DNA: Damage and Repair Mechanisms in Humans
Shrinivas Sudhir Ambekar*, Shanta Shrisel Hattur and Prajakta Budha Bule
Government College Of Pharmacy,Maharashtra India
Submission: June 27, 2017; Published: July 27, 2017
*Corresponding author: Shrinivas Sudhir Ambekar ,Shanta Shrisel Hattur and Prajakta Budha Bule, Government College Of Pharmacy,Maharashtra, India, Tel: 7507599947; Email: ambekarshree.ambekar@gmail.com
How to cite this article: Shrinivas S A ,Shanta S H, Prajakta B B. DNA: Damage and Repair Mechanisms in Humans. Glob J Pharmaceu Sci. 2017; 3(2): 555613. DOI: 10.19080/GJPPS.2017.03.555613
Abstract
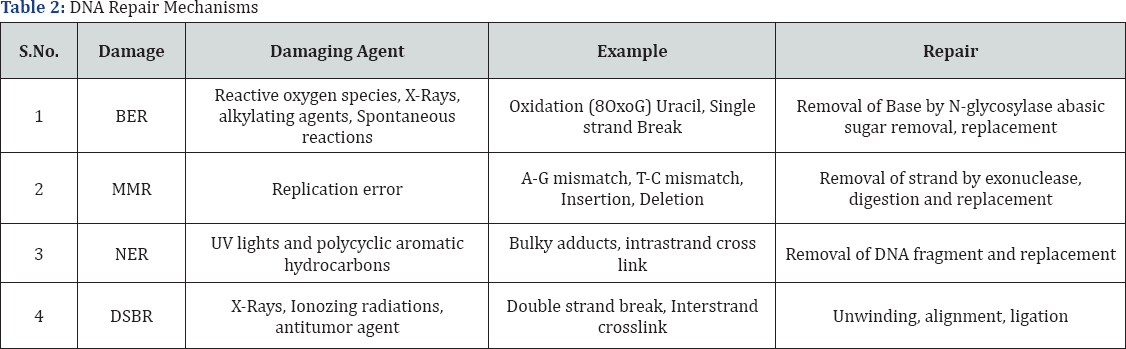
Carrier of genetic instructions used in development, functioning and reproduction of all leaving organisms is a macromolecule called DNA. Alteration in chemical structure of DNA is DNA damage. It is due to two types of factors-Endogenous and Exogenous. DNA impairment affects reproductive system, cardiovascular system, lymphatic system, Nervous system and causes cancer, ageing and various diseases. DNA is repaired by various ways such as Base Excision Repair (BER), Nucleotide Excision Repair (NER), Mismatch Excision Repair (MMR) and Double Strand Break Repair (DSBR). BER mechanism is used when DNA is affected by reactive oxygen species, alkylating agents by oxidation or single strand break. This mechanism involves removal of base and then replacement. NER mechanism is used when DNA is affected by UV light and polycyclic aromatic hydrocarbons by forming bulky adducts and intra-strand crosslink. This mechanism involves removal of DNA fragment and replacement. MMR mechanism is used when mismatch occurs within bases such as A-G pair and T-C pair. This acted by removal of strand followed by digestion and replacement. DSBR is another repair mechanism for damaged DNA by Homologous and Non-Homologous end joining. It includes unwinding, alignment and Ligation. In this Review we will study about DNA repair mechanisms and its consequences.
Keywords: Nitrogen bases; DNA damage; Apoptosis; V(D)J recombination; p53; Xeoderma Pigmentosum; CPD; TATA-BOX; BER; N-Glycosylase; MMR; DAM; NER; XPA; DSBR; NHEJ; DNA-PKcs; XLF; RAD
Abbreviations: MMR: Mismatch Excision Repair; BER: Base Excision Repair; NER: Nucleotide Excision Repair; DSBR: Double Strand Break Repair; DSB: Double Strand Breaks; HR: Homologous Recombination; DAM: Deoxy-Adenine-Methylase; NER: Nucleotide Excision Repair; DSBs: Double Strand Breaks; NHEJ: Non-Homologous End Joining
Introduction
DNA contains mainly three components viz. Pentose Sugar, Nitrogen Base and Phosphate group. DNA determines the growth, development function and reproduction in living beings. It is mainly nucleic acid and proteins, lipids and carbohydrates too. These macromolecules are essential for living beings. It has two strands coiled around each other forming a doublehelix structure. Since, two strands are formed from monomer units called nucleotides, they are termed as polynucleotides. Nucleotide is aggregation of nitrogen bases such as Purines (A,G) and Pyrimidines (C,T,U), Deoxyribose sugar and Phosphate group [1-7].
Number of nucleotide units form the backbone of a DNA helix, by forming phosphodiester bonds. Two nitrogen bases of complementary strand are joined by forming two (A≡T) or three (G=C) hydrogen bonds. Same information is carried by both anti-parallel strands of DNA. Each turn of DNA helix contain ten nucleotide units which are spaced at distance of 3.4Å. Hence, length of one turn is 34Å. Angle between two nucleotides is 36 °C. DNA is wrapped around histone proteins forming a chromatin network (Figure 1).

DNA impairment/damage
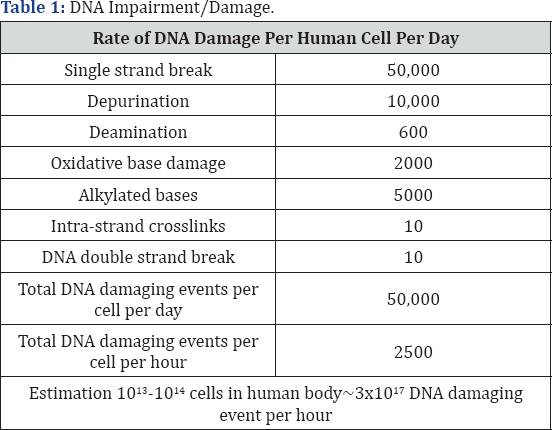
Damage to DNA is nothing but effect on primary structure of double helix. It occurs due to environmental factors and normal metabolic processes inside the cell. It occurs at rate 10,000 to 10,00,000 molecular lesions per cell per day. The modification in the bases occurs and they can disturb the molecule's regular helical structure by forming non native chemical bonds or bulky adducts [8]. DNA damage can occur due to two main agents viz. endogenous cellular processes and exogenous agents. The endogenous cellular processes include oxidation of nitrogen bases and generation of DNA strand interruptions from reactive oxygen species, alkylation of bases, hydrolysis such as deamination, depurination, depyrimidation, bulky adducts formation, mismatch of bases due to error in the replication process of DNA, monoadduct damage due to change in mononitrogen base and also diadduct damage (Table 1).
Whereas damage due to exogenous agents is by UV radiations such as UV-B light which cause direct DNA damage by crosslinking between C and T bases creating pyrimidine dimers and indirect damage by creating radicals. Ionizing radiations damage DNA by radioactive decay or cosmic ray causing strand breaks. Depurination and single strand breaks caused by thermal disruption at elevated temperature can affect the DNA helix structure. Industrial chemicals such as vinyl chloride, hydrogen peroxide and polycyclic aromatic hydrocarbons present in smoke, soot and tar create huge diversity of DNA adducts such as ethenobases, oxidized bases, alkylated bases, phosphotriesters and cross linking of DNA.
Nuclear DNA damage occurs inside nucleus while mitochondrial DNA damage occurs inside mitochondria. Reactive Oxygen species or free radicals create highly oxidative environment and cause mtDNA damage. Senescence is irreversible process by which cell doesn't divide and damaged DNA can't be further replicated.DNA damage is totally different from mutation. It occurs physically and can be repaired but mutation is change in base sequence and can't be repaired.
Consequences of DNA impairment
DNA Damage in mammalian reproductive cell: The damage in mammalian germ cells can be almost completely repaired in short period to provide maintenance to genomic heredity [9,10]. The DNA damage in germ cell is of three main types viz. the damage which is immediately repaired, survival; despite the damage which may cause unexpected death, apoptosis.DNA repair system in reproductive cell achieves repair of genomic mutations formed by genotoxicant factors or serves limited number of mutations. The main repair mechanisms used in germ cells are NER, BER, MMR, DSBR, etc. DNA damage in male germline cell is associated with poor fertilization rates following IVF, defective pre-implantation, embryonic development and high rates of miscarriage and morbidity in the offspring including childhood cancer.
This damage is poorly characterized but is known to involve hypomethylation of key genes, oxidative base damage, endonuclease mediated cleavage and formation of adducts with xenobiotic and the products of lipid peroxidation. There are many possible causes of DNA damage including abortive apoptosis, the oxidative stress associated with male genital tract infection, exposure to redox cycling chemicals and defects of spermiogenesis associated with retention of excess residual cytoplasm. Physical factors such as exposure to radio frequency, electromagnetic radiation or mild scrotal heating can also induce DNA damage in mammalian spermatozoa. Although the underlying mechanisms are unclear, ultimately resolving the precise measure of DNA lesions present in spermatozoa of infertile men will be an important step towards uncovering the etiology of this damage and developing strategies for its clinical management.
Ageing: Loss of homeostasis and increasing probability of illness and death followed by a progressive organic functional decline is termed as Ageing [11,12]. Accumulation of DNA damage is a consequence of ageing and not a leading cause. Factors such as diet, lifestyle and exposure to radiation and genotoxic chemicals seem to have significant influence on the relationship between cumulative DNA damage and age. Ageing and associated diseases are driven by the same pathway that drives developmental growth. Nutrient sensitive signaling network that control growth drives cell hyper-function and thus reproduction. It includes insulin, insulin like growth factor 1 (IGF-1) and in particular target rapamycin (TOR) kinase pathway. Many theories of ageing are based on DNA changes. These theories explain age related changes in DNA as consequence of stochastic events. Main stochastic explanation of DNA damage and other macromolecular damage accumulation with age is oxidative stress theory.
Ageing occurs due to continued activity of pathways and processes during adulthood involving optimization of development to this life stage and thus not due to accumulation of DNA damage. Age associated diseases are linked with increase of DNA damage and mutation or a decrease of DNA repair. Deficient DNA repair cause tissue degeneration and premature ageing is indicated by number of human genetic defects such as CS and XP. XP patients having skin and eye photosensitivity exhibit premature cutaneous ageing, increased incidence of basal cell carcinoma and melanoma. Different DNA repair defect represent only a partial but accelerated contribution to natural ageing process.
Lymphoidal cells and DNA damage: It is related with programmed DNA breaks in lymphoid cells. DNA double strand breaks (DSB) are the most toxic form of DNA damage. Such breaks are also generated in a programmed manner in mammalian lymphoid cells. B-receptor and T-cell receptor loci consist of variable (V) diversity (D) and joining (J) genes that recombine together in a process call V(D)J recombination. This is mechanism that generates a wide repertoire of β cells and T cells which enables these cells to recognize almost unlimited number of different antigens. The process of this recombination is tightly controlled and DSB must be repaired correctly to avoid the persistence of deleterious DNA lesions or resulting translocation. Mature β-cells undergo two genomic alteration: class switch recombination and somatic hyper-mutation [13].
This has severe consequences for affected individuals ranging from immune deficiency and neurological defect to predisposition to the development of malignancies. Human lymphoid deficiencies encompass a wide variety of disorders with broad symptoms. Aetiology of these diseases is not completely known. However the advent of exome sequencing has allowed for the advancement of mutation identification. Genes involved in the non-homologous end joining DNA repair pathway which facilitates repair of programmed DNA breaks, play a major role in β-cell and T-cell disorder. In recent years there have been many reports describing how proteins involved in the NHEJ pathway function and what their contribution is to suppressing human immune deficiencies and also cancer.
Neurodegenerative disorders: Accumulation of DNA lesions in neurons is associated with neurodegenerative disorders including ataxias together with Alzheimer's, Huntington’s and Parkinson's disease. The reason for this may be that neurons generally exhibit high mitochondrial respiration and associated with reactive oxygen species production that can damage mitochondrial DNA. This defect in the DNA trigger neuronal dysfunction and degradation also. Nervous system is particularly vulnerable to DNA damage because the limited capacity of nervous system for cell replacement in adults. Nervous cells are potentially differentiated and don’t repair by DSB but homologous recombination (HR) must be used for error prone NHEJ [14].
Neurons rely heavily on transcription and that oxidative DNA damage can block this. Thus accumulation of DNA lesions in repair defective patients and possibly lead to ageing. Normal individuals might progressively deprive neurons of vital transcripts leading to cell dysfunction or apoptsis. Such processes contribute to the neuro-degeneration observed in ataxias and in the Cookayne's syndrome caused by defects in DNA strand break repair and transcription couples NER respectively.
Genome instability and heritable diseases: DNA repeat instability causes some 40 known diseases that result from expansions or contractions of genetically unstable DNA repeat sequences usually a tri-nucleotide motif within a specific locus. For each disease this instability is thought to arise through repetitive nature of these regions allowing aberrant DNA secondary structure formation during DNA replication or its repair. Hence these neuromuscular and neurodegenerative diseases include fragile X-syndromes, friedrich ataxia, spinocerebellar ataxias, diabetes mellitus Type 2, creutz feldJakob disease, myotonic dystrophy and huntington's disease. Mutations and rearrangement of mtDNA can lead to impaired mitochondrial function such as amyotrophic lateral sclerosis, mitochondrial encephalomyopathy, leigh's syndrome, myoclonic epilepsy, leber's hereditary optic neuropathy and additional mjyopathies [15-18].
Cardiovascular diseases and metabolic syndrome: Proapoptic activity 53 (p53) induced cell death protects against tumerigenesis. p53 activity is harmful in settings such as stroke or heart attack. Induction of p53 by oxidative stress and other sources of DNA damage can also effect the development of atherosclerosis thus providing a link between DNA damage and cardiovascular disease. Growing evidence points to Human atherosclerosis being characterized by enhanced DNA damage and leads to senescence of vascular smooth muscle cells. Metabolic syndrome is common condition characterized by aberrant glucose metabolism, Insulin resistance. The failure of DNA repair generates in cell proliferation, apoptosis and mitochondrial dysfunction. This in turn leads to ketosis, hyperlipidism, increased fat storage and metabolic syndrome [19-22].
Cancer
Cancer and DNA damage has an intimate relationship. A fundamental feature of this is genome instability. For eg. Genomic instability in lymphoid tumors frequently corresponds to chromosomal translocation where proto-oncogenes loci are fused to those of antigens receptor apparently by aberrant antigen- receptor recombination. Chromosomal instability is seen in most sporadic solid tumors. It is likely that transient chromosomal instability arises when telomeres in a nascent tumor become critically short and prone to chromosomal fusion. Whereas activated oncogenes and ensuing DNA replication stress with DSB formation fuel chromosomal instability continuously. At larger stages of cancer progression chronic hypoxia and cycles of hypoxia and re-oxygenation might also contribute to genome instability [23-27].
Most carcinogens operate by DNA damage and cause mutations. The fundamental cause of cancer is damaged or faulty genes- the instructions that tell cells what to do. Genes are encoded with DNA so anything that damages DNA can increase the risk of Cancer. However the number of genes in the same cell need to be damaged before it can become cancerous. Most cancer that are caused by DNA damage that accumulates over a person's lifetime. Some cancers do however have a genetic risk factors. For eg. inherited mutations in some DNA repair genes have been associated with increased risk of Cancer.
Viral, parasitic and other diseases
DNA damage response proteins function in the life cycles of human parasites and pathogen suggesting that the DDR inhibitors could be used to treat their associated pathogenesis. For instance the reliance of HIV on host cell DNA damage response factor suggests a potential for inhibitor in AIDS therapy. The treatments for viral and parasitic diseases need to be evaluated for potential side effects [28,29].
UV induced DNA damage
UV source have higher proportion of radiation bordering between UV-B and UV-A. Such solar UV radiation should produce a higher proportion of Dewar isomers. Since, the photo- isomerisation is most efficient around 320nm, which correspond to the UV absorption maximum of 4-4 PPs. Consequently it has been suggested that all 6-4 PPs should be converted into Dewar isomers upon exposure to sunlight. The CPD and 6-4 PPs make up around 75 and 25% respectively of UV-induced DNA damaged products. Both can distort DNA helix. The ability of UV radiation to damage a given base is determined by flexibility of DNA. The nature of the bases plays a major role since a distribution of the dimeric photoproduct strongly depends on the pyrimidine bases involved. Sequence that facilitate the binding and unwinding are favorable site for damage formation [30,31].
For eg. CPDs formed at higher yields in single stranded DNA and at flexible ends but not in their rigid centres. CPD formation is less frequent when there is binding of the DNA towards the minor groove. One of the transcription factors having the direct formation of 6 PPs is in the TATA BOX where the DNA is bent, but CPDs are formed at the edge of TATA BOX and outside where DNA is not bent. CPDs have been reported to be formed preferentially at the major p53 mutational hotspot in UV-B induced mouse skin tumors. The biological effect of CPDs have been extensively studied in microbes and mammalians. CPDs have been reported to inhibit the programmes of DNA polymerase. The DNA lesions, if unpaired may interfere with DNA transcription and replication and can lead to misreading of genetic code and cause mutation and death.
DNA repair mechanisms
DNA repair can occur by one of two fundamental mechanisms that involve either the reversal of DNA damage or the excision of damaged elements [32]. To use a simplistic analogy if we consider a damage to be represented by a knot (Reversal of damage) whereas in others it is necessary to cut out a piece of string containing knot (Excision of damage) and replace it with a new segment of twine[33-35] (Table 2).
Common Pathway of DNA repair mechanisms
a. Lesion detection: Proteins are bound to DNA lesion
b. Damaged DNA removal: Nucleases, glycosylases etc remove damaged part
c. Repair/Resynthesis: DNA ligase, DNA polymerase
d. Effects on other cellular processes: Replication and/or cell division to allow more time for repair to occur
e. Consequence: Accurate repair-Survival, Inability to Repair-Cell death, Misrepair-Genome Instability
Base excision repair: The name itself implies that it is the predominant mechanism responsible for the repair of damaged DNA bases. In this repair mechanism the DNA helix or the backbone of DNA is not cut out, only the base is cut from the target site. It is opposite to NER. It contains enzymatic reactions pathway. It only removes the damaged bases by cleaving the N-glycoside linkage. There are five main enzymes included in the BER pathway. These are N-glycosylases, AP-endonuclease, flap endonuclease, DNA polymerase and ligase. At least twelve glycosylase have been identified. It acts on the particular base damaged site. The glycosylase enzyme remove wrong base and make AP-site i.e. apurinic or apyrimidinic site. This AP-site is acted by the AP-endonuclease-1(APE-1). APEX-1 and APEX-2 are endonucleases present in human [36-38] (Figure2).
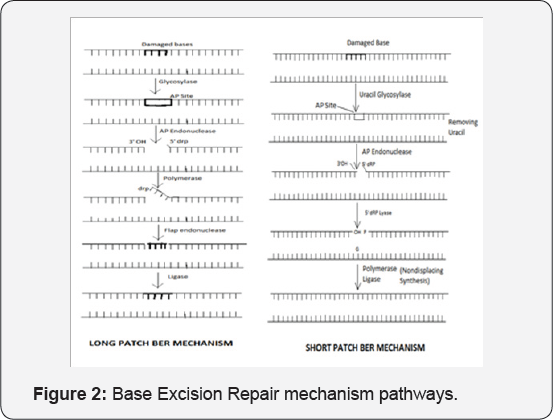
This enzyme hydrolyse the phosphodiester backbone immediately 5' to AP site. Some glycosylases have AP lyase activity. It cleaves AP-sites by β-elimination reaction and produce 3'-phospo-α,β-unsaturated aldehyde and 5'-phosphate at the margins of the break. Enzyme NEIL-1 and NEIL-2 catalyze β and δ elimination by converting the 3'-phospho-α,β-unsaturated aldehyde to 3'-phosphate. After the action of AP-endonuclease on the DNA helix, DNA polymerase acts on this site by 5' to 3' end. Then the enzyme ligase reaction occur. The DNA damage occurs by various other bases. These bases having ability to produce the hydrogen bond to the corresponding nucleotide and form incorrect base pairing. For e.g. A=U. Other examples of base lesions repaired by BER are Oxidized bases such as 8-oxoguanine, 2,6-diamino,4-hydroxy,5-formamido pyrimidine (Fapy G, Fapy A), Alkylated bases such as 3-methyladenine, 7-methylguanosine, Deaminated bases such as hydroxanthine formed from deamination of adenine, Xanthine formed from deamination of Guanine. Uracil initiate the transcription process.
A DNA strand break must have a hydroxyl on its 3' end and a phosphate on its 5' end for ligation to occur. The PNKP (Polynucleotide Kinase Phosphate) is responsible for these ends during BER. The protein having a kinase domain phosphorylates 5' ends and phosphate domain which removes phosphate from 3' end. Many enzymes that repair the single strand interruptions are: Tyrosyl-DNA-phosphodiesterase (TdP-1) repairs several 3' blocking termini, Apartaxin (APTX) end process enzyme which specially repair 5'-adenylate intermediate of DNA ligase activity after that strand break through DNA synthesis and ligation.
This step includes two different types of pathways which are long patch BER and short patch BER. When a single nucleotide is repaired, it is called as Short patch and bulk of nucleotides are repaired, it is called as Long patch repair. In short patch single nucleotide gap filling i.e. AP sites' filling with removal of 5'-drp by DNA polymerase β and successive ligation of the DNA bases by the DNA ligase 1 or complex of DNA ligase 3 and XPCCI occurs. It represents 80-90% of BER. In long patch it is only initiated when a 5' blocking lesions occur. The enzyme PEN-1 removes 5' Flap formed during long patch BER. These endonucleases show a great preference for a long 5' Flap adjacent to 1-nt, 3'-Flap. DNA ligase-1 ligate the break in long patch BER. Other proteins included in BER are poly ADP-ribose polymerase-1 (PARP-1) involved in X-Ray repair. PARP-1 is released by dense negative charge of PARP, responsible for repair of proteins to the damaged DNA sites. Thus the overall BER mechanism is sequential steps of enzymatic reactions.
Mismatch repair: Mismatch are the wrong base pairings occurred during replication. It plays major role in repair of misincorporated bases escaped in proofreading activity of replication polymerase after replication. The mechanism is acted either by insertion or deletion of misincorporated bases with help of MMR proteins. The mechanism is acted stepwise and include three main steps: First one is recognition, second excision step i.e. removal of error containing strand and finally third step being gap repair by filling it with resynthesized DNA. Mutator genes conducting proteins (Mut Proteins) such as Mut S, Mut L and Mut H are involved in running three stages. Parent DNA contains GATC palindrome sequence. Adenine is methylated by enzyme deoxy-adenine-methylase (DAM). Hence the parent strand is methylated. During replication, we get two new strands [39-43] (Figure 3).
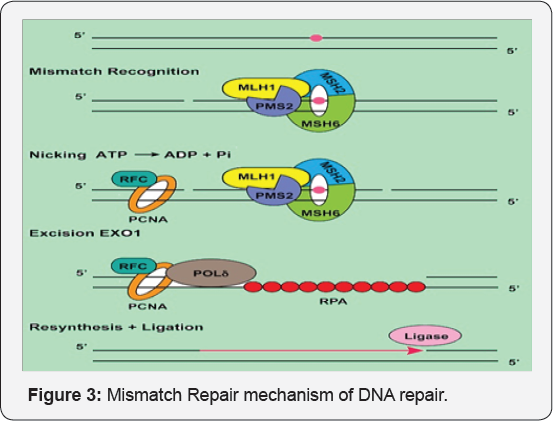
Parent strand is methylated and new strand is not methylated. Hence, alteration in the base is identified and repaired by methyl mismatch repair. Mut S protein recognize the mismatch and bind the respective region. Another protein Mut L couple the recognisation of mispaired base by the Mut S complex to downstream. The function of Mut S is activated by MSH 2, MSH 6, heterodimer and also Mut S α. The damaged site recognized by Mut S α is small IDLS of one or two nucleotides and the MSH 2-MSH 6 recognized longer IDLS. Formation of Mut S DNA complex is followed by ATP dependent requirement of Mut L homolog (MSH) complex.
Mut L will recognize the parent strand. Mut L brings the mispaired area closer and form a loop like structure around the area. Mut H protein provide a vital activity of cleavaging. It is a kind of endonuclease enzyme. Mut H cleave the DNA backbone around the misinocorporated base. Once cleavaging done, the another protein UVr-D functioning as helicase, act and release the cut strand out, forming a gap within. New and corrected nucleotide sequence is added by DNA polymerase 1 and joined by ligase. MMR deficient cells are set to display mutatory phenotype characterized by irreversibly microsatellite instability and elevated mutation frequency. Germline mutations in MMR genes are redisposed to variety of cancers.
Nucleotide excision repair: NER is one of the important repair system which is mainly meant for removal of bulky DNA lesions that can be caused by UV radiation (Cyclobutane- Pyrimidine Dimer) and 6,4- PP (Pyrimidine 6,4-Pyrimidine dimer). Other NER substrate includes bulky chemical adducts, DNA intrastrand links-crosslinks and some oxidative damage resulting form of NER. The mechanism is multistep process. Various proteins are required for action of NER that includes damage recognisation, local opening of double helix of DNA around the lesion, dual incision of damaged DNA strand, gap repair, synthesis and strand ligation. The NER mechanism process occur in the sequential manner. Initially the lesion to DNA such as Thymine dimer formation is recognized. In the NER mechanism of mammalian DNA, more than 30 proteins are required. During NER damaged DNA is recognized by the following binding proteins such as RPA, XPA and XPC-TFIIH, which assemble at the damaged site randomly [44-46].
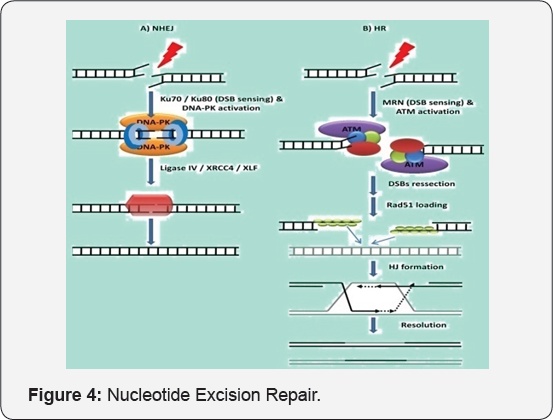
The two proteins uvrB and uvrC self attach and form a complex. For this energy is required and is fulfilled by ATP hydrolysis. The uvrBC complex scan for DNA and specifically bind to the DNA lesion and the uvrC is loaded to actual lesion point. These two processes need energy and thus two ATPs are hydrolysed. It cleaves the DNA backbone around the lesion site about 20bp around uvrD which is a functional helicase, separates two strands of DNA and separate lesion containing DNA. When lesion is removed the gap formed is fulfilled by accurate nucleotide sequence by DNA polymerase 1 and the cut is joined by ligase. Such step require numerous energy and utilize ATP by forming ADP and Pi. Hence, DNA is repaired. This mechanism is highly versatile (Figure 4).
Double strand break repair: The most hazardous type of DNA damage are Double Strand Breaks (DSBs). For ex. Single DSB can cause cell death. Inaccurate repair can lead to deletion or chromosomal aberrations which leads to cancer and other genetic instability syndromes. Thus for cell survival and maintaining genome integrity, DSB repair is critical. DSB repairs by two main mechanisms: NHEJ and HR. These two differ in requirement for homologous template DNA and DNA repair fidelity. HR is error free as it utilize information from undamaged sister chromatid as template. NHEJ is error prone and eliminate DSBs by direct ligation and broken ends in contrastive HR. HR restricted to late S and G2 phases while NHEJ in mammalian cell operating in all phases and cell cycles.
Non-homologous end joining: Small number of essential factors recruited sequentially in molecular mechanism of NHEJ. Recognition and binding of the Ku 70/Ku 80 heterodimer (Ku) to exposed DNA termini of DSB initiates NHEJ process. Preformed ring structure completely encircling DNA duplex is adopted by Ku structurally. DNA-PK holoezyme having protein kinase activity is generated by Ku-DNA complex recruiting catalytic subunit of DNA dependent protein kinase (DNA-PKcs) upon DNA binding. DNA-PKcs contact DNA termini by inward translocation of Ku along DNA induced by DNA PKcs recruitment. Synapsis of DNA- PKcs starts autophosphorylation of DNA-PKcs giving access to DNA termini. Depending on DSB break type and complexity, DNA ends are modified before ligation. Ex.DNA polymerase induced fill in or nucleolytic resection ligates DNA termini containing single strand overhangs. Two X family DNA polymerases carrying resynthesis of missing nucleotide in NHEJ are Pol μ and Pol λ [47-49].
Artemis can excise single strand overhangs and is acquired by DNA-PK phosphorelation. Artemis acts by DNA-PK independent 5' to 3' exonuclease and DNA-PK dependent endonuclease too. Lesion specific BER enzymes: APE-1, Tdp-1, PNKP, two functional exonucleases Exo 1 and WRN-mutated in Werner Syndrome patients can also participate in DNA end 'Cleaning'. Same enzymes in end processing step of NHEJ pathway carry overhanging mispairing and gain/loss of nucleotides associated with NHEJ repair consequently. At end, DNA ends' ligation occur by DNA ligase 4 with its binding partner XRCC4 after appropriate DNA termini processing. It is also promoted by an additional factor XLF (XRCC4-like Factor).
Homologous repair: It is knowledge available is based on studies in bacteria and yeast where it is most efficient. It is divided into three phases- Presynaptic, synaptic and postsynaptic. In presynaptic phase, DSB surrounding ends of DNA are processed through 5’ to 3' end resection generating 3' single stranded tails molecule. Resection involving 5'ends on either side of DSB, trimming back to create short 3' overhangs of single strand DNA is brought about by heteromeric MRN complex (Mre11-Rad50- Nbs1) along with CtIP (RBBP8). BLM helicase (Bloom syndrome, RecQ Helicase-like) and Exo1 exonuclease on combined action brings second step in 5' to 3' resection.RPA binds to single stranded DNA tails removing disruptive secondary nucleus which obstruct Rad51 recombined binding after the resection in conjunction with several mediator proteins-Rad52, BRCA2 and proteins' group called Rad51 paralogs (RAD51B, RAD51C, RAD51D, XRCC2 and XRCC3). Rad51 replaces RPA subsequently. DNA sequence homology search, central reaction of HR, is executed by Rad 51/nucleoprotein filament i.e. Rad51 coated single stranded DNA tail [50].
Invasion reaction is mediated by Rad 51 after identification of homologous DNA: wherein template DNA duplex (sister chromatid) is invaded by damaged DNA strand. DNA polymerase η carry DNA synthesis from 3’ end at invading strand and DNA ligase -1 successively ligate them yielding a four way junction intermediate strand called Holiday Junction. This junction is resolved in three ways: one BLM-Top3-α complex mediated dissolution, second GEN1/Yen1 or Slx1/Slx 4 symmetric cleavage and third structure specific endonuclease Mus81/ Eme1 cleavage asymmetrically giving error free DSB correction.
Conclusion
Various research papers and reviews have been published till today on damage and repair mechanism of DNA. This article provides a brief review on consequences of DNA damage such as effects on nervous system, cardiovascular system, lymphatic system and various disorders such as ageing, cancer, genome instability etc. This article also focuses on the various innate mechanisms of DNA repair with respect to specific DNA damage.
References
- Albert LL, David LN, Michael (2009)M Cox - Lehninger Principles of Biochemistry Pp. 28-31.
- Travers A, Muskhelishvili G(2015) DNA structure and function. FEBS J 282(12): 2279-2295.
- Sinden RR, Pearson CE, Potaman VN, Ussery DW (1998) DNA: Structure and function. Advances in Genome Biology 5(C): 1-141.
- Wolffe AP (1995) DNA structure and function. The FEBS Journal 20(8): 330-331.
- Points K, Korf BR, Irons MB (2012) DNA Structure and Function. Human Genetics and Genomics.
- Herbert A, Rich A (1999) Left-Handed Z-DNA: Structure and Function. Genetica 1999 106(1-2): 1-19.
- Rich A, Zhang S (2003) Z-DNA: the long road to biological function. Nat Rev Genet 4(7): 566-572.
- Hakem R (2008) DNA-damage repair; the good, the bad, and the ugly EMBO J 27(4): 589-605.
- Ozturk S, Demir N (2011) DNA repair mechanisms in mammalian germ cells. Histol Histopathol 26(4): 505-517.
- Jackson SP, Bartek J (2009) The DNA-damage response in human biology and disease. Nature 461(7267): 1071-1078.
- Soares JP, Cortinhas A, Bento T, Leitao JC, Collins AR, et al. (2014) Aging and DNA damage in humans: a meta-analysis study. Aging (Albany NY) 6(6): 432-439.
- Menck CF, Munford V (2014) DNA repair diseases: What do they tell us about cancer and aging? Genet Mol Biol 37(1 Suppl): 220-233.
- Prochazkova J, Loizou JI (2016) Programmed DNA breaks in lymphoid cells: repair mechanisms and consequences in human disease. Immunology 147(1): 11-20.
- Jeppesen DK, Bohr VA, Stevnsner T (2011) DNA repair deficiency in neurodegeneration. Prog Neurobiol 94(2): 166-200.
- Alt FW, Zhang Y, Meng FL, Guo C, Schwer B (2013) Mechanisms of programmed DNA lesions and genomic instability in the immune system. Cell 152(3): 417-29.
- Ghosal G, Chen J (2013) DNA damage tolerance: a double-edged sword guarding the genome. Transl Cancer Res 2(3): 107-129.
- Tubbs, Nussenzweig A (2017) Endogenous DNA Damage as a Source of Genomic Instability in Cancer. Cell 168(4): 644-656.
- Giuseppe PS, Giovana B, Ramiro BN, Thiago GH, Claudia RR (2015) Metabolic Syndrome and DNA Damage: The Interplay of Environmental and Lifestyle Factors in the Development of Metabolic Dysfunction. OJEMD 5(7): 65-76.
- Shimizu I, Yoshida Y, Suda M, Minamino T (2014) DNA Damage Response and Metabolic Disease. Cell Metab 20(6): 967-977.
- Mercer JR, Cheng KK, Figg N, Gorenne I, Mahmoudi M, et al. (2010) DNA Damage Links Mitochondrial Dysfunction to Atherosclerosis and the Metabolic Syndrome. Circ Res 107(8): 1021-1031.
- Prasad M, Bronson SC, Warrier T, Badarinath A, Rai S, et al. (2015) Evaluation of DNA damage in Type 2 diabetes mellitus patients with and without peripheral neuropathy: A study in South Indian population. J Nat Sci Biol Med 6(1): 80-84.
- Lakin ND, Jackson SP (1999) Regulation of p53 in response to DNA damage. Oncogene 18(53): 7644-7655.
- Lee SC, Chan JC (2015) Evidence for DNA damage as a biological link between diabetes and cancer. Chin Med J 128(11): 1543-1548.
- Torgovnick A, Schumacher B (2015) DNA repair mechanisms in cancer development and therapy. Front Genet 6: 157.
- Gasser S, Raulet D (2006) The DNA damage response, immunity and cancer. Seminars in Cancer Biology 16(5): 344-347.
- Stoyan C, Rumena P, George ChR, Nikolai Z (2014) DNA repair and carcinogenesis. Biodiscovery 12: 1.
- Brégnard C, Benkirane M, Laguette N (2014) DNA damage repair machinery and HIV escape from innate immune sensing. Front Microbiol 5: 176.
- Ellis LR, Robert H, Roger J (2016) Grand -Activation of the DNA Damage Response by RNA Viruses. Biomolecules 6(2).
- Sinha RP, Häder DP (2002) UV-induced DNA damage and repair: A review. Photochem Photobiol Sci 1(4): 225-236.
- Ravanat JL, Douki T, Cadet J (2001) Direct and indirect effects of UV radiation on DNA and its components. J Photochem Photobiol B 63 (13): 88-102.
- Jacobs AL, Schär P (2012) DNA glycosylases: in DNA repair and beyond. Chromosoma 121(1): 1-20.
- DNA damage and DNA repair.
- U Satyanarayana, U Chakrapani (2013) Biochemistry (4th Edn)Elsevier Publication Pp. 534-539.
- Thomas S. Dexheimer. DNA Repair Pathways and Mechanisms.
- Kim YJ, Wilson DM (2012) Overview of Base Excision Repair Biochemistry. Curr Mol Pharmacol 5(1): 3-13.
- Seeberg E, Eide L, Bj 0ras M (1995) The base excision repair pathway Trends Biochem Sci 20(10): 391-397.
- Zharkov DO (2008) Base excision DNA repair. Cell Mol Life Sci 65(10): 1544-1565.
- Li GM (2008) Mechanisms and functions of DNA mismatch repair. Cell Res 18(1): 85-98.
- Buermeyer AB, Deschenes SM, Baker SM, Liskay RM (1999) Mammalian DNA mismatch repair. Annu Rev Genet 33: 533-564.
- Jun SH, Kim TG, Ban C (2006) DNA mismatch repair system: Classical and fresh roles. FEBS J 273(8): 1609-1619.
- Kolodner RD, Marsischky GT (1999) Eukaryotic DNA mismatch repair. Curr Opin Genet Dev 9(1): 89-96.
- Kunkel TA, Erie DA (2005) DNA mismatch repair. Annu Rev Biochem 74: 681-710.
- Marteijn JA, Lans H, Vermeulen W, Hoeijmakers JH (2014) Understanding nucleotide excision repair and its roles in cancer and ageing. Nat Rev Mol Cell Biol 15(7): 465-481.
- Sancar A, Tang MS (1993) Nucleotide Excision Repair. Photochem Photobiol 57(5): 905-921.
- Reardon JT, Sancar A (2005) Nucleotide Excision Repair. Progress in Nucleic Acid Research and Molecular Biology 79: 183-235.
- Sugasawa K (2010) Regulation of damage recognition in mammalian global genomic nucleotide excision repair. Mutat Res 685(1-2): 29-37.
- Sonoda E, Hochegger H, Saberi A, Taniguchi Y, Takeda S (2006) Differential usage of non-homologous end-joining and homologous recombination in double strand break repair. DNA Repair (Amst) 5(9- 10): 1021-109.
- Davis AJ, Chen DJ (2013) DNA double strand break repair via non- homologous end-joining. Transl Cancer Res 2(3): 130-143.
- Lieber MR (2010) The Mechanism of Double-Strand DNA Break Repair by the Nonhomologous DNA End Joining Pathway. Annu Rev Biochem 79: 181-211.
- Jasin M, Rothstein R (2013) Repair of Strand Breaks by Homologous Recombination. Cold Spring Harb Perspect Biol 5(11): a012740.






























