sessa F, et al. (2015) "Lupara Bianca” a way to hide cadavers after Mafia homicides A cemetery of Italian Mafia A case study. Leg Med (Tokyo) 17(3): 192197.






























