Relationship between the Quantitative Results of HBV DNA and HBV Markers (HBV M) in Population with low-level HBsAg
Xiaoxiao Jiang1*, Yuzhu Dai2*, Changgui Sun2*, Dawei Cui3, Xujian Xu4 and Jun Cheng2
1Department of Gastroenterology, 117th Hospital of PLA, China
2Clinical Experimental Center, 117th Hospital of PLA, China
3Medical College of Zhejiang University, China
4Department of Biotechnology, University of Tokyo, Japan
Submission: February 06, 2018; Published: February 13, 2018
*Corresponding author: Jun Cheng, Clinical Experimental Center, the 117th Hospital of PLA, China, Tel: +86-571-87348812; Fax: +86-571-87348812; Email: cj1171967@163.com
How to cite this article: Xiaoxiao J, Yuzhu D, Changgui S, Dawei C, Xujian X, Jun C. Relationship between the Quantitative Results of HBV DNA and HBV Markers (HBV M) in Population with low-level HBsAg. Glob J Nano. 2018; 3(5): 555623. DOI: 10.19080/GJN.2018.03.555623
Abstract
Background and aim: Low-level HBsAg has brought new challenges to clinical laboratory and the treatment of HBV infectious diseases. The study was to analyze the quantitative differences between hepatitis B virus markers (HBV M) and HBV DNA in population with high-level or low-level HBsAg and to reveal the characteristics of HBV DNA and HBV M in population with low-level HBsAg.
Methods: A total of 264 chronic HBV-infected patients were enrolled in our study, and were divided into low-level HBsAg group (147 cases) and high-level HBsAg group(117 cases) based on the HBsAg level or were divided into immune tolerance stage, immune clearance stage and nonactive stage based on the natural history. Real-time PCR and microparticle enzyme immunoassay (MEIA) were used to determine the content of HBV DNA and HBV M in serum samples of high-level and low-level HBsAg patients, and then the quantitative results were compared.
Results: There were statistically significant differences between HBV DNA and HBV M( anti-HBs, HBeAg and anti-HBe) of low-level HBsAg patients in immune tolerance stage(7 cases) and immune clearance stage(4 cases)(t=2.531~9.181, P<0.01~0.05 ). HBV DNA lower than 105copies/L was found in 94.1%(128/136) of serum samples in non-active stage, and the detection rates of direct PCR and enriching PCR were 10.3%(14/136) and 10.3%(47/136) in low-level HBsAg group, respectively, and there were no correlations between HBV DNA and each of HBV M(P>0.05). In high-level HBsAg group, HBV DNA was positively correlated with HBeAg and anti-HBe in 25 cases of immune tolerance stage(r=0.744~0.772, tr=3.858~4.207, P<0.01), and was only negatively correlated with anti-HBs in 46 cases of immune clearance stage and 46 cases of non-active stage(r=-0.693—0.598, tr=- 4.616—3.936, P<0.01~0.05).
Conclusion: Population with low-level HBsAg has specific serological characteristics, which may be correlated with organic immune tolerance and individualized immune responses.
keywords: Hepatitis b virus marker; Chronic HBV Infection; HBsAg
Introduction
It has been demonstrated that the proportion of population with low-level HBsAg accounted for 6.68%-23.16% among HBsAg positive population [1-4]. The detection and reporting of low- level HBsAg have brought new challenges to clinical laboratory and new thinking to the diagnosis and treatment of infectious diseases, so it has attracted the concern of domestic and foreign experts [5-11]. Currently, some quantitative assessments about HBV DNA and HBV M in population with high-level HBsAg have been reported [12], so, the relationship between the quantitative results of HBV DNA and HBV M in population with low-level HBsAg is worth to be discussed and studied. Therefore, we divided the patients into different groups based on the HBsAg level and the natural history [13,14] of chronic HBV- infected patients, then, real-time PCR and microparticle enzyme immunoassay (MEIA) were used to quantitatively determine the content of HBV DNA and HBV M in serum samples of low-level HBsAg patients, in order to understand the relationship between HBV DNA and HBV M in population with low-level HBsAg.
Materials and Methods
Serum samples
A total of 147 serum samples with low-level HBsAg were collected from outpatients, inpatients and medical examination population with chronic HBV-infection as low-level HBsAg group(According to the standard of Clinical Laboratory Center, Ministry of Public Health, the content of HBsAg<5μg/L was considered as low-level. Low-level HBsAg was less than 72±6.8S/N detected by Abbott Axsym immune analyzer after ELISA screening, and 4 cases of positive HBsAg/HBeAg/anti- HBc, 137 cases of positive HBsAg/anti-HBe/anti-HBc and 6 cases of positive HBsAg/anti-HBc were confirmed by neutralization test). Low-level HBsAg samples emerged in early stage of HBV infection and HBsAg/anti-HBs conversion stage of recovery stage, and long-term concurrent HBsAg/anti-HBs positive samples were eliminated. Samples were divided into 4 cases of immune tolerance stage, 7 cases of active stage, and 136 cases of non-active or low/non-replicative stage (non-active stage) based on the natural history [6].
There were no liver-protection therapy, enzyme reduction therapy and application history of immunomodulators and antiviral drugs before sample collection. In addition, 117 serum samples with high-level HBsAg of chronic HBV infection were collected as high-level HBsAg group (the content of HBsAg>79S/N), including 25 cases of immune tolerance stage, 46 cases of active stage and 46 cases of non-active stage; 31 cases of positive HBsAg/HBeAg/anti-HBc and 86 cases of positive HBsAg/anti-HBe/anti-HBc. Serum samples were sub-packaged and stored at -70-
Reagents and instrumentso ABI 7000 fluorescent quantitative PCR machine (ABI, USA); nucleic acid concentrated solution, reagents for HBV DNA fluorescent quantitative PCR (Shanghai Shenyou Bioengineering Co., Ltd.); AXSYM autoimmunal analyzer and reagents for HBsAg, anti-HBs, HBeAg, anti-HBe, and anti-HBc detection (Abbott, USA).
Methods
a. Real-time fluorescent quantitative PCR was adopted to determine the content of HBV DNA in serum samples of these two groups. Enriching method was used in low-level HBsAg group without detected HBV DNA and high-level HBsAg group: 500ul serum was mixed with 500ul nucleic acid concentrated solution (The procedures of critical control, negative control and positive control were same to serum samples and the reference materials were not necessary to deal with). The mixture was mixed well by shaking, and then stood at 400c for 5min, then centrifuged at 15000rpm for 5min. The supernatant was discarded, and 50ul lysis buffer was added into the pellet (Direct method: 500ul serum was mixed with 500ul lysis buffer, and the following steps were same to enriching method). The mixture was vibrated severely or broken by tip until no sediment. Then the sample was centrifuged briefly, and incubated in boiling water for 10min, then centrifuged at 15000rpm for 5min. The supernatant was stored at -700c for detection within one month.
b. MELA was used to determine HBV M in serum samples of these two groups. The detail experiments were performed according to the manufacturer's protocol of reagents.
c. The quantitative results of HBV DNA and HBV M in serum samples of these two groups were collected and divided according to the copy number of HBV DNA: HBV DNA less than 105copies/L was included in Group A, and HBV DNA less than 106copies/L was included in Group B. In the same way, HBV DNA more than 1011copies/L was included in Group G.
Statistical Analyses
HBV DNA was expressed with logarithmic mean of copy number. The logarithm of samples without detected HBV DNA was calculated as 0. HBsAg(S/N), anti-HBs (IU/L), HBeAg (S/N), anti-HBe (S/CO) and anti-HBc (S/CO) were expressed by mean ± standard deviation. Different means were compared by T test, and χ2 test was used to compare the detection rates of HBV DNA between two groups of non-active stage. Correlation analysis between HBV DNA and HBV M was carried out. Statistical analyses of all data were performed with SPSS12.01 software package.
Results
From the quantitative results of HBV DNA and HBV M in serum samples of 4 cases in immune tolerance stage and 7 cases in immune clearance stage in low-level HBsAg group, we observed that HBV DNA in immune tolerance stage was found in Group D and E, and HBV DNA in immune clearance stage was found in Group B-D. According to the comparison of means of parameters, we found that there were statistically significant differences among logarithmic means of HBV DNA (t=3.708, P=0.005), anti-HBs (t=2.531, P=0.032), HBeAg (t=9.181, P=0.002), and anti-HBe (t=7.943, P=0.004) between two groups. The results were shown in Table 1.
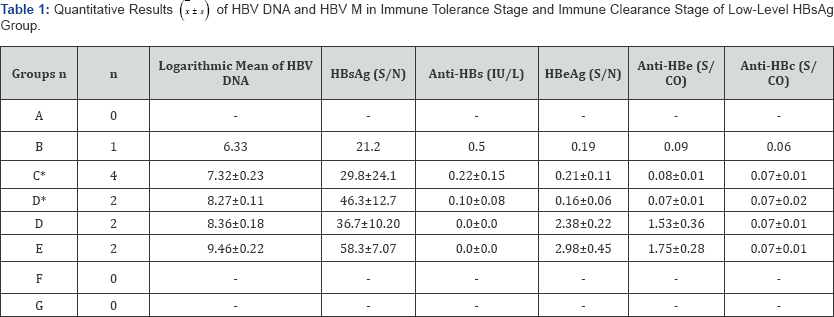
*Determination results of immune clearance stage.
From the quantitative results of HBV DNA and HBV M in 136 serum samples in non-active stage in low-level HBsAg group, we observed that HBV DNA lower than 105copies/L was found in 94.1% (128/136) of samples. Only a detection rate of 10.3%14/136) for HBV DNA was obtained by direct PCR and the detection rate for HBV DNA in non-detection samples could be increased to 34.6% (47/136) by enriching PCR. Correlation analysis showed no correlations between HBV DNA and each parameter of HBV M in low-level HBsAg group (P>0.05). The results were shown in Table 2.
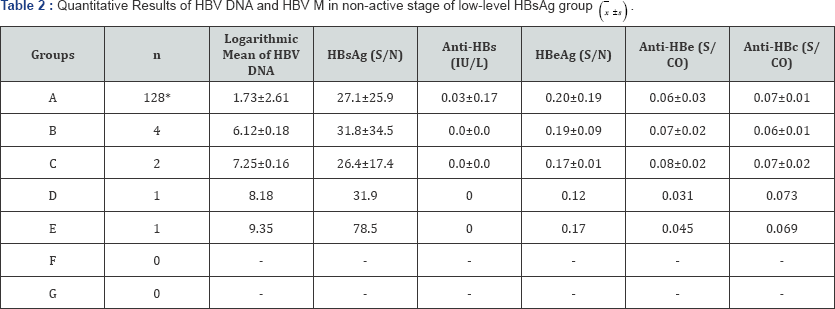
*HBV DNA was detected in 6 of 128 cases by direct method and in 33 cases by enriching method.
From the quantitative results of HBV DNA and HBV M of each stage in high-level HBsAg group, we observed that HBV DNA was positively correlated with HBeAg (r=0.744, tr=3.858, P=0.002) and anti-HBe (r=0.772, tr=4.207, P=0.001) in immune tolerance stage, and was only negatively correlated with anti-HBs (r=- 0.693/-0.598, tr=-4.616/-3.936,P=0.000/0.017) in immune clearance stage and non-active stage. The results were shown in Tables 3-5.

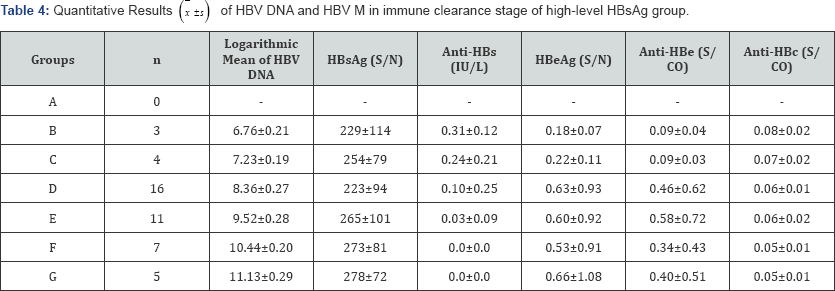
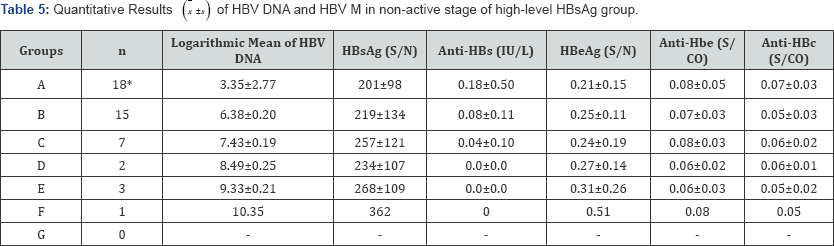
*HBV DNA was detected in 2 of 18 cases by direct method and in 9 cases by enriching method.
Discussion
HBV DNA and HBV M were quantitatively determined in a total of 264 serum samples of immune tolerance stage, immune active stage and non-active stage in two groups with different concentrations of HBsAg. The results indicated that the HBV DNA of immune tolerance stage and clearance stage in low-level HBsAg group were distributed at 108-9copies/L and 106- 8copies/L, respectively. However, they were distributed at 107- 11copies/L and 106-11copies/L in high-level HBsAg group. There were significant differences between these two groups in both the number of observation cases and distribution ranges of HBV DNA. HBV DNA and HBV M in 128 serum samples of non-active stage in low-level HBsAg group were quantitatively determined. The detection rates were 10.3% and 34.6% by direct PCR and enriching PCR, respectively. There were statistically significant differences as compared with the high-level HBsAg group (65.2% by direct method, 84.8% by enriching method) (P<0.01).
Correlation analysis showed no correlations between HBV DNA and each parameter of HBV M in low-level HBsAg group (P>0.05, Correlation analysis was not performed in immune tolerance stage and immune clearance stage due to the few groups). However, HBV DNA of immune tolerance stage in high-level HBsAg group was correlated with HBeAg and anti-HBe (r=0.744-0.772, tr=3.858-4.207, P<0.01), which was not absolutely same as the pervious literatures [12]. The main reason was the difference caused by grouping. The grouping in some literatures was according to model of HBsAg/HBeAg/ anti-HBc for acute hepatitis B. HBV DNA in immune clearance stage and non-active stage was only negatively correlated with anti-HBs (r=-0.693~-0.598,tr=-4.616~-3.936, P<0.001~0.05), which was probably due to the multiple transformation between immune clearance stage and non-active stage. However, HBV DNA in non-active stage in low-level HBsAg was not correlated with anti-HBs (P>0.05). According to the literatures [15], it was considered to be relevant to the immune tolerance caused by its low dose and the less transformation to the active stage due to the long-term tolerance status.
The results in Table 1 also indicated that there were statistically significant differences between HBV DNA of immune tolerance stage and immune clearance stage in low-level HBsAg group and the mean value of HBsAg quantitative determination (P<0.05). However, there were no differences among either two stages in high-level HBsAg group. Because of the small sample size, it still needs to be further confirmed whether there exists such kind of differences. From the clinical diagnosis and typing, patients with low-level HBsAg (The low-level HBsAg emerged in the early stage of HBV infection or HBsAg/anti-HBs transformation stage of recovery stage were eliminated, and the very few cases with low-level HBsAg [16-19] existing as a longterm HBsAg/anti-HBs positive model caused by S genovariation were also eliminated) should belong to patients with chronic HBV infection. 90% of them belong to asymptomatic carriers (ASC), and only about 5% belong to chronic active hepatitis B (CHB). Chronic HBV infection is the most complicated problem in clinical medicine and epidemiology. Grouping and studying population with chronic HBV infection based on the HBsAg level and natural history of chronic HBV-infected persons could eliminate other factors besides different HBsAg levels and reveal some valuable information.
In our study, while dividing 147 cases with low-level HBsAg and 117 cases with low-level HBsAg based on the natural history, we found that HBV DNA was lower than the staging standard of "Guidelines for Prevention and Cure of Chronic Hepatitis B" (hereinafter referred as "the Guidelines") in 2 cases of immune tolerance stage and 7 cases of immune clearance stage in high- level HBsAg group, however, HBV DNA was higher than the staging standard of "the Guide" in 28 cases of non-active stage (The performance model of HBV M, ALT and liver histology were in accordance with the standard) [13,14]. Similarly, HBV DNA was lower than the staging standard of "the Guidelines" in 5 cases of immune clearance stage in low-level HBsAg group, and HBV DNA was higher than the staging standard of "the Guidelines" in 8 cases of non-active stage in low-level HBsAg group. These cases could not be classified as other stages.
Therefore, we considered that while dividing the chronic HBV infected persons based on the natural history, it should take issue with the determination of HBV DNA level. Otherwise, a small percentage of cases were difficult to divide according to the natural history. In addition, "HBV DNA of non-active stage was less than the lower limit" in "the Guidelines" is also worth to be discussed. With the enhancement of PCR sensitivity, HBV DNA can be detected by routine PCR in quite a few of serum samples of non-active stage actually (In our study, the detection rate were 10.3% and 34.6% in low-level HBsAg by direct method and enriching method, respectively, and 65.2% and 84.4% in high- level HBsAg respectively.), so, we suggested to revise the part of "natural history" in "the Guidelines"
In summary, chronic HBV infected population with low- level HBsAg is a special group formed during the HBV infection, dissemination and natural clearance with the increasing ages. Most of them are in non-active stage with the phenomenon of immune tolerance (The capability of immunocomplex formation and clearance is decreased with complete or incomplete T, B lymphocyte tolerance [15]). The non-active stage switches alternately and repeatedly with immune clearance stage in high-level HBsAg group. There exists immune response but not immune tolerance, and it has essential difference with immune tolerance stage in high-level HBsAg group (difference in age and high-dose tolerance [15]). Therefore, systemic studies for population with low-level HBsAg including the distribution in natural population, clinical features, performance model of serologic HBV marker, serotype, genotype, humoral and cellular immune function, S gene sequence, and even the molecular biological mechanism and epidemiologic feature of whole genome and so on, contributes to clarify the mechanism of immune tolerance of population with low-level HBsAg, and there may be important clinical and epidemiological significances in clearing chronic HBV infection and preventing the dissemination of HBV through breaking immune tolerance.
Acknowledgment
This work was supported by the Medical Science Foundation of Nanjing Military Command (NO.12MA117), and the Natural Science Foundation of Zhejiang Province (N0.Y15H200001). We thank Mr. Qiang-Lin Duan for his editorial assistance and revision of the paper.
References
- Cheng J, Sun GZ, Chen Y (2001) Pattern analysis of seven Hepatitis B markers in high and low concentrations of HBsAg serum. Chin J Cellular and Molecular Immunology 17(5): 443-444.
- Chen Y, Zhong BY, Xu GY (2001) Determination of low level HBsAg in serum by MEIA. Chin J Lab Med 24(1): 39-41.
- Cheng J, Sun cg, Chen Y, Dai YZ, Xu ZL, et al. (2009) Molecular analysis on chronic hepatitis B patients with low- level HBsAg. Chin J Lab Med 32(10): 1128-1132.
- Cheng J,Sun CG,Chen Y,Xu ZL,Wang GZ,et al. (2010) Analysis on clinical features and immunity in chronic hepatitis B virus infected patients with low-level HBsAg. Afr J of Microbiol Res 4(7): 547-550.
- LI JM (2006) Attention to the confirmative tests of weak reactive samples should be paid in serological testing for infectious diseases. Chin J Lab Med 29(7): 577-580.
- Ozdil B, Cosar A, Akkiz H (2009) Negative correlation between viral load and HBsAg levels in chronic HBV-infected patients. Arch Virol 154(09):1451-1455.
- De Gascun CF, Fraher M, Crean M, Connell J, Hall WW (2010) The importance of being earnest: following up a low level hepatitis B surface antigen (HBsAg) result. J Clin Virol 49(2): 79-81.
- Tseng TC, Liu CJ, Yang HC, Chan CL, Hsu CA, et al. (2013) Serum Hepatitis B Surface Antigen Levels Help Predict Disease Progression in Patients With Low Hepatitis B Virus Loads. Hepatology 57(2): 441450.
- Martinot-Peignoux M, Lapalus M, Asselah T, Marcellin P (2014) HBsAg quantification: useful for monitoring natural history and treatment outcome. Liver Int 34 (Suppl 1): 97-107.
- Huang G, Lau WY, Zhou WP, Shen F, Pan ZY, et al. (2014) Prediction of Hepatocellular Carcinoma Recurrence in Patients With Low Hepatitis B Virus DNA Levels and High Preoperative Hepatitis B Surface Antigen Levels. JAMA Surg 149(6): 519-527.
- Yang XH, Shi XF (2016) Significance of HBsAg quantification in guiding clinical treatment of chronic hepatitis B. Zhonghua Gan Zang Bing Za Zhi 24(4): 317-320.
- Cheng J, Sun CG, Wang GZ, Tuncel U, Sunbul M, et al. (2007) Relationship of the quantitative results between two serologic patterns of HBV DNA and HBV M. J Cellular and Molecular Immunology 23(7): 681-683.
- Chinese Society of Hepatology,Chinese Medical Association; Chinese Society of Infectious Diseases,Chinese Medical Association (2015) The Guideline of Prevention and Treatment for Chronic Hepatitis B. Beijing,CMA.
- World Health Organization (2015) Guidelines for the prevention,care and treatment of persons with chronic hepatitis B infection. Geneva, WHO, France, pp. 1-166.
- Cheng J, Sun CG, Chen Y (2008) Analysis on clinical features and immune function in population with low concentration of HBsAg. J Cellular and Molecular Immunology 24(12): 1187-1189.
- Kohno H, Inoue T, Tsuda F, Okamoto H, Akahane Y (1996) Mutations in the envelope gene of hepatitis B virus variants co-occurring with antibody to surface antigene in sera from patients with chronic hepatitis B. J Gen Virol 77(Pt8): 1825-1831.
- Pollicino T, Amaddeo G, Restuccia A, Raffa G, Alibrandi A, et al. (2012) Impact of hepatitis B virus (HBV) preS/S genomic variability on HBV surface antigen and HBV DNA serum levels. Hepatology 56(2): 434443.
- Chen Y, Qian F, Yuan Q, Xuefen Li, Wei Wu, et al. (2011) Mutations in hepatitis B virus DNA from patients with coexisting HBsAg and anti- HBs. J Clin Virol 52(3): 198-203.
- Liu W, Hu T, Wang X, Huang M, Yuan C, et al. (2012) Coexistence of hepatitis B surface antigen and anti-HBs in Chinese chronic hepatitis B virus patients relating to genotype C and mutations in the S and P gene reverse transcriptase region. Arch Virol 157(4): 627-634.






























