Polyol Pathway Shed Light on Non-Obstructive Azoospermia Testicular Sperm Extraction (TESE) Negative ROS Imbalance
Kambiz Gilany*1,2
1Reproductive Biotechnology Research Center, Avicenna Research Institute, Iran
2Metabolomics and Genomics Research Center, Endocrinology and Metabolomics Molecular Cellular Sciences Institute, Tehran University of Medical Sciences, Iran
Submission: July 05, 2018;Published: August 24, 2018
*Corresponding author:Kambiz Gilany, Reproductive Biotechnology Research Center, Avicenna Research Institute, ACECR, Tehran, Metabolomics and Genomics Research Center, Endocrinology and Metabolomics Molecular Cellular Sciences Institute, Tehran University of Medical Sciences, Tehran, Iran.
How to cite this article:Kambiz G. Polyol Pathway Shed Light on Non-Obstructive Azoospermia Testicular Sperm Extraction (TESE) Negative ROS Imbalance. Glob J Reprod Med. 2018; 5(5): 555675. DOI: 10.19080/GJORM.2018.05.555675.
Abstract
In our recent study regarding non-obstructive azoospermia by metabolomics technology, we clearly demonstrated that testicular sperm extraction negative patients are in extreme oxidative imbalance. With a reanalysis of our data and the integration of proteomics data, we found the Polyol pathway is up-regulated in non-obstructive azoospermia testicular sperm extraction negative patients. In this regard, we suggest the contribution of this biochemical pathway in the extreme oxidative imbalance of non-obstructive azoospermia testicular sperm extraction negative patients.
Keywords: Azoospermia; Metabolomics technology; Testicular sperm extraction; Sperm production; Non-obstructive azoospermia; Fingerprinting; Non-invasive method; Proteomics; Polyol pathway, Diabetic patient’s; Plasma proteome database; Male reproductive system; Extreme oxidative imbalance; Human protein; Glutathione deficiency; Glutathione redox state; Proteomics analysis
Abbrevations: NOA: Non-Obstructive Azoospermia; TESE: Testicular Sperm Extraction; UPP: Up-Regulated Polyol Pathway
Introduction
Non-obstructive azoospermia (NOA) refers to absence of sperm in the semen because of the abnormal sperm production. It is estimated that 6-10% of infertile men are NOA [1]. In our recent study, we used the untargeted metabolomics and metabolic fingerprinting approach for of seminal plasma in order to develop a non-invasive method for detection of spermatogenesis [2,3]. Following a reanalysis of untargeted metabolomics data, we found that sorbose was up-regulated in NOA the testicular sperm extraction (TESE) negative patients (NOA TESE(-)). This data was in line with the proteomics analysis of seminal plasma from NOA patients conducted by Batruch et al. [4]. With the Integrating of the proteomics and metabolomics data, we found that polyol pathway is up-regulated in NOA TESE(-) patients. It is an old fact that glucose and fructose concentrations are increased in azoospermia patients [5].
To the best of our knowledge, no studies have been done regarding the measurement of redox potential in azoospermia or in NOA. However, it is known from diabetic patient’s data redox potential (NAD/NADH and NADPH/NADP) in up-regulated polyol pathway (UPP) is in imbalance [6]. It is known that NADPH acts to promote glutathione redox state while its deficiency causes glutathione deficiency [7]. Bhardwaj et al. [8] have shown that azoospermia patients, in fact have glutathione deficiency. Additionally, we have already shown that NOA TESE (-) are in extreme oxidative imbalance by metabolic fingerprinting. It is postulated that NADH oxidase (NOX1) is responsible for the increased level of ROS in UPP [9]. With a search in seminal plasma proteome database, we found NOX1 activator protein (NOXA1) [10]. Furthermore, with a search in the Human Protein Atlas database we also found that NOX1 is expressed in male reproductive system [11]. By integrating these data, we suggest a hypothesis how UPP leads to extreme oxidative imbalance in NOA TESE(-) patients (Figure 1).
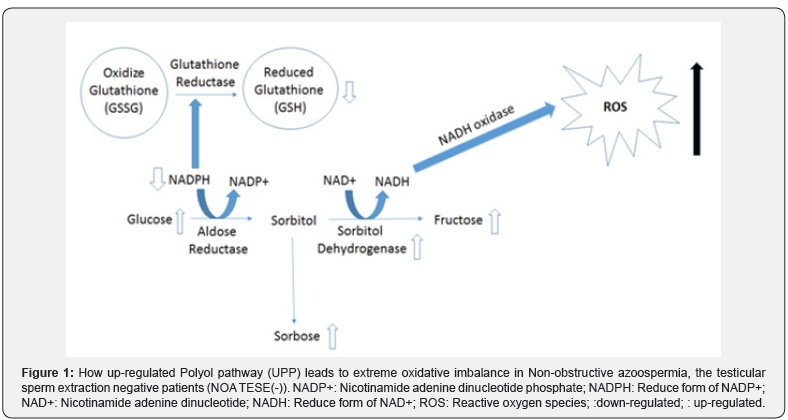
References
- Costabile RA, Spevak M (2001) Characterization of patients presenting with male factor infertility in an equal access, no cost medical system. Urology 58(6):1021-1024.
- Gilany K, Mani Varnosfaderani A, Minai Tehrani A, Mirzajani F, Ghassempour A, et al. (2017) Untargeted metabolomic profiling of seminal plasma in nonobstructive azoospermia men: A noninvasive detection of spermatogenesis. Biomed Chromatogr 31(8).
- Gilany K, Jafarzadeh N, Mani Varnosfaderani A, Minai Tehrani A, Sadeghi MR, et al. (2018) Metabolic Fingerprinting of Seminal Plasma from Non-obstructive Azoospermia Patients: Positive Versus Negative Sperm Retrieval. J Reprod Infertil 19(2):109-114
- Batruch I, Smith CR, Mullen BJ, Grober E, Lo KC, Diamandis EP, et al. (2012) Analysis of seminal plasma from patients with non-obstructive azoospermia and identification of candidate biomarkers of male infertility. J Proteome Res11(3): 1503-1511.
- Buckett W, Lewis Jones D (2002) Fructose concentrations in seminal plasma from men with nonobstructive azoospermia. Arch Androl 48(1): 23-27.
- Obrosova IG, Stevens MJ (1999) Effect of dietary taurine supplementation on GSH and NAD (P)-redox status, lipid peroxidation, and energy metabolism in diabetic precataractous lens. Invest ophthalmol Vis Sci 40(3):680-8.
- Vogel R, Wiesinger H, Hamprecht B, Dringen R (1999) The regeneration of reduced glutathione in rat forebrain mitochondria identifies metabolic pathways providing the NADPH required. Neuroscience letters. 275(2): 97-100.
- Bhardwaj A, Verma A, Majumdar S, Khanduja K (2002) Status of vitamin E and reduced glutathione in semen of oligozoospermic and azoospermic patients. Asian J Androl 2(3): 225-228.
- Tang W, Martin KA, Hwa J (2012) Aldose reductase, oxidative stress, and diabetic mellitus. Front Pharmacol 3: 87.
- Gilany K, Minai Tehrani A, Savadi Shiraz E, Rezadoost H, Lakpour N (2015) Exploring the human seminal plasma proteome: an unexplored gold mine of biomarker for male infertility and male reproduction disorder. J Reprod Infertil 16(2): 61-71.
- Uhlen M, Oksvold P, Fagerberg L, Lundberg E, Jonasson K, et al. (201) Towards a knowledge-based human protein atlas. Nat Biotechnol 28(12):1248-1250.






























