Molecular Characterization of Protease Producing Stenotrophomonas and Their Cytotoxic Effects on Cancer Cell Lines
Anamika Thakur, Sanju Kumari, Utkarshini Sharma, Rohit Krishna, Kanak Sinha and Santosh Kumar*
Department of Botany, B R A Bihar University, India
Submission: September 24, 2018;Published: October 24, 2018
*Corresponding author: Santosh Kumar, Biotechnology Unit, Department of Botany, B R A Bihar University, Muzaffarpur 842001, India.
How to cite this article: Anamika Thakur, Sanju Kumari, Utkarshini Sharma, Rohit Krishna, Kanak Sinha, Santosh Kumar. Molecular Characterization of Protease Producing Stenotrophomonas and Their Cytotoxic Effects on Cancer Cell Lines. Int J cell Sci & mol biol. 2018; 5(2): 555660. DOI: 10.19080/IJCSMB.2018.05.555660.
Abstract
Proteases are highly required where proteins need some sort of alteration either structurally or functionally. They are widely used in household laundry, paper and leather industries as well as in several pharmaceutical compounds. Thus, the present study was confined to screen more than fifty isolates of microbes from the rhizospheres of fruit orchard for proteolytic activity on skimmed milk agar medium. Isolates SK701 and SK709 were found producing halo regions around the streaked mycelia. However, only isolate SK709 displayed maximum 64.6 U/ml protease activities in starch casein broth after 96 hours of culture at optimum conditions of 35°C and pH 7.6. It was identified as a gram-negative bacterium on the basis of staining, morphological, cultural, biochemical and physiological characteristics. The cytotoxic activity of cell free extracts of isolate SK709 against EAC, SiHa & Hep G2 cell lines in terms of IC50 was found to be 15.1, 26.3 and 35.5μg/ml, respectively. The 16S rRNA gene sequencing, molecular characterization and the BLAST operation as per NCBI directive to construct phylogenetic tree confirmed the putative protease producing isolate SK709 to be the species of Stenotrophomonas. The difference of 2% in the gene sequence with the closest member, Stenotrophomonas sp. MCYF1(KC734883.1) which amounted to 21 out of 1022 nucleotides, made the isolate SK709 qualified for obtaining a separate NCBI accession number KF484911..
Keywords: Stenotrophomonas;Protease; Cytotoxicity; EAC; SiHa and HepG2 cell lines; 16S rRNA gene sequencing
Introduction
Enzymes are biocatalysts obtained from plants, animals & microorganisms. Microbes are known for rich source of natural products with biomedical properties. They are important source of bioactive compounds especially antibiotics [1] and extracellular enzymes [2]. The ongoing research on microbes has brought a technical platform for industries by way of producing industrial enzymes viz., keratinases, pectinases, xylanases, amylases, lipases, phytases, cellulases and more prominently proteolytic or proteases (caseinases and gelatinases). Microbial exploitation for the production of these protease enzymes is highly attractive for applications in fruit, detergent, pharmaceuticals, paper industries and leather [3]. Microbes producing proteases are preferred for their easy isolation, limited space requirements and optimum temperature near 30°C [4]. The physical factors such as temperature, pH, incubation time, agitation, inoculums density and the availability of carbon and nitrogen source in the media also influence its production.
Proteases almost constitute 25-30% of total global enzyme sales [5]. In recent years many authors have characterized the microbial proteases as an important group of secondary metabolites for industrial purposes. Actually, the importance of proteases lies in its mode of action: it hydrolyses bonds of amino acids, either at terminal ends (exopeptidases) or in the middle of chain (endopeptidases) [6]. The other group of proteolytic enzymes, proteinases hydrolyses the secondary structure of protein. Thus, proteolytic enzymes are highly required where proteins need some sort of alteration either structurally or functionally [7], they actually destroy cell components such as lipoprotein membranes and immunoglobulins [8]. The cytotoxic study of the culture filtrate of Stenotrophomonas maltophila caused vigorous endocytosis in cancer cell lines like Vero and HeLa cells [9]. The function of proteases is largely pH dependent, so based on optimum pH, proteases have been recognized as acid proteases, neutral proteases and alkaline proteases [5,6]. Alkalophilic microorganisms such as Streptomyces, Micromonospora, Stenotrophomonas etc., are prone to produce alkaline proteases into extracellular media which are widely used in biotechnological processes.With this background the present study has been aimed to screen the isolated strains of microbes from the rhizosphere of Muzaffarpur (mainly from fruit orchards) for the production and purification of proteases and its characterization with respect to alteration in protein configuration to assess the cytotoxic effect on cancer cell lines.
Materials and Methods
Collection of soil samples
The soil samples were collected from rhizosphere of different locations of Muzaffarpur (26°7’N &85°25’E) district in Bihar, India. The soil samples were taken from 2-5 inches below the surface using sterilized spatula and zipped bags. The samples were kept first at 70°C for 15 minutes to evaporate moisture. It was then mixed thoroughly and sieved through a domestic 1.5mm pore size sieve to get rid of large debris.One gram of thus obtained sieved soil sample was suspended in conical flask having 10ml sterile distilled water. The flask was thoroughly shaken. It was then filtered through a two layered muslin cloth. Thereafter the sample was diluted to 10-5 dilutions and stored in sterilized test tubes at 4°C for 24h.
Isolation of microbes from soil samples
Aliquots (ml) of each dilution were spread on the surface of starch agar medium (Starch-9g, L-asparagine-9g , Ammonium sulphate -2g,Tris -2g, Sodium chloride –1g, Dipotassium sulphate -0.5g, Magnesium sulphate -0.2g, Calcium chloride -0.1g, Trace solution - 1ml, Potassium Dihydrogen Phosphate -0.5g, agar -15g, dissolved in one litre distilled water; pH -7.0) plates, supplemented with antifungal nystatin (50μg ml-1) and incubated at 35±2ºC for seven days. Plates with around 200 colonies were selected. Streaking of single colony was carried out to purify the selected colonies on the same medium. Gram staining smears were prepared and examined under the microscope to ascertain the nature of isolates.
Screening of proteolytic microbes
More than fifty isolates, purified from the soil samples were screened by streaking the mycelium on skim milk agar medium (Skimmed milk powder-100g, peptone -5g, agar-15g, dissolved in one litre Distilled water, pH 7.0) andinoculated at 35±2°C for 5-7 days. The culture plates showing transparent halo zone against an opaque non-hydrolysed medium indicated the protease producing strains. Proteolytic activities of the isolates were measured by diameter of clear zone around colony on skim milk agar plates and finally two isolates, SK701 and SK709 were selected for further investigation.
Cultural and morphological studies of proteolytic microbes
The colour of colony, aerial mycelium and soluble pigment of the selected isolates when grown on starch agar medium were recorded after 96h of incubation. Smears of isolates were prepared for gram staining and spore chain morphology studied under a compound light microscope (Nikon, Japan) using 1000× magnification power. Colour of the substrate mycelia (reverse of the plate) was also observed along with diffusible pigments. Various biochemical and physiological tests performed for the identification of the potential isolates were indole production, methyl red, Voges Proskauer, citrate utilization, casein hydrolysis, starch hydrolysis, urea hydrolysis, gelatin hydrolysis, H2S production, temperature tolerance, utilization of carbon and nitrogen sources and Sodium chloride resistance.
Optimization for protease production in broth culture
Optimum conditions for growth (optical densities at 540nm) and proteinase production of the selected isolates, SK701 and SK709 were carried out in starch casein broth (soluble starch: 10g, K2HPO4: 2g, KNO3: 2g, casein: 0.3g, MgSO4.7H2O: 0.05g, CaCO3: 0.02g, FeSO4.7H2O: 0.01g, Distilled water: 1litre, pH:7.0) by incubating on shaker at 35°C for 7 days. Effects of different incubation temperatures (25, 30, 35, 40, 45, 50, 55 and 60°C) and initial pH (4, 5, 6, 7, 8 and 9) of the medium on protease estimation just after 96 h were screened in broth. Effect of casein degradation on pH was also noticed on daily basis. Different carbon sources such as 1.0% glucose, sucrose, fructose, lactose, and mixture of glucose + sucrose, both at 0.5% were also tested. Nitrogen sources were 0.5% yeast extract, ammonium sulphate, sodium nitrate, urea and ammonium acetate. All the experiments were carried out in triplicate and averages were reproduced.
Assay for protease enzyme
The isolates, SK701 and SK709 with protease activities were allowed to grow starch casein broth medium at 35°C for 2 days on a rotary shaker at 150rpm. Then 1ml of such grown bacterial suspension was again cultivated in starch casein broth medium (40mL) in 250mL Erlenmeyer flasks for 7 days at 37°C at 200rpm. After due period, centrifugation of broth suspension was carried out at 10,000rpm for 15min and cell free supernatant was separated for protease activity assay. The method of Tsuchida et al. [10] for protease estimation was followed. 250ml of starch casein broth was prepared and inoculated with SK701 and SK709 separately and incubated at 37°C for 7 days in incubator shaker at 180rpm. After that the whole broth was centrifuged at 10000rpm at 4°C for 20min and the clear supernatant was taken as crude enzyme and was stored at 4°C. A mixture of 500μl of 1% (w/v) of casein in 50mM phosphate buffer (pH 7) and 200μl crude enzyme extract were incubated in a waterbath at 40°C for 20min. After that the enzyme reaction was terminated by adding 1ml of 10% (w/v) TCA and was kept at room temperature for 15min. The reaction mixture was centrifuged to separate the unreacted casein at 10,000rpm for 5min. The supernatant was then mixed with 2.5ml of 0.4M Na2CO3. 1ml of 3-fold diluted Folin and Ciocalteus Phenol reagent was added. The resulting solution was incubated at room temperature in dark for 30min and the absorbance of the blue colour, thus developed, was measured at 660nm against a reagent blank using tyrosine.Since SK709 had shown maximum proteolytic activity and thus selected for further investigation.
Extraction of bioactive compounds
10ml of overnight culture of the isolate SK709 was transferred into conical flask containing 500ml of Bennet medium (sterilized with steam explosion) individually and incubated for 7 days at 24°C at 200rpm and pH (7.0±0.2). Further the mass cultured broth was filtered through cheese cloth to get clear filtrate. Further, equal volume (1:1) of ethanol was added & mixed by vortexing & kept without disturbance. The organic phase was collected & evaporated at 70°C in an incubator. The obtained residue was stored at-20°C for cytotoxicity assay.
Cytotoxicity assay
EAC (Ehrlich Ascites Carcinoma), SiHa (Human cervical cancer) and HepG2 (Hepatic carcinoma) cell lines were procured from NCCS Pune, India. Cells were cultured and maintained in Dulbecco’s Modified Eagle’s Medium (DMEM) with 10% Fetal Bovine Serum and antibiotics in a humidified atmosphere of 5% CO2at 37°C in culture dishes/flasks. Stock culture was maintained in the exponential growth phase by passaging as monolayer culture using in 0.02% EDTA. The dislodged cells were suspended in complete medium and reseeded routinely.
The cytotoxic effects were assessed against EAC, SiHa and HepG2 cells at the varying concentrations of plant extracts by the MTT assay. The 3-(4,5-dimethyl-2-yl)-2,5-diphynyl tetrazolium bromide (MTT) is metabolic substrate which is reduced by the mitochondrial succinate dehydrogenase enzyme and forms formazan crystal. In brief cells were seeded overnight and then incubated with various concentration of extracts for 72h. At the end of the treatment, medium was removed, and cells were incubated with 20μl of MTT (5mg/ml in PBS) in fresh medium (50μl) for 4 hrs in CO2 incubator. After the treatment formazan crystal, formed by mitochondrial reduction of MTT were solubilised in DMSO (150μl /well) and the absorbance was read at 570nm after 10min incubation on the iMark Microplate Reader (Bio-Rad, USA). Percent cytotoxicity (IC50) values were determined by extrapolating the graph.
Molecular characterization
Genomic DNA extraction
Genomic DNA of SK709 was isolated using standard protocol [11] and purity of DNAwas checked A280/A260. Further bands were visualized by running on (0.8% w/v) agarose gel electrophoresis.
16S rRNA gene amplification and sequencing
16S rRNA gene of isolated genomic DNA of SK709 was amplified [12] in a PCR thermal cycler (Genetix) by universal primers, 9F (5’GAGTTTGATCCTGGCTCAG3’) & 1541R (5’AAGGAGGTGATCCAACC3’). The reaction mixture contained 2μg of dNTPs mixture (1.25mM each), 1μg of each of the primers, 2.5μg of DNA, 1μl of Taq polymerase and sterile deionised water to make final volume 100μl. PCR consists of an initial denaturation at 94°C for 1min, annealing at 63°C for 1min and 72°C for 1min & final 5min extension at 72°C. The PCR amplified product were stained with ethydium bromide and run on agarose (1.2%) gel electrophoresis and examined under gel documentation system. The complete 16S rRNA gene of SK709 was sequenced by using PCR products at Samved Biotech, Ahmedabad.
Construction of phylogenetic tree
The 16S rRNA gene sequence of SK709 were compared with other bacterial sequences by using NCBI BLAST programme. Multiple sequence alignment & phylogenetic tree were constructed using software programme MEGA 4.0 [13]. The 16S rRNA sequence of strain SK709 was submitted to GenBank (Nucleotide database of NCBI). The sequence was accepted and the accession number (KF849411) for SK709 was obtained.
Results
Isolation & Identification of protease producing microbes
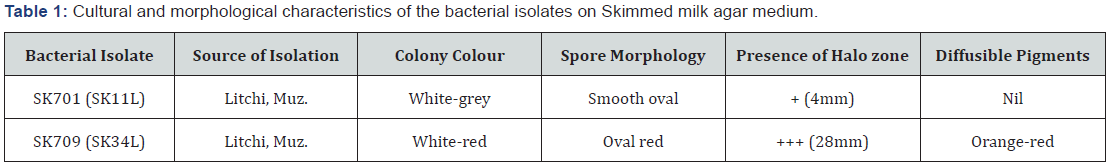
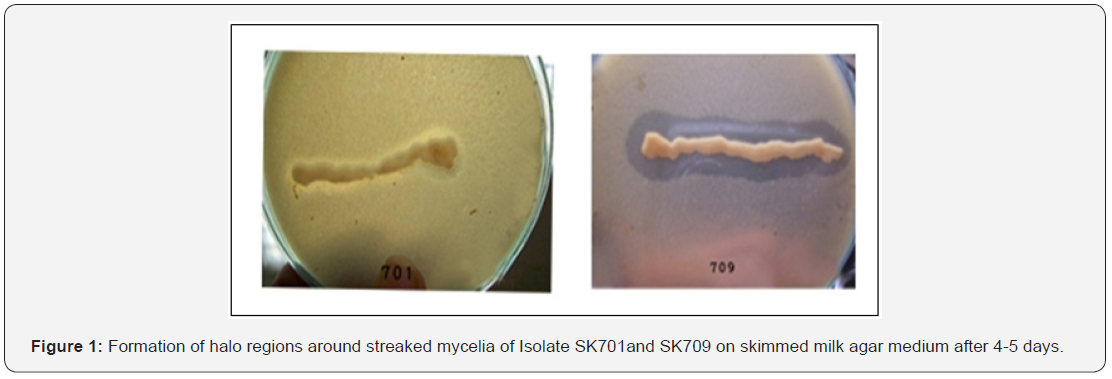
Bacterial strains, isolated from the rhizosphere of Muzaffarpur were grown on starch agar medium, supplemented with nystatin and were screened colony colour, spore morphology, [14,15] mycelia colour, etc. (Table 1). Two bacterial strains, SK701 and Sk709 of litchi orchard of Muzaffarpur were confirmed with proteolytic activities as they developed halo regions on Skimmed milk agar plates.But isolate SK709 was more prominent having approximately 28 mm clear halo zones around colonies or streaked mycelia on fourth day of incubation (Figure 1). Purified strains were examined under microscope for gram staining and Isolate SK709 was found to be gram negative whereas isolate SK701 with many others were found to be gram positive. Further both isolated reacted positively in different biochemical and physiological tests (Table 2). Sucrose was found to be the optimum source among carbon compounds and beef extract for protease production. The optimum growth of bacterial isolates was found at pH 7.8 and temperature at 35±2°C.
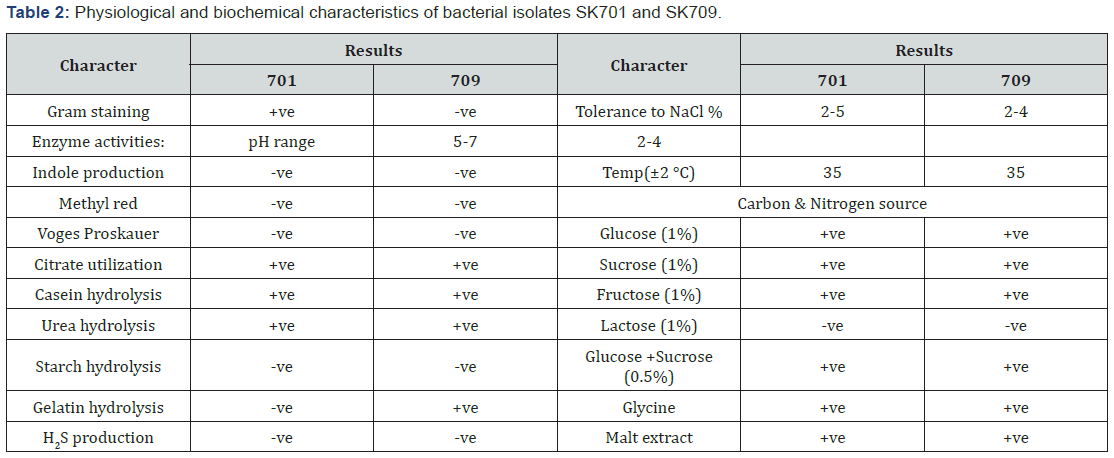
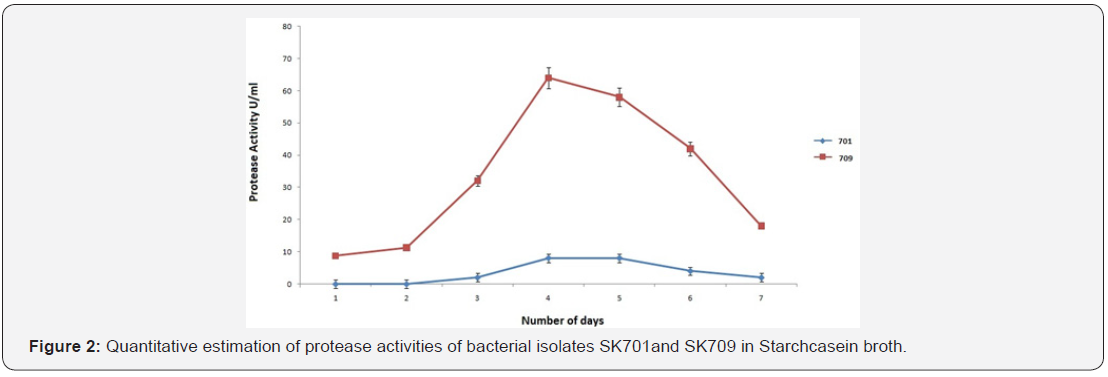
Proteolytic activities
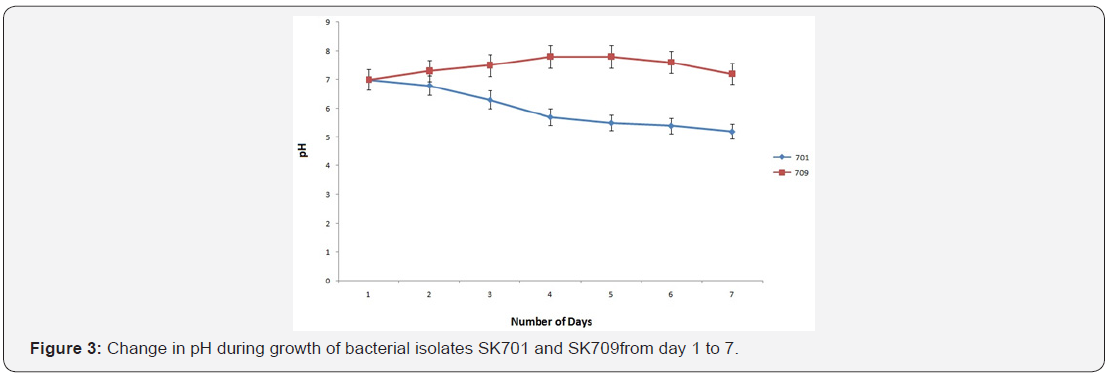
Growth of the bacteria and protease production were found to be maximum on 4th day of incubation. The casein-degradation activities by releasing proteinase was found maximum (64.6 U/ ml) on 4th day (Figure 2).The pH above 7.0 was required in case of gram negative isolate SK709 whereas pH with less than 7.0 favoured the growth of other bacterial strains including SK 701. The pH of the broth started to increase with the growth of isolate SK709, the value reached to 7.8 on 4th day after 96h of incubation (Figure 3). The effective temperature for maximum protease activities was noticed at 35±2°C. At high temperature (>45°C), the protease activities dropped to the level of 20% (Figure 4).

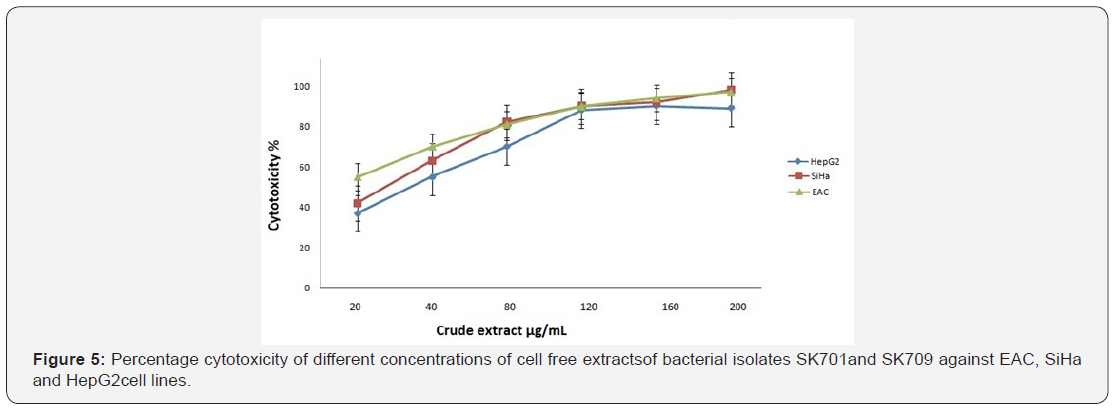
Cytotoxicity
The cytotoxicity effect of crude extract of isolate SK709 was assayed against EAC, SiHa and HepG2 cells at the varying concentration by MTT assay. The data of MTT assay were used to extrapolate IC50 of the extracts of these cell lines, which was found dose (200 μg/ml) and time dependent (72h). Among the three cancer cell lines (Figure 5), EAC showed the minimum concentration IC50value (17.2μg/ml) followed by SiHa (26.5μg/ ml) and Hep G2(35.5μg/ml).
DNA Amplification
The genomic DNA of the proteolytic isolate SK709 was isolated by standard protocol [16] and the samples were run on 0.8% agarose gel electrophoresis. Thereafter PCR amplification using universal primers resulted into a single discrete amplicon band of 1.5kb of 16S rRNA gene (Figure 6).

Molecular characterization of the selected isolate
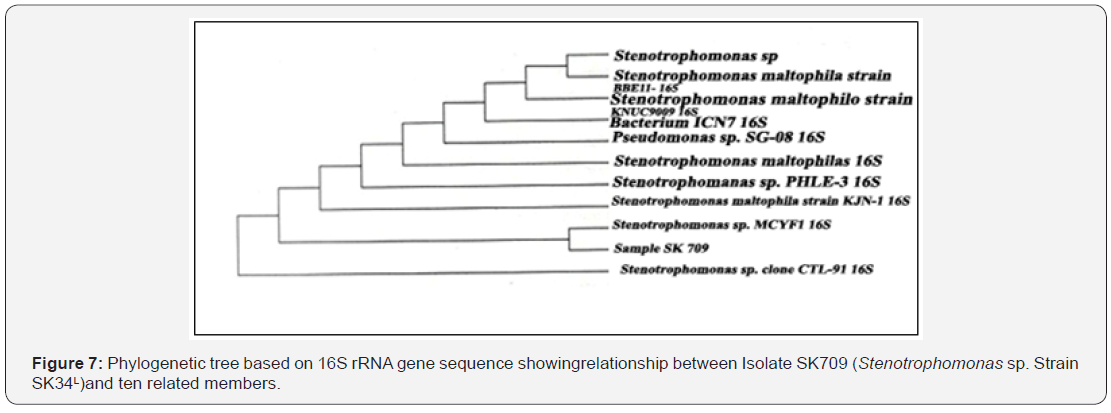
The NCBI BLAST homology analysis of 16S rRNA gene of SK709 showed maximum similarity (≥98%) with Stenotrophomonas sp. Based on the result, phylogenetic tree was constructed using neighour joining method (Figure 7).Thesequence (Table3) was published after trimming and assembling to NCBI(http//: www. ncbi.nlm.nih.gov). It was revealed that the strain SK709 showed 1001 base similarity out of 1022 (98%) with Stenotrophomonas sp. MCYF1(NCBI Accession no. KC734883.1). Thus, the difference of 21 nucleotides in the 16S rRNA gene qualified the strain SK 709 to earn a separate NCBI Accession no. KF484911.

Discussion
Microorganisms are usefulness in food and dairy industries, pharmaceutical sectors, leather industries etc. They produce a variety of bioactive substances and secondary metabolites. Nowa- days, they are an important producers of industrial significant enzymes viz. protease, amylase, phytase, cellulase, lipase, etc. [17,18]. Proteases are one of such novel enzyme that is useful in day-to-day activities. Most of the bacterial strains were isolated on starch agar medium. In order to obtain a specific byproduct, the medium requires some modifications. Thus, a specific protease screening medium (skim milk agar medium) was used for screening casein degrading isolates. Cultural, morphological, physiological, biochemical properties revealed the strains SK701 & SK709, isolated from rhizosphere of litchi orchard of Muzaffarpur to be the species of genera Streptomyces (actinomycetes group) and Stenotrophomonas (Pseudomanad group), respectively. The generic status of these strains SK701 and SK709 was not only confirmed by physiological and cultural characterization but also by the Burgey’s manual of Determinative Bacteriology [16,17]. However, the final support of the recognition Stenotrophomonas was established on the basis of phylogenetic tree constructed using 16S rRNA gene sequencing [19,20].
The proteolytic activity in broth was observed after 96h of incubation during the stationary phase. The protease activity was optimum at 35°Cand showing maximum activity pH 7.8, confirming alkaline nature of protease enzyme [21]. Different alkaline protease producing alkalophilic strains have been reported [5].Alkaline proteases showed better resistance to alkali and other denaturing chemicals in the reaction mixture & have higher affinity towards proteinaceous substrates thus it could be proved as a very good detergent supplement. The organism showed optimum growth near 35±2°C indicating its mesophilic nature. In present investigation optimum protease production was obtained by combination of sucrose (carbon source) & beef extract (nitrogen source).
In present study an attempt was made to find cytotoxic activity from the extract of Stenotrophomonas sp. Earlier Figueiredo et al. [9] have reported cytotoxic activity in Stenotrophomonas maltophila. The bacterial isolate showed IC50 value 17.2μg/ml, 26.5μg/ml and 35.5μg/ml in EAC, SiHa and Hep G2 cancer cell lines, respectively. Suffness et al.[22], have reported that the IC50 values less than 30μg/ml in cancer cell lines can be considered as potent for anticancer drug development[23]. This report augurs that the extract of SK 709 may prove a beneficial chemical to fight against cancer and the protease may have the ability to make alteration in protein configuration in cancer cell lines leading to apoptosis and thus hindering the growth of tumour cells.
Conclusion
The isolate SK709 with proteolytic activity on skimmed milk agar medium was identified as Stenotrophomonas on the basis of morphological, cultural, biochemical and physiological characteristics as well as 16 S rRNA gene sequencing. On the basis of 98% blast homology data base similarity with Stenotrophomonas sp.MCYF1(KC734883.1), the isolate SK709 was christened as Stenotrophomonas sp strain SK 34L and because of the difference of 21 nucleotides which amounts to 2%, the strain earned a fresh accession no. KF849411.
Conflict of interest statement
The authors have no conflict of interest associated with this study.
Acknowledgements
Authors are thankful to the Head of the Department of Botany for providing proper facilities to work. SKas Project Fellows and US as as P I, DST WOS-A,are thankful to the University Grants Commission, New Delhi for granting financial support under UGC SAP (DRS-Phase 1) scheme to the University Department of Botany, B R A Bihar University Muzaffarpur (File No. F.3-13/2009- SAP-II) and to the Department of Science and Technology, New Delhi (File No. SR/WOS-A/LS-289/2011), respectively.
References
- Sethi S, Kumar R, Gupta S (2013) Antibiotic production by microbes isolated from Soil.International Journal of Pharmaceutical Sciences and Research 4: 2967-2973.
- Cherry JR, Fidants AL (2003) Directed evolution of industrial enzymes: an update. Curr Opin Biotechnol 14: 438-443.
- Jeyadharshan VN (2013) Production & partial purification of protease by actinomycetes species. International Journal of Scientific & Research Publications 3(4):1-3.
- Jain R, Agrawal SC, Jain PC (2008-2009) Proteolytic actinomycetes from Indian habitats. Journal of Culture Collection 6:28-37.
- Mehta VJ, Thumar JT, SinghSP (2006) Production of alkaline protease from an alkalophilic actinomycete. Bioresource Technology 97:1650- 1654.
- Guravaiah M, Daniel K, Prabhakar T, Hatti I, Sirisha P(2012) Screening, isolation, characterization and strain improvement of Streptomyces indicus GAS-4. Indian Journal of Science 1:77-83.
- Mathew CD, Gunathilaka (2015) Production, purification and characterization of a thermostable alkaline serine protease fromBacillus lichniformisNMS-1. International Journal for Biotechnology and Molecular Biology 6: 19-27.
- Edberg SC, Gallo P, Kontnick C (1996) Analysis of the virulence characterstics of bacteria isolated from bottled water cooler and tap water. Microbial Ecology in Health and Diseases 9:67-77.
- Figueiredo PMS, Furumura MT, Santo AM, Sousa ACT, Kota DJ(2006) Cytotoxic activity of clinical Stenotrophomonas maltophila. Applied Microbiology 43:443-449.
- Tsuchida O, Yamagata Y, Ishizuka J (1986) An Alkaline proteinase an alkalophilic Bacillus sp. Current Microbiology 14:7-12.
- Wang TY, Wang L, Zhang JH, Dong WH (2011) A simplified universal genomic DNA extraction protocol suitable for PCR. Genetics Molecular Research10:519-525.
- Olmezoglu E, Herand BK, Oncel MS, Tune K, Ozkan M (2012) Copper bioremoval by novel bacterial isolates and their identification by 16S rRNA gene sequence analysis. Turkish Journal of Biology36:469-476.
- Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA 4 molecular evolutionary genetics analysis (MEGA) software version 4.0. Molecular Biology Evolution 24(8): 1596-1599.
- Phridham TG, Tresner HD (1974) Genus I Streptomyces. In: Waksman & Henrici, Buchnan RE, Gibbons NE (Eds.), Bergey’s Manual of determinative bacteriology (8th edn), Williams & Wilkins, Baltimore, UABaltimore, USA,pp. 748-829.
- Holt JG, Krieg HR, Sneath PHA, Staley JT, WilliamsST(1994) Bergey’s manual of determinative bacteriology,(9thedn). Lippincott Williams & Wilkins, Baltimore, USA.
- Kieser T, Bibb MJ, Buttner MJ, Chater KF, Hopwood DA (2000) Practical StreptomycesGenetics. The John Innes Foundation, Norwich, England.
- Sharma U, Kumari S, Sinha K, Kumar S (2017) Isolation and molecular characterization of phytase producing actinobacteria of fruit orchard. The Nucleus 60:187-195.
- Kumari S, Sharma U, Krishna R, Sinha K, Kumar S (2018) Screening and molecular characterization of cellulase producing actinobacteria from Litchi Orchard. Current Chemical Biology 12(2).
- Saitou N, Nei M (1987) The neighbor-joining method: a new method for constructing phylogenetic trees. Mol Biol Evol 4: 406-425.
- Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences J Mol Evol 16:111-120.
- Viswanathan K,Rebecca LJ,Arumugan P (2017) Optimization of protease enzyme production by marine actinomycetes. International Journal of Pharmacy and Biological Sciences 8: 188-194.
- Suffness M, Pezzuto JM (2009) Assay related to cancer drug discovery, In:Hostettemank (Edt.), Methods in plant biochemistry: assay for bioactivity. Academic Press, London, England, pp. 71-133.
- Ravikumar S, Fredimoses M, Gnandesigan M (2012) Anticancer property of sediment actinomycetes against MCF-7 & MDA-MB-231 cell lines. Asian Pac Jof Trop Biomed 2: 92-96.






























