Identification and Analysis of Salt Responsive Candidate Gene Based on SSRs from Tomato (Solanum lycopersicum L.)
Jaafar Umar*1, Aliero Adamu Aliyu2, Shehu Kasimu1 and Abubakar Lawal3
1Department of Biology, Federal University, Nigeria
2Department of Biological Sciences, Usmanu Danfodiyo University, Nigeria
3Department of Crop Science, Usmanu Danfodiyo University, Nigeria
Submission: August 09, 2018;Published: August 30, 2018
*Corresponding author: Jaafar Umar, Department of Biology, Federal University, Birnin Kebbi, Nigeria.
How to cite this article: Jaafar U, Aliero A A, Shehu K, Abubakar L. Identification and Analysis of Salt Responsive Candidate Gene Based on SSRs from Tomato (Solanum lycopersicum L.). Int J cell Sci & mol biol. 2018; 5(1): 555653. DOI: 10.19080/IJCSMB.2018.05.555653.
Abstract
Salinity inhibition of plant growth is the result of osmotic and ionic effect and different plant species have developed different mechanisms to cope with those effects. With the discovery of molecular markers and marker assisted selection technology, it is possible to develop markers that identify salt tolerance. This study aimed at identification salinity responsive gene based SSRs. In this study, genetic diversity of twenty tomato genotypes (landraces/accessions in Nigeria) DNA of the 20 accessions were isolated using Bioland Plant Genomic DNA protocols, Primers were designed from 15 different salt responsive candidate genes. All 15 primers sets generated shows clear distinct polymorphic profiles as evident from the 6% agarose gel profile. Data obtained were analyzed using SSRs polymorphic markers and Unweighted Pair Group Method with Arithmetic Mean. The lowest polymorphic information content (PIC) value of 0.01 was exhibited from AI486387.1 gene while highest value of 0.903 was obtained from AI778183.1 gene with tri-nucleotide motif. The dendogram generated shows two clusters, one containing only the accessions and the other one containing both landraces and accessions. In conclusion, the present study represents the potential of salt responsive candidate gene based SSR (cgSSR) markers to be utilized as remarkable candidate for diversity analysis among tomato genotypes differing in salinity response and the genetic distance information reported in this study might be used by breeders when planning future crosses among tomato genotypes.
Keywords: Tomato; Salinity; DNA; Polymorphic information content; Candidate gene
Introduction
Tomato (Solanum lycopersicum L.), belong to the Solanaceae family which is one of the most important vegetables being widely grown in both fields and under protected cultivation. Most tomato cultivars are sensitive to moderate levels of salinity [1]. Indeed, all plant development stages, including seed germination, vegetative growth and reproduction, show salinity sensitivity, that leads to poor harvests and reduced economic yield [2]. Tomato is considered as a vegetable model and has thus been subjected to molecular investigation resulting in abundant genomic information. In addition to its worldwide agricultural and economic importance as a crop, tomato is a pre-eminent model system for genetic studies in plants. The use of molecular markers in breeding by means of marker assisted selection (MAS) could improve performance under extreme environments [3].
Tomato (Solanum lycopersicum L.), a major horticultural crop consumed all over the world, suffers heavy losses due to salinity. USP (universal stress protein) family proteins, first identified in prokaryotes, appear to play an active role in abiotic stress response, but their function remains largely unknown in plants [4]. A USP gene (SpUSP), cloned from wild tomato (S. pennellii) and functionally characterized in cultivated tomato exhibited increased expression under dehydration stress, salinity, oxidative stress and phyto-hormone ABA treatment. With the discovery of molecular markers and marker assisted selection technology, research has entered in to a new era and has made it possible to develop new and more informative PCR-based markers, including simple sequence repeats (SSRs), and to further facilitate the use of markers in tomato breeding. Genomic microsatellite markers are an elite group of markers, but there is possible uncertainty of linkage with the important genes. In contrast, there are better possibilities of linkage detection with important genes if SSRs are developed from candidate genes [5].
Deoxyribonucleic acid (DNA) polymorphisms provide a powerful tool for quantifying the existing levels of genetic variation in plant germplasm [6]. Molecular markers can provide an effective tool for efficient selection of desired agronomic traits because they are based on the plant genotypes and thus, are independent of environmental variation. It is suggested that the variation or polymorphism of SSRs are as a result of polymerase slippage during DNA replication or unequal crossing over [7]. SSRs are not only very common, also are hyper variable for numbers of repetitive DNA motifs in the genomes of eukaryotes [8,9]. Development of SSR markers based on QTL or candidate genes related to an important agronomic trait is useful in marker-assisted breeding programs for the concerned trait. In line with this, SSR markers were combined with morphological traits to assess the genetic diversity of cultivated and wild tomatoes [10]. The use of molecular markers can facilitate tomato breeding by means of marker assisted selection (MAS) to improve agronomical important traits such as yield, fruit quality, and disease resistance.
Materials and Methods
Plant materials plant materials
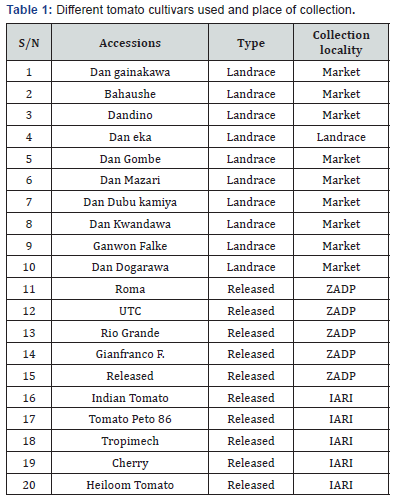
The seeds of the selected landraces tomato accessions were obtained from local markets around Sokoto and Zamfara metropolis and the released cultivars seeds were obtained from Zamfara State Agricultural Development Project, Gusau and Indian Agricultural Research Institute (IARI), New Delhi. A total of 20 genotypes of tomato grouped into landraces and released were used for the diversity analysis. The collection locality, type and common name of each cultivar are summarized in Table 1.
Isolation of genomic DNA
The DNA was isolated following a protocol of Bioland Plant Genomic DNA. Fresh green leaves were collected from twenty selected tomato plant samples and weighed (100mg), in 2.0mL micro centrifuge tube and immediately 600μl Buffer PL. 1 was added. The samples were then incubated at 65 °C for 10 minutes. The samples were mixed twice during incubation by vertexing the tube, 2μl of RNase A to remove RNA. Thereafter, 140μl Buffer PL. 2 was added and mixed by vertexing for 10 seconds and centrifuge at 11,000 rpm for 10 minutes. The supernatant was carefully transferred to a clean 2.0mL tube. Then 0.7 volume of isopropanol was added and mixed by vertexing for 5 second and centrifuge at 12,000rpm for 2 minute to precipitate the DNA. The supernatant was carefully decanted and discarded. Moreover, 300μl of elution Buffer (Pre heated) was added and vortexed for 10 second to mix the DNA well. Then 150μl of Buffer PL. 3 and 300μl of ethanol ware added and mixed by vertexing for 5 second, the samples were transferred into column and centrifuge at 11,000rpm for 1 minute. Therefore, 600μl of DNA washed Buffer was added and centrifuge at 11,000rpm for 20 second and the flow through was discarded. The column was transferred to a clean 1.5mL tube and pre-warmed (65 °C) 100μl elution buffer was added and immediately centrifuged at 11,000rpm for 1 minute to elute the DNA. The eluted DNA was added back to the column for second elution.
Retrieval of salt tolerant gene sequences, simple sequence repeats detection and primer design
Nucleotide sequences conferring salt tolerance in tomato were downloaded from National Center for Biotechnology Information (NCBI). The downloaded nucleotide sequences were used to mined simple sequence repeats [11,12]. The gene sequences were used to mined SSRs in SSR identification tool [13]. Respective references of those candidate genes which have been found to contain microsatellite repeats were used. Primers was designed manually with the following parameters: primer length 18-30bp, melting temperature 50-60 °C, GC percentage 40-60 and product size- 160-500bp using Vector NTI software.
Pcr amplification and 3% agarose gel electrophoresis
PCR amplification was perfumed on 20 genotypes with 15 pairs of SSR primers in a total volume of 25μl using a C1000 Thermal Cycler (Bio Rad, USA). Each 25μl volume of reaction mixture contained 50ng of genomic DNA as template, 1X Taq polymerase buffer, 2mM MgCl2, 0.2mM dNTPs mix, 0.4pM each of the forward and reverse primer, 1 U of Taq polymerase. The optimized condition was initial 5 minutes incubation at 97 °C for complete denaturation, followed by 38 cycles consisting of 94 °C for 1min, 55 °C- 60 °C (vary with the primer pair) for 1min, 72 °C for 2min and finally 72 °C for 10min. The experiments were repeated twice. Resolving of all PCR products were performed in a vertical non-denaturing 3% Agrose gel electrophoresis system at constant 90V with 1X TAE (Tris acetate EDTA) buffer (pH- 8.0). The gel was stained with ethidium bromide solution and visualized in gel documentation system (Protein Simple, USA) according to [14].
Allele scoring
Molecular weights of the amplified bands were determined by the number of base pair were multiplied by the average molecular mass of one base pair (660g/mol) to get the approximate mass of the whole double-stranded DNA molecule. Molecular weights of the amplified bands were determined based on the relative migration of standard 100bp DNA ladder (Thermo Scientific, USA) in the gel. Presence and absence of a particular allele was denoted as 1 or 0 respectively. Allele exclusively found in one genotype, it was designated as unique allele, in less than 5% of genotypes were designated as rare [11].
Data analysis
Polymorphism information content (PIC) value of each primer pairs was calculated according to the formula: PIC = 1- Σ pi-2, where pi is equal to the frequency of the allele of a particular locus [14] The dendrogram was generated using Unweighted pair group method analysis (UPGMA).
Result
Allelic variation among the polymorphic simple sequence repeat loci
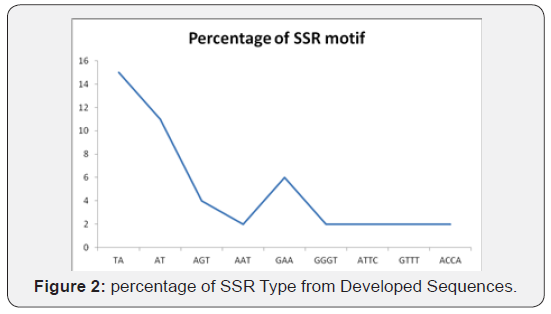
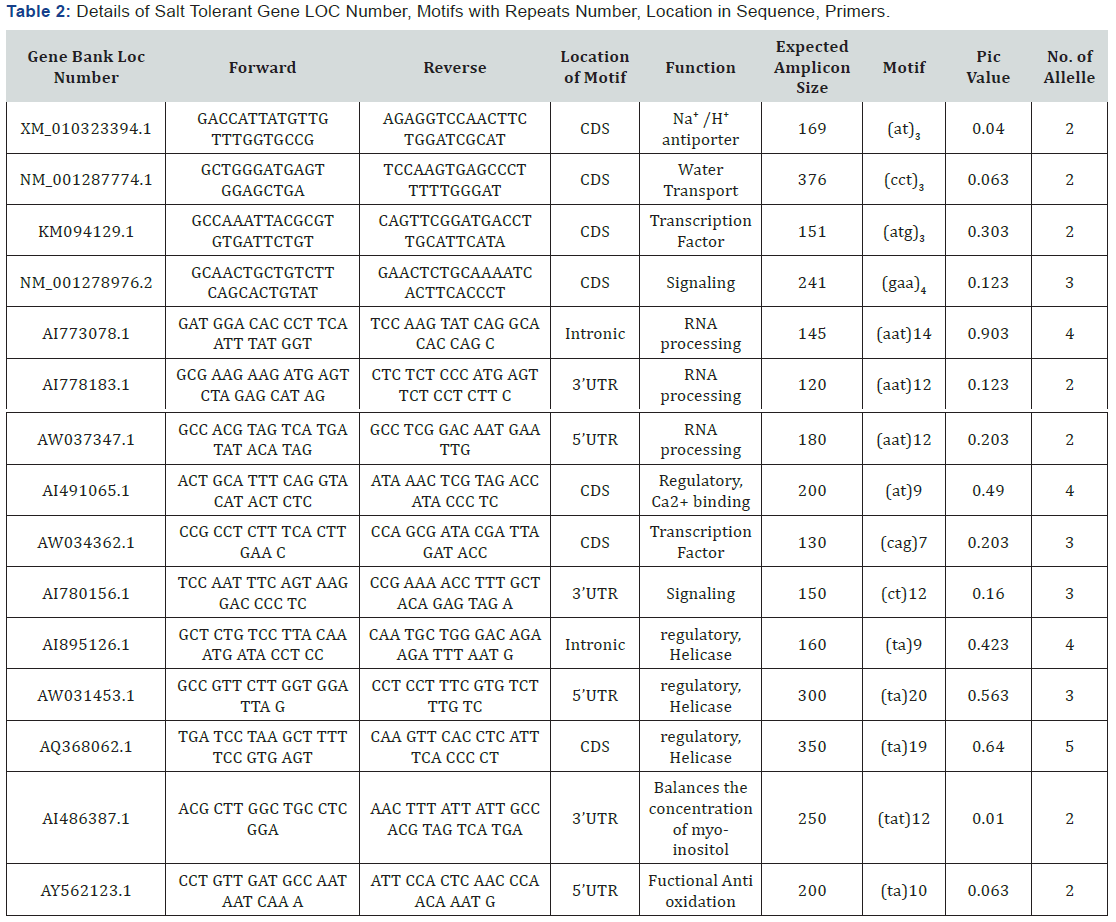
A total of 144 alleles were detected including 2 rare alleles with no unique allele. The cgSSR from XM_010323394.1 gene produced the lowest number of 4 alleles, followed by NM_001287774.1, AI486387.1, AY562123.1 with 5 alleles. The cgSSR from AI773078.1 gene gave rise to the highest number of alleles (19). In this research, only di-tetra nucleotide repeats and reiteration of motifs less than 5 times was excluded. Di -nucleotide motifs were found to be the largest with 175 SSR loci and tetra-nucleotide motifs formed the smallest group with 48 SSR loci (Figure 1). List of those genes with SSR loci with their respective gene bank LOC number, function, number, types and location of motif found were detailed in Table 2. Out of the different kinds of motifs found, TA motifs (38.9%) were most frequent, followed by AAT (25%) and AT, TAT and CT (8%) motif (Figure 2).

Type of simple sequence repeats, number of repeats and number of nucleotide per repeat
The sequences from the twelve randomly selected alleles were inserted in to simple sequence repeats identification tools, to identify and analyzed the SSR types, number of number of repeats and number of nucleotide per repeats. To validate the present gene with the original candidate genes. Di-nucleotide recorded the highest number of 78 repeats followed by trinucleotide with 55 repeats and tetra-nucleotide with 5 repeats. The highest percentage of 15 SSR motif was found in TA followed by AT with 11 SSR motifs. The lowest percentage of 2 SSR motifs was found in GTTT, ATTC, TTTG and ACCA respectively (Figure 2).

Simple sequence repeats markers characterization based on salt responsive gene
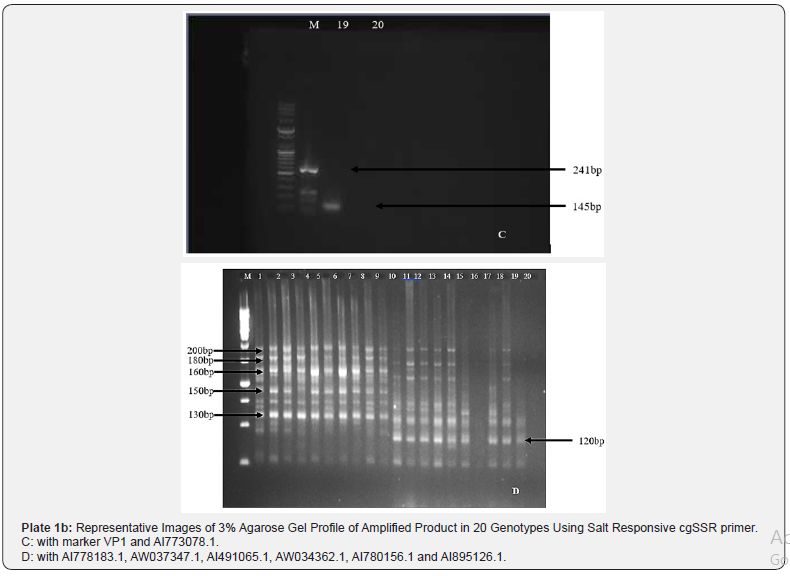
All the 15 different primer sets amplified to different respective cgSSR loci among tomato genotype and were used in the analysis of genetic diversity among the selected tomato genotypes (10 accessions and 10 landraces genotypes). Primer sets generated clear distinct polymorphic profiles as evident from the 3% agarose gel profile (Plate 1a & 1b). A total of 144 were detected by these polymorphic markers and the PIC value denotes the allelic diversity and frequency among genotypes. The lowest PIC value of 0.010 was exhibited by the cgSSR from AI486387.1gene, while highest value of 0.903 was obtained with the cgSSR from tri-nucleotide motif of AI773078.1 gene. Primer designed from di-nucleotide motif loci of XM_010323394 had lower PIC value of 0.040. Details of primers and their corresponding PIC values were presented on Table 2.
Genetic diversity analysis based on SSR
The data matrix generated from 15cg SSRs profiling of 20 genotypes were utilized to study the genetic diversity by cluster analysis. The dendrogram generated through unweighted pair group method of arithmetic mean (UPGMA) showed the similarity among the tomato genotype. The dendrogram exhibited four distinct clusters, interestingly none of the genetypes from different panel (i.e accessions and landraces) intermixed with one another (Figure 3). It was observed that dandino, dan kwandawa, dan mazari, dan eka and dan dubukamiya having same collection locality formed a separate subgroup. However, the same pattern of distinct subgroup was obtained under a subgroup with UTC and UC82B accessions. It is distinct from the genetic diversity analysis using the 15 cgSSR markers that those markers are able to distinguish tomato genotypes on the basis their genetic diversity based on salinity responsive genes (Figure 3).
Discussion
Allelic variation among the polymorphic simple sequence repeat loci of salt responsive gene
An average polymorphic information content of 0.287 and allelic variation observed might be correlated with the number of repeats within a particular microsatellite locus. Smulder et al. [15]and He et al. [16] reported a positive relationship between the number of repeats and the PIC value of tomato screened using SSR. The gene AI486387.1 had lower PIC (0.01) with 12 repeats compared to AI780156.1 which has PIC of 0.16 with 12 repeats. Similar observation was reported by He et al. [16] with genes with the same number of repeats but has different PIC value in Lycopersicum esculentum cultivars. Most of the tri- nucleotide motifs were found in CDS, in this respect, the result of this study is in agreement with an earlier report in inbred tomato with 57% tri-nucleotide found in CDS followed by 5´UTR with 21% (39). The phenomenon of conspicuous of tri-nucleotide repeats in CDS could be attributed to the selection pressure against frame shift mutation in coding regions resulting from length changes in non-triplet repeats [17]. In this study, TA repeat was the most-frequent type of SSR in the genotype, followed by AAT repeat. Wang et al. [18] reported the most-frequent type of microsatellite repeat as the AT/TA repeat. This shows that AT/ TA motifs are most frequent in tomato salinity responsive gene; in this regard breeding tomato with this motif may alleviate the effect of salinity in tomato production.

Phylogenetic analysis of tomato using simple sequence repeats markers
In the phylogenetic analysis, most of the tomato landraces and accessions were clustered together in respect to their genetic variation in response to salinity responsive gene and might have a similar genetic background. Those clustered within the same group or subgroups are mostly from the same origin and those, which are distantly grouped, might be genetically distinct. The relationship was also observed in similarity of the landraces genotypes in terms of their growth habit. In group IV, Bahaushe, Dandino and Dan eke are from the same location and exbited same growth habit. Similar result was reported on tomato. Cultivars from same geographical locations were group in a cluster of the dendrogram.
Conclusion
However, SSR based dendrogram showed clear relationship among the two panels (i.e accessions and landraces) in group I. interestingly, none of the landraces intermixed with the accessions in a sub group. This could help to improve genetic diversity analysis in tomato and the markers obtained could be used in a wide range of identification and pre-screening for salinity responsive gene in tomato.
References
- Singh J, Sastry EVD, Singh V (2012) Effect of salinity on tomato (Lycopersicon esculentum Mill.) during seed germination stage. Journal of Plant Physiology and Molecular Biology 18: 45-50.
- Zhang X, Lu G, Long W, Zou X, Li F, et al. (2014) Recent progress in drought and salt tolerance studies in Brassica crops. Journal of Science Breeding 64: 60-73.
- Bohnert HJ, Nelson DE, Jensen RG (1995) Adaption of Environmental Stresses. Plant and Cell 7: 1099-1111.
- James C (2006) Global Status of Transgenic Crops, Various Global Review Briefs from 1996 to 2006. International Service for the Acquisition of Agri-Biotech Application (ISAAA).
- Munns R (2002) Comparative physiology of salt and water stress. Journal of Plant Cell Environment 25: 239-250.
- Benor S, Zhang M, Wang Z, Zhang H (2008) Assessment of genetic variation in tomato (Solanum lycopersicum L.) inbred lines using SSR molecular markers. Journal of Genetics and Genomics 35: 373-379.
- Levinson G, Gutman GA (1987) Slipped-strand mispairing: A major mechanism for DNA sequence evolution. Journal of Molecular Biology and Evolution 4: 203-221.
- Hamada H, Petrino MG, Kakunaga T, Seidman M, Stollar BD (1984) Characterization of genomic poly(dT-dG) poly(dC-dA) sequences: Structure, organization and conformation. Journal of Molecular and Cell Biology 4: 2610-2621.
- Van der Schoot J, Pospiskova M, Vosman B (2000) Development and characterization of microsatellite markers in black poplar (Populus nigra L.). Theoretical and Applied Genetics 101: 317-322.
- Zhou H, Hanson T, Zhang J (2016) Generalized accelerated failure time spatial frailty model for arbitrarily censored data. Lifetime Data Anal 23(3): 495-515.
- Ali Molla K, Ananda BD, Showkat AG, Tapan KM (2015) Identification and analysis of novel salt responsive candidate gene based SSRs (cg SSRs) from rice Oryza sativa L.) BMC Plant Journal of Biology 15: 122- 130.
- Kawahara Y, De la Bastide M, Hamilton JP, Kanamori H, Mc Combie WR (2013) Improvement of the Oryza sativa Nipponbare reference genome using next Generation Sequence and Optical Map Data. Asian Journal of Agricultural Research 6(1): 4-14.
- Temnykh S, De Clerck G, Lukashova A, Lipovich L, Cartinhour S, et al. (2014) Computational and Experimental Analysis of Microsatellites in rice (Oryza sativa L.) Frequency, Length variation, Asian Journal of Molecular Biology 31: 31-40.
- Botstein D, White L, Skolnick M, Davis RW (1980) Construction of a Genetic Linkage Ma using restriction Fragment length Polymorphisms. American Journal of Human Gene 32(3): 314-31.
- Smulders MJM, Bredemeijer G, Rus Kortekaas W, Arens P, Vosman B (1997) Use of Short Microsatellites from Database Sequences to Generate Polymorphisms Among Lycopersicon esculentum Cultivars and Accessions of other Lycopersicon species. Theoretical and Applied Genetics 97: 264-272.
- He C, Poysa V, Yu K (2003) Development and Characterization of Simple Sequence Repeat (SSR) Markers and their Use in Determining Relationships among Lycopersicon esculentum cultivars. Theoretical and Applied Genetics 106: 363-373.
- Metzgar D, Bytof J, Wills C (2000) Selection Against Frameshift Mutations Limits Microsatellite Expansion in Coding DNA. Genome Resources 10: 72-80.
- Wang Z, Weber JL, Zhong G, Tanksley SD (1994) Survey of plant short tandem DNA repeats. Theoretical and Applied Genetics 88: 1-6.






























