Nanoplateforms of Mitochondrial Dysfunction Oxidative Stress in Cancers
Rakesh Sharma1*, Yashwant Pathak2, Arias Tzika1 and Anna Gradvojakova3
1Plastic Surgery Fellow, Shriner’s Children Hospital, 1Massachusetts General Hospital, Harvard Medical School, Boston MA, USA
2Taneja College of Pharmacy, University of South Florida, USA
3Pharmacobiochemical Laboratory of Third Medical Department, Medical Faculty, Comenius University, NMR Department, Slovak Technical University, Bratislava, Slovakia
Submission: November 08, 2023; Published: December 01, 2023
*Corresponding Address: Rakesh Sharma, 1Plastic Surgery Fellow, Shriner’s Children Hospital, Massachusetts General Hospital, Harvard Medical School, Boston MA, Email: rksz2009@gmail.com
How to cite this article: Rakesh Sharma*, Yashwant Pathak, Arias Tzika and Anna Gradvojakova. Nanoplateforms of Mitochondrial Dysfunction Oxidative Stress in Cancers. Canc Therapy & Oncol Int J. 2023; 25(4): 556167. DOI:10.19080/CTOIJ.2023.25.556167
Abstract
Mitochondrial major functions are ATP synthesis, regulation of Ca2+ homeostasis, generation of reactive oxygen species and programmed cancer cell death. They play as essential significant role in the generation of cell activation nano signals via released reactive species, as well as small GTPases, mitogen activated kinases located in the outer and inner membranes of cancer cell mitochondria. Several mutations in mitochondrial DNA led to synthesis of malfunctional proteins characterized by impaired energy metabolism due to presumed genetically based oxidative phosphorylation (OXPHOS) dysfunction. These proteins are potential nanoplatforms to indicate distinct cellular/organelle pathological changes. Several mitochondrial metabolic protein nanoplatforms screen the mitochondrial dysfunction induced oxidative damage to DNA, electron transport chain and structural proteins as etiologically important events in development and progression of mitochondrial driven cancer diseases. Literature data suggests several mechanisms of mitochondrial dysfunction indicated by protein and ROS nanoplatforms. One of the main challenges in using nanoplatforms seems to assess timely the functional/structural modulation to mitochondrial proteins and damage to DNA as potential causes or consequences of cancer diseases. Upon mitochondria dysfunction in oxidative stress mechanistic issues are resolved, use of mitochondria as a pharmacological nanoplatform target holds promise in the treatment of diseases. This paper presents a glimpse on application of mitochondrial nanoplatforms with newest supporting protein-based mechanisms of mitochondria in oxidative stress, inflammatory and apoptosis-based diseases, clinical management guideline and therapeutic failure if any.
Keywords: Mitochondria; Chemotherapeutics; Oxidative Stress; Lipid Metabolism; Oncometabolites
Abbreviations: OXPHOS: Oxidative Phosphorylation; TCA: Tricarboxylic Acid; SDH: Succinate Dehydrogenase; FH: Fumarate Hydratase; ROS: Reactive Oxygen Species; ETC: Electron Transport Chain; NOS: Nitric Oxide Synthase; TRAP-1: TNF Receptor-Associated Protein-1; HSP90: Heat Shock Protein 90; AML: Acute Myeloid Leukemia; IONP1: Ion Peptidase-1; PCR: Polymerase Chain Reaction; DHPLC: Denaturing High Performance Liquid Chromatography; DGGE: Denaturant Gradient Gel Electrophoresis; TGGE: Temperature Gradient Gel Electrophoresis; TTGE: Temporal Temperature Gradient Gel Electrophoresis; SSCP: Single-Strand Conformation Polymorphism; RFLP: Restriction Fragment Length Polymorphism; ASO: Allele Specific Oligonucleotide; FTICR-MS: Fourier Transform Ion Cyclotron Resonance Mass Spectrometry; ESI-TOFMS: Electrospray Ionization Time-of-Flight Mass Spectrometry; MERRF: Myoclonic Epilepsy and Ragged Red Fibers; PEO: Progressive External Ophthalmoplegia; POLG1: Polymerase Gamma; MRS: Magnetic Resonance Spectroscopy; TNBC: triple negative breast cancer; TFAM: Transcription Factor A Mitochondrial DNA; POLRMT: Mitochondrial RNA Polymerase; OMM: Outer Mitochondrial Membrane; CGN: Cerebellar Granule Neuron; ISP: iron-Sulfur Protein; LHON: Leber Hereditary Optic Neuropathy
Introduction
Mitochondria are unique energy storage organelles. Mitochondrial dysfunction and oxidative stress play a role in promoting tumor propagation and metastasis activity through ROS production and oncogenic transformation. Excessive mitochondrial oxidative stress triggers cell death in both transformed and non-transformed cells through proliferation, migration/invasion, angiogenesis, inflammation and cancer cell immnoescape towards adoption of new environment. As a result, oxidative stress causes malignant transformation to metastasis, dormancy and resistance to chemotherapeutics. In this process, ROS species and several proteins participate that serve as signals on nanoplatforms to detect physiological stresses and changes in metabolic states of the cell [1]. These signals on nanoplatforms respond to new metabolic requirements by controlling ATP levels, lipid metabolism and/or promoting oxidative signaling during cancer diseases, cell migration, inflammatory responses, nutrient starvation, low oxygen, and oxidative stress or tumor progression during other diseases [2]. In adaptive mechanism, cancer cell mitochondria sustain tumor dissemination and optimize the production of oncoproteins, oncometabolites, energy, Ca2+ metabolism and generation of reactive species during cancer progression [3]. Mitochondrial functions proteins perhaps control the cellular responses to oncogene stimuli and regulation of cellular metabolism [4]. Mutations in mitochondrial enzymes of the tricarboxylic acid (TCA) cycle, such as succinate dehydrogenase (SDH), fumarate hydratase (FH), and isocitrate dehydrogenase 1, and 2 suggests that mitochondrial dysfunction enhances tumor growth or promotes cancer progression. Mitochondrial function, including oxidative phosphorylation (OXPHOS), is essential for cancer cell viability because the elimination of cancer cell mitochondrial DNA (mtDNA) reduces their growth rate and compromises tumorigenesis. These physiological processes affect cancer growth by modulating biosynthetic and cell signal pathways, transcription factors. Cancer cells have high anerobic glycolysis capacity znd active metabolic and proteotoxic stress. Presently, mitochondrial stress response targeting serves as an anti-cancer therapeutic method based on tumor cell growth suppression and prevention of metastasis.
Mitochondrial Electron Transport Chain and Tumor Initiation
Elevated reactive oxygen species (ROS) byproduct of OXIPHOS due to defective ETC activity drives towards more oxidized state in signal transduction cascades regulating cell proliferation, differentiation, oncogenic transformation and death [2]. The excessive ROS can cause DNA mutations in ETC genes that may contribute to genomic instability. Various cellular processes suggest a tight control for ROS production and function. There are at least three distinct signal mechanisms for ROS production in malignant transformations: 1) in cancer cells mutations in superoxide and SDH, FH and ICDH1 and oncometabolites 2, Land D-2-hydroxyglutarate, succinate, fumerate, all these alter the signal transduction and mutation gene-expression regulation of hypoxia-inducing factor (cancer epigenome) towards low oxygen metabolic remodeling of malignant transformation. These accumulated oncometabolites inhibt Jumonji C domain-containing histone lysine demethylases; and suppress KG-dependent 10-11 translocation methylcytosine nucleotide dioxygenase that drives oncogenic transcriptional program. Thus high superoxide, ROS and oncometabolites contribute to malignant transformation (biosignal); 2), ROS are produced by various oxidases including lipoxygenases, cyclooxygenases, xanthine oxidase and nitric oxide synthase (biosignals); and 3) mitochondria are the most important source for ROS in every cell type, where 1-5% of the transported electrons are diverted to the formation of superoxide (O2•-) instead of water [5]. Thus, accumulated mitochondrial oxidative stress causes metabolic re-programming leading to enzyme defects as observed mutations in SDH, FH, ICDH and ROS sensitive bio signal nanoplatforms to monitor the mitochondrial electron transport chain dysfunctions in human tumors.
Defects in the function of the mitochondrial electron transport chain (ETC) have been monitored as remarkable characteristics of several pathological conditions ranging from ischemia-reperfusion damage [6], diabetes [7], hypertension [8], liver damage by hepatitis B [9], allergic inflammation [10], cancer [11], cardiac impairment due to aging [12], neurodegenerative diseases such as Alzheimer’s disease, Parkinson’s disease and amyotrophic lateral sclerosis [13], among others. Mitochondrial nanoplatforms may detect mitochondrial dysfunctions in above pathological conditions. A common feature among these pathological conditions is the increased generation of ROS, which in some cases arise not only from the mitochondrial ETC but from other biosignal sources like NADPH oxidases (Nox) enzymes, cytochrome P450 enzymes, cyclooxygenases or uncoupled nitric oxide synthase (NOS) [14]. Therefore, ETC dysfunction during these pathologies has been monitored using biosignal nanoplatforms as ROS attack on the different components of the ETC. Furthermore, despite the prominent role attributed to mtDNA mutations in mitochondrial dysfunction, the direct damage by ROS on membrane lipids and proteins may be more relevant biosignals since it has been observed by nanoplatform experiments that accumulation of damage in ETC components is not codified by mtDNA [15].
Usually, it is assumed that ROS react preferentially with the bio signal molecules on nanoplatforms surrounding their generation site due to the short lifetimes of most of the radicals. For example, the steady-state concentration of hydroxyl radical (OH•) is practically zero because it reacts with very high-rate constants with almost every type of biomolecule [16]. In contrast, superoxide radical (O2•-) is far less reactive with non-radical species and cannot cross membranes due to its negative charge. However, its reactivity and membrane permeability depend on the pH of the nanoplatform environment. Thus, at low pH values hydroperoxyl radical (HO2•), the protonated form of superoxide, can cross the membranes more readily than O2•- given its uncharged nature [16]. Hence, the low pH value generated by membrane potential in the intermembrane space might contribute to the formation of HO2• from extra-mitochondrial sources to enhance the deleterious effects of non-mitochondrial-generated O2•-. It has also been proposed that O2•- can indirectly exert damaging effects on mitochondrial function through the opening of the mitoKATP channels, which may lead to increased K+ influx, matrix alkalinization, swelling and ROS production [17]. Also, extra-mitochondrial-generated O2•- has harmful effects on the ETC through its combination with NO• to form ONOO-, which can cause damage to respiratory complexes [18] possibly via tyrosine nitration [19]. Therefore, mitochondrial ETC dysfunction due to unregulated production of O2•- by extra-mitochondrial bio signal systems such as Nox enzymes, cyclooxygenases or uncoupled NOS, may exacerbate and/or contribute to the loss of homeostasis observed in many of the pathological conditions mentioned above. For this reason, one of the main challenges in using mitochondrial nanoplatforms is to evaluate the exact contribution of the different extra-mitochondrial bio signal sources in ETC dysfunction during the development of diseases.
Mitochondrial ROS may be central in the cell death transduction pathways. Most investigators believe that most of the oxidants are produced by electron chain transport. The accumulation of high levels of ROS would induce necrosis, while programmed cell death (apoptosis) would be predominant in the presence of low levels of ROS. The ubiquinone site as nanoplatform in complex III appears as the major source of mitochondrial ROS production as this site catalyzes the conversion of molecular oxygen to O2•-, which can lead to the formation of other potent ROS such as hydrogen peroxide (H2O2) and OH•. The release of cytochrome c must lead to a breakdown of the mitochondrial electron flow downstream of the ubiquinone site which in turn would result in an increased generation of ROS [20]. The decision of a cell to undergo apoptosis is mediated by a complex integration of signal transduction pathways [21] (Figure 1). Rho GTPases have been implicated as bio signals in the apoptosis of many cell types, but the mechanism by which they act is not fully understood. Several groups have reported both positive and negative involvement of Rho proteins in apoptotic program, depending on the cell line investigated. Thus, human RhoA, RhoC, Rac1, Rac2, and Cdc42 proteins have been involved in the induction of apoptosis in diverse cellular systems, including fibroblasts [22], thymocytes [23], neurons [24], and epithelial cells [25].

Tumor Growth
i. Mitochondrial enzymes for amino acid, fatty acids, androgen, estradiol synthesis in cancer cells support rapid cell division and membrane integrity in breast and prostate tumor progression as oncogenic factors for genetic alterations and metabolic rewiring.
ii. Major changes happen to be hypoxia, glucose and NADPH deficiency in tumor cell mitochondria to cause stress on TCA cycle intermediates needed for amino acid, nucleotides, lipids. So, mitochondria in tumors utilize lactate, serine, glycine and fatty acid oxidation in the pyruvate deficient acidosis conditions.
iii. In hypoxic conditions, mitochondria activate reductive carboxylation of glutamine metabolism for anaplerotic circuitry of TCA intermediates that render cancer cells reliant on reductive carboxylation of glutamine-derived α-ketoglutarate to fatty acids and redox balance for proliferation.
iv. Serine hydroxymethyltransferase-2 for serine catabolism provides NADPH/NADP while phosphoglycerate dehydrogenase for serine synthesis enhances ketoglutarate to support cancer growth.
v. Tumor cells secrete lactate and absorb it to fuel OXIPHOS at low glucose conditions.
vi. Cytosolic lactate makes pyruvate by LDH and in mitochondrial pyruvate makes acetyl-CoA by pyruvate dehydrogenase.
vii. In cancer cells, acetyl-CoA synthesizes fatty acids, isoprenoids, cholesterol to support proliferation in stressful conditions.
viii. Mitochondrial pyruvate carriers mediate pyruvate oxidation as on-off switch between OXIPHOS and glycolysis to support proliferation in breast, prostate, colon and liver cancers.
Metastasis and Survival
In nutrient starved, low oxygen and cytotoxic oxidative stressed harsh conditions leads to cancer cell death. However, metastatic cells easily evade these stressors to cause irreversible mitochondrial damage. Above all, malignant cells acquire alterations to maintain mitochondrial integrity and immortality. Hypoxia affects survival and metastasis in low oxygen fluctuating hypoxia-inducing conditions by oncogenic signaling pathways during tumor progression. These hypoxia-inducing factors HIF- 1 trigger ROS production, proliferation, angiogenesis, migration, invasion, poor patient prognosis and therapeutic resistance. In cancer cells, the HIF-1 activates glucose transporters, hexokinase-2, pyruvate dehydrogenase kinase to accelerate glycolysis. HIF-1 reduces electron transport chain or mitochondriqal respiration by inhibiting pyruvate dehydrogenase to prevent oxidative stress. HIF-1 transcription is increased by mutations in TCA cycle SDH and FH genes to accumulate TCA cycle succinate and fumerate intermediates. These intermediates inhibit prolyl hydroxylase to disrupt HIF-1 activity. Cancer cells also activate HIF-1 to accelerate glycolysis and more ATP or relief from oxidative stress.
Mitochondrial Protein Quality Control in Cancer
Cancer cells evade environmental stress by monitoring the defensive mitochondrial repair defects before their cell death trigger. Initially, mitochondrial chaperones and proteases (mtUPR enzymes) degrade misfolded proteins to form protein aggregates in mitochondria by proteotoxic oxidative-hypoxic stressors. The heat shock protein 90 (HSP90) and TNF receptor-associated protein-1 (TRAP-1) chaperones are overexpressed in glioma cell mitochondria to control protein folding quality in tumor progression, survival and maintenance of OXIPHOS capacity during hypoxia and low glucose conditions. TRAP-1 induces HIF- 1 stabilization and reduces mitochondrial respiration to confer tumorigenic potential.
Mitochondria dynamics in cancer cells promote mitophagy and remove damaged mitochondria during cytotoxic stress and survival. During this mitochondrial damage, proapoptotic factors release out to induce apoptosis and turnover older to new cells. Higher Ion peptidase-1 (LONP1) and Caseinolytic mitochondrial matrix peptidase proteolytic subunit (C1pP) amounts are in matrix of human colorectal cancer, glioma and melanoma as correlates of proliferation, high tumor grade, poor patient prognosis and survival. Low oxygen induced lower LONP1 expression arrests proliferation by OXIPHOS reduction and degrade misfolded ETC subunits. Thus, LONP1 impairs mitochondrial respiration, suppresses cell proliferation and cell metastatic dissemination, increases oxidative stress in human acute myeloid leukemia (AML). The C1pP expression is increased in breast, lung, liver, prostate, thyroid and melanoma cancers. In fact, LONP1 and C1pP genes share OXIPHOS, amino acid and lipid metabolism to maintain proteins in cancer cells to survive and evade the metabolic, oxidative and cytotoxic stress depleted.
Mitochondrial Dynamic Changes in Cancer
Continuously, hormones and cell signaling pathways alter cancer cell mitochondrial dynamics during tumor progression and survival in stress. The mitochondria change their shapes by fusion mitofusin (Mfn) 1 and 2 and OPA1 proteins and fission dynamin protein-1, mitochondrial fission factor proteins after lipid oxidation and mtDNA mutations in mitophagy process in liver, colorectal, brain, lung, breast cancers. Above proteins regulate mitochondrial pyruvate carrier in oncogenic pathways:
i. ERK1/2 signaling pathways responsible for MAPKtransformed tumor growth in melanoma and pancreatic cancers.
ii. CDK5 activates dynamin protein-1 phosphorylation to show poor outcome in glioblastoma cancer.
iii. PI3K-Akt signaling in fragmented mitochondrial networks.
iv. Increased dynamin protein-1 expression or activated fission in lymph node metastasis in breast, thyroid, brain, prostate cancers.
v. Androgens hormones influence prostate cancer cell survival by androgen-receptor upregulated signaling in mitochondrial fission dynamics.
vi. The dynamin-1 protein enhances prostate cancer cell proliferation but weakens cell survival in hypoxic-oxidative stress, autophagy through voltage-dependent anion channelmitochondrial pyruvate translocation carrier complex and TCA cycle rewiring.
vii. The dynamin-1 protein expression activates MFN2 downregulation to promote tumor survival or cancer related mortality through fragmented mitochondria in triple negative breast cancer (TNBC) cell mitochondria in hypoxic stress.
viii. The dynamin-1 receptor protein activates receptor MFF-VDAC complex protein phosphorylation through AMPProtein Kinase in Myc targeting MFF and overexpressing it to induce mitochondrial fission, cell death in malignant primary and metastatic cancers. Thus, MFF upregulation and repression limits tumor growth by impairing cell division and loss of self-renewal.
Mitophagy in Cancer
Autophagosomes remove damaged mitochondria during hypoxia, nutrient deficiency and DNA damage in tumors by continued mitophagy. In case of mitophagy defects, dysfunctional mitochondria accumulate or excessive mitophagy induces loss of functional mitochondria by mitochondrial fission resulting with tumor cell death. Several mechanisms describe mitophagy:
i. An AMPK pathway initiates autophagy by unc-51, ULK-1 or TG1 (autophagy activating kinase-1 but AMPK inhibits rapamycin complex-1 (mTORC1). The absence of ULK1 in cancer cells does remove defective mitochondria and tumor cells die with oxidative stress-induced cell death by excessive lipid peroxidation.
ii. PTEN-induced kinase-1/ Parkin-mediated depolarized mitochondria accumulates PINK-1 on outer mitochondrial membrane to catalyze mitochondrial protein ubiquitination by E3 ubiquitin protein ligase and delivering defective mitochondria to autophagosomes.
iii. Parkin increases oxidative metabolic levels and inhibits aerobic glycolysis by p53 tumor suppressor during mitochondrial fission.
iv. Parkin binds to ubiquitin and degrades MFN2 (PINK1) during melanoma tumor formation and metastasis. The PINK1 silencing induces ROS overproduction to reduce lung cancer cell migration and its invasion capacity otherwise Parkin-deficient breast cancer cells show low proliferation and metastasis potential.
v. Many receptors NIX, BNIP3 and FUNDC1 bind to dynamin DRP1 on outer mitochondrial membrane to enhance mitochondrial fission during mitochondrial fission in hypoxic regions in glioblastoma for cancer stem cell maintenance, cell survival. The loss or inhibition of NIX slows down proliferation by redox homeostasis in pancreatic ductal adenocarcinoma.
vi. The BNIP3 inhibition reduces mitophagy in hypoxic adenoid cystic carcinoma invasion. Cisplatin promotes BNIP3-
NIX co-upregulation to support mitophagy in lung cancer cells and depletes FUNDC1 expression to inhibit cell proliferation in cervical and colorectal cancers.
vii. Parkin protein prevents mitochondrial fission and fragmentation by dynamin DRP1 ubiquitination.
Cell Death Regulation
i. Cancer cells struggle till highest mitochondrial integrity threshold and slowly die (apoptosis) and finished (necrosis) through common stress response pathways and survival promoting B-cell lymphoma BCL-2 proteins against oxidative stress death. Mitochondrial fragmentation network enhances mitochondrial threshold for cell death, tumor expansion, metastasis and drug resistance. It all happens in a sequence of 1. Pro-apoptosis and antapoptosis balance due to mitochondrial permeability loss and induced targeting BH3 downstream cell-death proteins such as antiapoptotic BCL-2, BCL-XL, MCL1 regulator proteins (decreased mitophagy) and pro-apoptotic BAX, BAK proteins and upstream BAD, BIM, PUMA, BID, NOXA regulator proteins, to sense apoptosis and membrane permeability.
ii. Anti-apoptosis BCL-2 proteins (BCL-XL and MCL-1 complex) bind Parkin and stop Parkin translocation to depolarized mitochondria (fragmented across permeable outer membrane) while BH3 proteins enhance Parkin translocation.
iii. Molecular chaperone HSP60 enhances tumor growth. The HSP60-BAX upregulation (low HSP60 expression) due to stress-induced mitochndria damage quickly depletes mtDNA activity (BAX mitochondrial accumulation and cytochrome C caspase activation) to induce apoptosis and pro-apoptotic p53 protein function to prevent cancer cell death.
iv. Chaperone HSP10 overexpression accumulates antiapoptotic BCL-XL and BCL-2 but reduces pro-apoptotic BAX.
v. Chaperone HSP90 inhibits cytochrome c-mediated protease cascade.
vi. Mitochondrial cyclophilin-D bound HSP90/60 promotes tumor growth, and its disruption causes caspase-dependent apoptosis.
vii. HSP27 overexpression delays caspase activation in etoposide treated cytotoxicity in human leukemia.
viii. High HSP70 inhibits mitochondrial BAX translocation, TRAP1 expression (silencing TRAP1 phosphorylation) and opens mPTP by apoptosis inducing drug to prevent stress-induced cell death. PINK1 protein phosphorylates TRAP1 and slows cytochrome c loss from mitochondria to protect cells from stress.
ix. BAX-Dynamin DRP1 colocalizes to reduce mitochondrial fission and caspase-dependent cancer cell death in autophagy.
x. In MFF deficient prostate cancer, AMPK signaling pathway and dynamin DRP1 depletion causes autophagy. The gene silencing or drugs inhibit autophagy and enhance caspasedependent cell death.
Mitochondrial DNA in Cancer
Circular 16569 bp sized mitochondrial DNA encodes 13 ETC OXIPHOS proteins, two rRNA and 22 tRNAs. Mitochondrial DNA homeostasis is balanced by mtDNA repair and replication, mt DNA copy number alteration, mtDNA mutations and regulation of transcription-translation of mtDNA in tumorigenesis, tumor survival during dissemination and metastasis. The oxidative damage and mtDNA replication errors cause mutations. Thus, mtDNA integrity is related to mitochondrial function. So, mitochondrial dynamics (mitochondrial fission-fusion) and mitophagy (mitochondrial stress response) maintain mtDNA integrity, copy number, shape by fission-fusion processes to remove damaged mtDNA and accumulated mutations in high stressed conditions. Thus, increased mtDNA damage by mitochondrial fusion and reduced mitophagy produce bioenergetic defects of metabolic disorders and cell death.
Mitochondrial DNA mutation and Copy Number in Cancer
Mitochondrial DNA is most abundant in ovarian and lung adrenocarcinoma cancers and least abundant in myeloid, bladder, breast, kidney cancers. These mutations show specific patterns:
i. The low mtDNA content, heteroplasmic single nucleotide variant or D-loop mutations and copy number reflect the high cancer risk of lymphoma, breast cancer and colorectal cancer patients.
ii. The mtDNA mutations in 16S rRNA, ND2 and ATPase 6 genes reflect breast tumor small deletions and insertions in coding and non-codng regions.
iii. Synonymous and non-synonymous mtDNA mutations regulated by mtDNA repair in non-oncogenic mutations to keep tumor energy metabolism up in stressed conditions in kidney tumorigenesis.
iv. Low mtDNA deletions in dysfunctional mitochondria and apoptosis sensitized tumors. It helps to remove cancer cells having large scale mtDNA mutations during early tumor progression.
Mitochondria DNA Transcription and Translation
The role of mitochondrial transcription and translation in Mito ribosomes during cancer progression includes:
i. The mitochondrial gene regulation by chemical modification in heavy strands after inhibition of mitochondrial transcription and translation to proteins in Mito ribosomes affects cancer progression and metabolism.
ii. Mitochondrial DNA transcription factor A (TFAM), mitochondrial RNA Polymerase (POLRMT), mitochondrial transcription B1, B2 factors facilitate transcription in cancer. IMT1 inhibitors of POLRMIT sensitive OXIPHOS system inhibit mtDNA expression and OXIPHOS in tumors.
iii. The G1 arrest and growth inhibition through ROSinduced JNK/38 MAPK signaling leads to downregulation of TFAM in lung cancer cells.
iv. TFAM inhibition induces apoptosis and increases cisplatin sensitivity to mtDNA transcription in non-small lung cancer treatment.
v. All above antitumor effects of mitochondrial transcription inhibitors increase mtDNA gene expression to make more cellular metabolites by inhibiting mtTFAM downregulation for resistance in tumors.
vi. Orthogonal RNA sequencing shows lower mitochondrial RNA levels in breast, esophageal, head-neck, kidney cell, liver cancers.
vii. mitochondrial DNA copy number and lower mitochondrial RNA expression compensates for low mitochondrial RNA transcript levels.
viii. The abnormal translation (altered initiation, elongation mtEF4 factor, termination, ribosome recycling, translation factors) of mitochondrial encoded proteins, mtDNA polymerase in OXIPHOS system and gene mutations or deletions may cause apoptosis in leukemia cancer progression through lower cellular redox, mtRNA expression. Some of the above events may serve as nanoplatforms in oxidative stressed cancer detection and monitoring.
Mitochondrial Oxidative Stress Cancer and ROS induced Immunoescape
During oxidative stress, reactive oxygen species (ROS) are formed in mitochondria and mediate proliferation, migration/ invasion, angiogenesis, inflammation and immunoescape in cancer cells to adapt to the stressed environment. ROS induces immunosuppression through immune cells, M2 macrophages, dendritic cells and T-cells to mitigate oxidative stress. Stress specific genome instability (replicative and mitotic stress), hypoxia (metabolic stress and angiogenesis), ROS production (mitochondrial stress) may cause carcinogenesis to support malignant behavior. Mitochondrial Lon-ROS supports cell proliferation, migration, angiogenesis, resistance towards apoptosis, inflammatory cytokine formation, mtDNA/Evs to escape immunosupressive environment in cancer. ROS-modulating anticancer immunotherapies show anticancer effects to avoid immunoescape and recurrence. In the next section, mitochondrial ROS signaling is described during cancer progression.
Rho Family GTPases in Mitochondria in Cancers
The Rho family GTPases are monomeric G-protein bio signals. These interact with intracellular target proteins or nanoplatform effectors to trigger a wide variety of cellular responses. Several responses may be the reorganization of the actin cytoskeleton, membrane trafficking, cell cycle progression, gene transcription, adhesion, migration, metastasis, and cell survival [26]. The three most studied Rho family member proteins as bio signals are RhoA, Rac, and Cdc42. Western blot analysis indicated the localization of Cdc42, Rac1, and Ras proteins in the mitochondrial fraction of pancreatic beta cells [27]. Besides, the presence of the Miro proteins (for mitochondrial Rho) has been identified as bio signal. Miro family members (1 and 2) function in a novel pathway that integrates cell bio signaling with mitochondrial dynamics and functions. These bio signal proteins are localized at the outer mitochondrial membrane (OMM).
Both bio signal proteins consist of 618 amino acid residues and were found to be 60% identical to each other [28]. Miro proteins are present in yeast (Gem1p) [1], Drosophila (dMiro) [29], and mammalian cells as well [28]. They have two potential Ca2+ binding domains, so called EF-hands. Functional EFhand motifs exposed to the cytoplasm would be able to sense changing cytosolic calcium levels, whereas localization of these motifs to the mitochondrial matrix could allow Miro to monitor organellar calcium levels [1]. In neurons, Miro protein supports the distribution of mitochondria to the dendrites close to the Ca2+ entry sites and promotes the calcium bio signal propagation to the mitochondria [4]. The transport of mitochondria to specific neuronal locations is critical to meet local cellular energy demands and for buffering intracellular calcium levels. Miro1 and the kinesis adaptor Grif-1 play an important role in regulation of mitochondrial transport in neurons through the movement along microtubules. Thus, Miro1 is a key determinant of how energy supply is matched to energy usage in neurons [30].
Rho Proteins and Apoptosis in Cancer
In cardiomyocytes, RhoA activation bio signals act through effector Rho kinase to induce p53-mediated increases in the level of activated Bax protein and an increase in the amount of Bax associated with the mitochondria. Over time, however, Bax up-regulation appears to be sufficient to shift the balance in Bcl-2 family protein expression, induce mitochondrial permeabilization, facilitate cytochrome c release, initiate activation of caspase-9 and -3, and ultimately induce DNA fragmentation and cell death. The sustained RhoA activation (48h) can also increase the mitochondrial distribution of Bax, and increased RhoA bio signal protein activity following ‘myocardial infarction’. It suggests that RhoA bio signal activation could play a role in ischemia/ reperfusion-induced cell death [31]. In addition, the involvement of RhoA protein in hypertrophy of neonatal ventricular myocytes has been established [32]. Cultured cerebral cortical neurons undergo mitochondria-mediated apoptosis upon exposed to phenylalanine effector. The effector induces the RhoA/ROCK activation. Inhibition of the RhoA/ROCK bio signal pathway with effector C3 transferase or Y-27632 prevented the phenylalanineinduced caspase-3 activation (using nanoplatform) with result of rescued neurons from apoptosis. It indicated that the RhoA/ROCK bio signal pathway plays an important role in phenylalanine-induced neuronal apoptosis [33]. Membrane blebbing, cleavage of caspase 3, and release of cytochrome c from the mitochondria are unique nanoscale bio signals reduced by ROCK inhibition mechanism. In clinical practice, inhibition of Rho/ROCK bio signals may improve the survival of grafted cells in cell replacement therapy [34]. Rho proteins as nanoscale bio signals play an important role in physiological regulation of the apoptotic response to stress-inducing test drugs or agents because a dominant-negative mutant of Rac1 interferes with the induction of apoptosis by tumor necrosis factor-α [22]. Other studies indicate that over expression of Cdc42 induced apoptosis in Jurkat cells [35]. Transgenic mice carrying an activated version of the Rac2 gene nanoplatform showed an increased apoptosis in the thymus. In fact, it was the first in vivo demonstration of the role of Rho protein bio signal in apoptosis [23].
How the expression of Rac1 gene can induce apoptosis on nanoplatform?
The gene expression towards apoptosis takes place by a mechanism that involves mRNA and protein synthesis, activation of caspase-3 but not caspase-1. It may require the simultaneous production of ceramides and the synthesis of FasL protein by JNK- and NF-kB-dependent signals with no release of cytochrome c from mitochondria. In fact, an independent bio signal pathway was observed before the onset of the apoptotic program. Possibly, it was suggested as a pro-apoptosis pathway independent of cytochrome c release [36]. The pro-apoptotic role of Rac protein was correlated with a loss of plasma membrane-associated Rho family GTPases. These pro-apoptosis induction towards the apoptosis was reported in primary cortical neurons and brain neuroblasts [37]. Other pro-apoptosis inactivation of Rac GTPase stimulated the Bax protein translocation to mitochondria. Concurrently, it promoted a conformational change in Bax protein that is commonly associated with its cytochrome c-releasing activity [38]. While loss of Rac enzyme activity coincides with induction of apoptosis, Cdc42 activity is regulated independently of apoptotic signals. These indicate that Rac is the major Rho family GTPase. It is involved in promoting cerebellar granule neuron (CGN) survival and secondary role for Cdc42 [38].
The Cdc42 protein is a major player in neuronal cell death. It involves the up regulation of the JNK pathway [24]. Two other proteins Ras and Rac1 are essential for Fas-mediated apoptotic death of Jurkat cells [39]. The ability of these GTP binding proteins to induce apoptosis originates from their ability to activate downstream stress-activated kinases such as p38 and JNK proteins [40]. The Rac1 and Cdc42 proteins belong to the same family of Rho GTPases. Only Rac1 protein activates the p38 pathway. Thus, regulation of specific targets/signals by Myc upstream of Rho GTPases leads to specific activation of Rac1 and not Cdc42 [21]. Aberrant neuronal apoptosis is the sole cause of onset and progress of neurodegenerative disorders of Parkinson’s disease and amyotrophic lateral sclerosis [38]. The Rho proteins may possibly regulate apoptosis [36].
Rho Protein Biosignals and ROS Species in Cancers
The ROS species are generated from nonenzymatic superoxide by-product of mitochondrial metabolism. The integrin proteins engage in mitochondria through the activation of Rac protein to change the mitochondrial function towards elevated levels of ROS [2]. The small GTPase Rac species mediates the ROS production in diverse signaling pathways. Eventually, ROS species trigger mitogenesis, gene expression, and stress responses [41]. Some are:
i. The ROS production in neutrophils evidenced the Rac2 binding to a NAD(P)H oxidase in response to above events [42].
ii. In Ras-induced NIH 3T3 cell transformation, Rac1 protein was necessary to induce the ROS production [43].
iii. The ROS species activated the Rho/ROCK pathway. The ROS species lead to vasoconstriction. In experimental animal systemic arteries evidenced a cold-induced vasospasm (Raynaud`s phenomenon). It was exogenous H2O2 activated Rho kinase to increase the expression of ROCK-1 protein. The rise in PO2 increased the endogenous production of ROS in the mitochondria. The O2-dependent increased mitochondrial ROS might trigger the activation/induction of Rho-kinase [44]. The ROS inhibits the RhoA protein function by inducing the tyrosine phosphorylation of the RhoA GTPase activating protein p190Rho- GAP [45]. The exogenous ROS activated the RhoA in the smooth muscle of rat aorta [46].
Iron-Mediated Mitochondrial Dysfunction and Nanoplatforms
Another cellular oxidative stress bio signal is a free iron. Free iron causes several pathologies of Parkinson´s disease [47,48], neoplastic cancers [49,50], haemochromatosis [51], type II diabetes [52], cardiac and hepatic failure [53] and Friedreich’s ataxia [54]. Free iron catalyzes the formation of highly toxic ROS like OH• radical, iron-centered ferryl and perferryl radicals [55,56]. Mitochondria plays an important role in iron mediated toxicity. The iron-driven oxidations require the interaction of iron metal with O2•- and H2O2. The mitochondrial ETC is major source of O2•- and H2O2 radicals [53]. Thus, mitochondrial compartments regulate iron homeostasis. The iron-sulfur (Fe-S) clusters and iron in protoporphyrin IX, both participate in heme synthesis in mitochondria [57]. Iron controls iron transport across the plasma membrane (i.e., Aft1/2p transcription factors), and transport into the mitochondrion (i.e., Mmt1/2p and Mrs3/4 transporters) to make Fe-S clusters. Inside mitochondria, iron is sequesterized by human mitochondrial protein frataxin. This protein participates in assembling of Fe-S clusters with other mitochondrial scaffold proteins Isu1/2p and ferredoxin Yah1p, the ferredoxin reductase Arh1p, and chaperone pair Ssq1p and Jac1p, to transfer the assembled Fe-S clusters from Isu1/2p scaffolds to apo-Fe-S proteins [57,58]. Thus, quantity of bio signal proteins in mitochondrial iron metabolism reflects the management of minimum mitochondrial free iron concentration to avoid mitochondrial ROS generation.
In an inherited neurological disorder FRDA, disrupted iron handling occurs in the mitochondria due to mutations in the frataxin gene [59]. Mitochondria in cultured fibroblasts and mitochondria from FRDA patients had more iron than mitochondria from control fibroblasts. Interestingly, cardiac and neural tissues had more iron deposits [59]. In affected tissues, a decrease in several Fe-S containing proteins was reported [60]. Moreover, skeletal muscle from FRDA patients presented a decreased oxidative metabolism. The FRDA patients had mutation in human frataxin gene with accumulated high levels of iron [61], reduced activity of several Fe-S mitochondrial proteins and defective mitochondrial respiration [62].
Molecular Aspects of ETC Dysfunction by Lipoperoxidation
As mentioned above, mitochondria are exposed to oxidative attack during several pathological conditions characterized by an increment in cytosolic and mitochondrial ROS production. Thus, exhaustive research has been done about the effects of ROS on ETC complex function and the mechanism by which oxidative alterations on ETC components lead to an increase in ROS production with the objective to identify the main molecular targets of ROS attack to design future strategies to minimize the damage in mitochondrial function and improve the bioenergetics status of the cell during such diseases.
Numerous reports have identified lipoperoxidation and sulphydryl group oxidation as the main factors that decrease ETC activity under oxidative stress conditions generated in vitro or during pathological situations. In the case of lipoperoxidation, this process has been generally stimulated through the addition of nanomolar and micromolar concentrations of Fe alone [63] or in combination with ascorbate [13,15,64,65]. ADP/NADPH [66] or ADP/ascorbate [67]. The mitochondrial lipoperoxidation has been observed with H2O2 [68], O2•- [69] and ischemiareperfusion models as well [70-72]. A remarkable feature of the mitochondrial inner membrane is the presence of cardiolipin, a tetra-acyl phospholipid essential for the proper structure and activity of ETC complexes except in succinate dehydrogenase (complex II) as recently reviewed [73]. Thus, lipoperoxidation has been associated with cardiolipin loss and inactivation of NADH: ubiquinone oxidoreductase (complex I) [72,74], ubiquinol: cytochrome c oxidoreductase (complex III) [70,71,75] and cytochrome c oxidase (complex IV) [76-78]. In agreement with the lack of cardiolipin requirement for proper function of complex II, lipoperoxidation does not participate in complex II inactivation given that the treatment with the anti-lipoperoxidative agent butylated hydroxytoluene cannot protect enzymatic activity the treatment with Fe2+ or Fe3+/ascorbate/ADP [63,67].
Another component of ETC, highly sensitive to lipoperoxidation, is the liposoluble electron carrier ubiquinone, which transfers electrons between complexes I and II to complex III. Oxidative modifications that impair the ability of ubiquinone for electron transfer, as well as the loss of the ubiquinone pool, have been associated with inactivation of partial reactions involving ubiquinone, i.e., NADH- and succinate-ubiquinone and cytochrome c reductases [67]. This fact and the finding that ubiquinone is more important for the protection of ETC against oxidative inactivation than the liposoluble antioxidant vitamin E, have led to the proposal that ubiquinone serves as an important lipophilic antioxidant system in mitochondria besides its function as an electron carrier [66]. Indeed, the maintenance of the ubiquinone pool in a reduced state through the feeding of ETC with succinate prevents lipoperoxidation and ubiquinone loss [64,67]. However, the ubiquinone loss during lipoperoxidation progression seems to be controversial since it has been observed in liver mitochondria treated with iron/ascorbate that the total ubiquinone content (reduced plus oxidized forms) remains constant and only the reduced fraction becomes exhausted. This observation seems to be tightly correlated with a rapid decay in the content of vitamin E [64]. Although this fact is also controvertial with the proposal that vitamin E is less important than ubiquinone for ETC protection as mentioned above, it was suggested instead that vitamin E and ubiquinone act through a synergistic fashion, where ubiquinone would act as a substrate to convert vitamin E radicals to reduced vitamin E and ubiquinone would not act as an antioxidant per se [64].
Despite the importance of ubiquinone and cardiolipin damage during lipoperoxidative injury to ETC, the damage to proteins is also relevant to the mechanism of ETC inhibition by lipoperoxidation since it is well known that lipoperoxidation inactivates several enzymes bound to membranes due to a decrease in membrane fluidity and the reaction of polypeptidic chains with several products of lipoperoxidation [79,80]. The analysis of the protein content of ETC complexes from liver mitochondria during lipoperoxidation induced by iron/ascorbate have revealed that protein degradation is more rapid and pronounced in complex I than in complex III, while complex IV is the least sensitive complex to lipoperoxidation [65]. These observations agree with the facts that these complexes require interaction with cardiolipin for a proper function and conformation [73] and hydrophobic antioxidants like butylated hydroxytoluene or vitamin E protect their activity from oxidative inactivation [63,81]. However, these observations do not correlate with the extent of loss of complex I and III activity observed during the treatment of rat liver mitochondria [64,82,83] and synaptosomes with iron/ascorbate. In contrast, the rate of complex IV and complex III degradation correlates well with the degree of its inactivation in beef heart [67], liver submitochondrial particles [66], rat testis mitochondria [15], liver mitochondria [64], gerbil liver [83] and Saccharomyces cerevisiae mitochondria with manipulated fatty acid content [63] treated with Fe3+/ascorbate/ADP, Fe3+/NADPH/ADP/, Fe2+/ ascorbate, Fe2+/ascorbate, subcutaneous injection of iron/dextran and Fe2+ alone, respectively.
It has been suggested that the lack of correlation between the decline in the complex III activity and the rate of its protein degradation could be due to specific impairment of the redoxactive centers (i.e., Fe-S clusters or cytochromes) caused by changes in phospholipid-protein interactions induced by lipoperoxidation [64]. In agreement with this idea, some studies have found that treatment with Fe3+/ascorbate/ADP or Fe2+ impairs electron transfer to cytochrome b without altering the concentration and structure of b hemes [63,67]. Furthermore, the analysis of cytochrome b reduction in mitochondria from the yeast S. cerevisiae, confirms that impairment in electron transfer to b hemes can be attributed to lipoperoxidation, because the treatment with Fe2+ does not impair cytochrome b reduction by succinate or dithionite in this organism [63], which is intrinsically resistant to lipoperoxidation due to its membrane fatty composition constituted by saturated and monounsaturated fatty acids [84].
Iron treatment also affects the content of hemes in both yeast and rat models. However, in contrast with hemes b, this effect cannot be attributed to lipoperoxidation given that c+c1 hemes loss was also observed in yeast mitochondria [63,67]. The alteration on electron transfer to cytochrome b by lipoperoxidation is consistent with its hydrophobic character [85] and the requirement of this subunit for lipids to achieve an adequate stabilization of its structure as delipidation of complex III impairs antimycin A binding in a reversible way through cardiolipin addition [86]. Therefore, these observations indicate that lipoperoxidation possibly alters electron transfer through destabilization of cytochrome b conformation without altering heme b center. However, this may impair redox potential of b hemes to an unfavorable value that impairs electron transfer to these prosthetics’ groups. A mechanism has been proposed by which lipoperoxidation may impair complex III activity and promote ROS generation in the basis of knowledge in the Q-cycle operation and the next observations:
i. Lipoperoxidation is associated with increments in the production of ROS [12,63,70,71,75].
ii. Lipoperoxidation impairs electron transfer to cytochrome, but it may not affect ubiquinol oxidation at the QO site [63,67].
iii. Lipoperoxidation induces resistance to the inhibitory effects of antimycin A in cytochrome c reduction [63].
iv. The subunits that constitute QO site require specific interactions with membrane phospholipids to achieve a proper conformation and function [86,87].
v. The disruption in the electron transfer between b hemes impairs the movement of the head domain of reduced iron-sulfur protein (ISP) [88].
In such a mechanism, lipoperoxidation would affect cytochrome b conformation, disrupting electron transfer to b hemes and the ISP movement between heme bL and cytochrome c1, which in turn, would lead to decreased catalytic cytochrome c reduction and accumulation of electrons at the QO site. The failure in electron transfer to cytochrome b and the impaired ISP movement would lead to a leak of electrons arising from partial ubiquinol oxidation by ISP to yield ubisemiquinone, which may react with oxygen to produce O2•-.
Thiol Oxidation in the Mechanism of ETC Inhibition by ROS in Cancer
Thiols (-SH) have been known to play an important role in regulating cellular function through redox changes in cysteine residues from peptides and proteins. Mitochondrial function seems not to be an exception since the existence of reactive – SH groups from cysteine residues of ETC complexes has been demonstrated which modulate their enzymatic activity through redox changes under oxidative stress conditions. Many ROS oxidize -SH directly to thiyl radical, yielding different protein -SH alterations including formation of mixed disulfides or internal disulfides from vicinal dithiols, S-nitrosation, and the formation of higher oxidation states [89]. H2O2, one of the most important ROS produced from the ETC, does not react at physiologically significant rates with -SH unless the reaction is catalyzed or under conditions that promote formation of thiolate (S-) species, which react with hydroperoxides at rates varying from ~10–105 M-1 · s-1 [90]. Thus, the reaction of –SH in mitochondria with H2O2 is highly favored by S- formation due to the high matrix pH (~8.0) and pKa values of protein -SH of 8–8.5 [91]. Nonetheless, although S- rather than -SH reacts with H2O2, the rate constants for some Sare still relatively slow and may require catalysis [90], which may explain the different sensitivity of –SH to oxidation as remarked below.
Proteomics approaches have demonstrated the presence of –SH on ETC complexes with different sensitivity to ROS attack. For example, in a study carried out in the presence of exogenous oxidants or stimulators of endogenous ROS production, it was found that core protein I of complex III is the only protein from the ETC that contains thiols sensitive to oxidation stimulated by endogenous ROS. Since two-dimensional electrophoresis may restrict detection of hydrophobic membrane proteins, there were no other identified ETC proteins with sensitive –SH groups [92]. In another study, the reactivity of thiol groups with lipophilic cations which accumulate in mitochondria in response to membrane potential and form stable adducts in a thiol-specific reaction was evaluated. This approach showed the presence of free reactive -SH residues on the matrix surface of complexes I, II, and IV [91]. Therefore, it can be concluded from these studies that all complexes from the ETC contain free -SH residues prone to oxidative modifications. Several functional studies indicate that reversible –SH oxidation modulates ETC activity at the level of all ETC complexes, except in complex IV. For example, the treatment of yeast mitochondria or rat synaptosomes with micromolar concentrations of hydrogen peroxide H2O2 or ascorbate/iron, respectively, inhibit activities of complex II and complex III [68,81]. A role was attributed to protein –SH oxidation as treatment with the thiol reductant β-mercaptoethanol in yeast almost completely reverted the inhibition in both complexes II and III [68], while in synaptosomes, glutathione alone prevented complex II inhibition and a mixture of glutathione and lipophilic antioxidants prevented complex III inhibition, suggesting that in complex III, lipoperoxidation and –SH oxidation impair enzyme function [81]. The presence of reactive –SH groups in complex II agrees with earlier reports about the existence of two classes of –SH groups in complex II with different reactivities to alkylating agents, with the most sensitive groups being those localized in the substrate site of the enzyme [93].
Despite the presence of reactive –SH groups in complex IV [91], these groups do not seem to participate in activity modulation since the treatment of mitochondria with milimolar concentrations of H2O2 or H2O2/protein ratios as high as 3000 nmol/mg protein caused a null or slight loss of activity, respectively [94]. One possible explanation for this phenomenon may reside in the fact that complex IV can function in the presence of H2O2 as a peroxidase using reduced cytochrome c as an electron donor. During complex IV turnover, the dioxygen bound to the Fe3+ atom from heme a3 (Fe3+-O-O=) undergo a double protonation to give a peroxide (Fe3+-HO-OH), which is unstable and suffers immediate heterolytic cleavage resulting in H2O release [95]. In a similar fashion, H2O2 can bind to heme a3, forming the complex Fe3+-HO-OH and undergo a heterolytic cleavage in the same way as the intermediate peroxide generated during normal complex IV turnover [96]. Thus, H2O2 can be considered a substrate of complex IV rather than an oxidant agent that modifies its structure and function, which may explain the observed resistance of this complex to high H2O2 concentrations.
In contrast, complex I activity is highly sensitive to –SH oxidation as demonstrated in dopaminergic cells subjected to oxidative stress through glutathione depletion. Because complex inhibition can be reversed by addition of the thiol reductant agent dithiotreitol, the participation of –SH oxidation through mixed disulfide or intramolecular disulfide formation from vicinal thiols has been suggested [97]. Additional evidence that complexes I is inactivated by –SH reversible oxidation due to glutathione depletion has been obtained from mitochondria subjected to ischemia-reperfusion [98] or haloperidol exposure [99]. A possible role for reactive –SH groups in the protection of complex I against oxidative damage has been proposed based on the observation that complex I activity is inhibited without promoting ROS production by selective glutathionylation of only two of six free cysteine residues (Cys-535 and Cys-704). Glutathionylation of these exposed surface –SH groups could proceed through oxidation of –SH to sulfonic acid and its posterior reaction with reduced glutathione to form a mixed disulfide, which then reacts rapidly with GSH in a reaction catalyzed by mitochondrial glutaredoxin Grx2 to regenerate thiol, thus creating a cycle of –SH oxidation-regeneration that may protect other essential amino acids residues from oxidative damage [100]. Therefore, the hypothesis is plausible that -SH groups from ETC complexes may serve as a redox sensor that modulates ETC activity in function of the redox state of mitochondria. This could decrease ROS generation by limiting electron fluxes to mitochondrial sites of ROS generation (i.e., quinone sites of complex III). The operation of this system would be optimal under high GSH/GSSG ratios and moderate levels of oxidative stress. Otherwise, -SH modification would be irreversible due to further –SH oxidation to sulfinic or sulfonic acid species, thus inactivating the protection to ETC conferred by -SH from oxidative damage to essential amino acid residues and leading to irreversibly damaged ETC complexes and further ROS formation. Hence, this picture may also, in part, explain the irreversible damage to ETC proteins observed under strong oxidative conditions like iron overload. However, it must be pointed out that although sulfinic or sulfonic acid formation in proteins has long been viewed as an irreversible process, it has been demonstrated that sulfinic acids can be reduced to thiols in vivo by an enzyme with sulfinic acid reductase activity called sulfiredoxin [101].
Despite the growing body of data showing the direct interaction of H2O2 with –SH groups, there is some contradictory evidence indicating that H2O2 inhibits ETC function without directly altering the enzymatic activity of ETC complexes [102,103]. In heart mitochondria, H2O2 exposure inhibited respiration in state 3 using glutamate/malate as a substrate. This effect was attributed to inhibition of α-ketoglutarate dehydrogenase activity and a consequent decrease in NADH production to feed the ETC. A short time of H2O2 exposure (3 min) may account for the lack of direct effects on ETC function, since in other reports, longer incubations were needed to observe direct effects on ETC activities. Similar results were obtained using α-ketoglutarate as a substrate, but in this case, succinate dehydrogenase and aconitase were also susceptible to H2O2 inactivation. Remarkably, enzyme activities and respiration were recovered several minutes after oxidant addition except for aconitase. Since this phenomenon was only observed in intact mitochondria but not in solubilized mitochondria, it was suggested that inactivation of enzymes was due to glutathionylation rather than direct interaction of H2O2 with proteins, while NAD(P)H depletion was also explained because of its utilization for a system present only in intact mitochondria (i.e., glutaredoxin) that carries out reversion of glutathionylation [102]. In summary, the ambiguous effects of H2O2 on mitochondrial function may be related to the concentrations and time of exposure used as well as whether H2O2 was added exogenously or generated through the inhibition of ETC complexes or reverse electron transfer, which may influence the proximity of –SH groups to the site of H2O2, its reactivity or its conversion to thiolate form.
Mitochondrial Dysfunction Evaluation Methods
Mitochondrial diseases are genomic in origin with multiorgan system manifestations and phenotypes. Diagnostic evaluation includes blood analytes and tissue histochemical measurements, neuroimaging, enzymatic assays in tissues or cultured cells and DNA analysis indicating grade of disease burden and its progress according to mitochondrial society in-depth evaluation stretagy [104]. Primary mitochondrial disorders follow patterns of maternal inheritance (i.e., mitochondrial DNA (mtDNA) mutations) or classical Mendelian inheritance (i.e., nuclear DNA (nDNA) mutations). Mitochondrial disease in children is caused by nDNA autosomal recessive pattern of inheritance confirmed by mtDNA analysis and molecular testing.
The biochemical analysis is done by lactate and pyruvate analysis. For it, sample is taken 30 minutes after placement of an intravenous catheter. Lactic acid is seen minimally elevated in mitochondrial polymerase gamma (POLG1)-associated diseases, Leber Hereditary Optic Neuropathy (LHON), Leigh disease, Kearns-Sayre syndrome, and complex I deficiency [105]. CSF lactate is elevated in CNS infection, stroke, malignancy, inflammation, and seizures [106]. Blood lactate/pyruvate ratios reflect the NAD+/NADH cytoplasmic redox state. Pyruvate is quite unstable. Blood and/or CSF pyruvate may be elevated in pyruvate metabolic defects such as pyruvate dehydrogenase deficiency, pyruvate carboxylase deficiency, or biotinidase deficiency. CSF lactate or pyruvate levels may be elevated in mitochondrial disease patients with predominant brain manifestations [107]. Pyruvate metabolism defects are genetic based mitochondrial diseases of oxidative phosphorylation dysfunction [108,109].
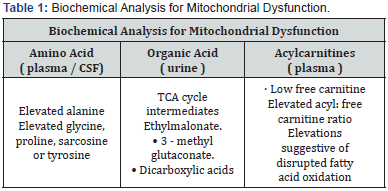
Amino acid analysis is performed using tendem mass spectrometry (MS/MS) for amino acid analysis of any change in cystine, homocystine, lysine, tryptophan, and GABA neurotransmitters involved in methylation reactions. Reversephase HPLC is another high throughput method for amino acid analysis. Elevated alanine: lysine ratio > 3:1 and alanine: phenylalanine + tyrosine (normal ratio < 4:1) indicate true hyperalaninemia [110]. An absolute elevation in alanine above 450 uM indicates Nijmegen diagnostic protocol. Aminoaciduria with renal tubular acidosis and glycosuria indicates renal Fanconi syndrome with mtDNA deletions [111]. Alanine, proline, glycine, and sarcosine amino acids elevate in mtDNA depletion syndrome (OXPHOS disease with severe infantile-onset liver disease). Elevated tyrosine withourt rise in succinylacetone in newborn screening studies indicate type 1Tyrosinemia [112].
Methods of organic acid quantification mass spectrometry extracted ion ratio method and semi-quantitative analysis utilizing gas hromatography flame ionization with mass spectrometry identify organic acid peaks [113]. Elevated organic acid mevalonate indicates diagnosis of mild mevalonic kinase deficiency or hyper IgD syndrome). Elevated CSF organic acid analysis has utility in select disorders of GABA metabolism. Urine organic acid analysis is routinely obtained in infants with suddenonset encephalopathy, organic acidopathies and dicarboxylic acid in β-fatty acid oxidation disorders. Increased excretion of tricarboxylic acid (TCA) cycle intermediates, ethylmalonic acid, and 3-methyl glutaconic acid jndicate renal immurity due to microsomal fatty acid oxidation [114-116].
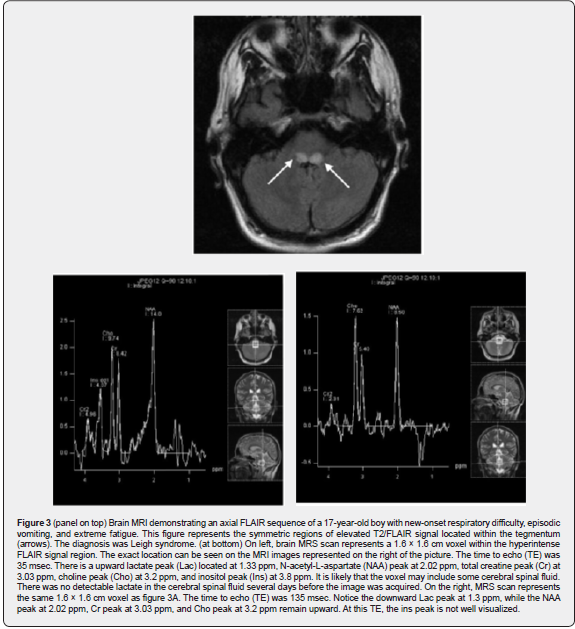
Blood total and free carnitine levels along with free and acylcarnitine ester profiling indicates fatty acid oxidation defects of primary aminoacidemias, organic acidemias, secondary fatty acid oxidation defects and carnitine deficiency in primary OXPHOS disorders. Quantitative acyl-carnitine analysis is performed by either tandem mass spectrometry (MS/MS) or HPLC followed by electrospray ionization (ESI)-MS/MS [117]. A filter paper blood spot can be used. (Figures 2-4).
Proton magnetic resonance spectroscopy (MRS) is used for neurometabolic evaluations of mitochondrial diseases [118-124] to detect lactate (1.33 ppm), N-acetyl-L-aspartate (2.02 ppm), succinate (2.39/2.40 ppm), total creatine (3.03 ppm), choline (3.22 ppm), and myo-inositol (3.55 pm). The author established a method of metabolite concentration based on peak area proportional to the number of spins to calculate disease burden. The prominent lactate peak resonance at 1.33 ppm at short TE times (i.e., 35 milliseconds) is inverted down at intermediate TE times (i.e., 135 milliseconds) and long TE time (i.e., 288 milliseconds) to reduce lipid contamination. Clinical spectroscopy centers still report peak area ratios. Lactate is elevated in anaerobic glycolysis and Leigh syndrome or mitochondrial myopathy, encephalopathy, lactic acidosis and stroke-like episodes (MELAS) in range of 0.5 – 1 mM as shown in Figure 3 by Barkovich [118-123]. Patients with respiratory chain deficiencies showed blood lactate elevated levels with moderate sensitivity18 % to 27% depending on acute, subacute, or chronic [124].
N-acetyl-L-aspartate (NAA) in neurons, axons, and dendrites is synthesized within the mitochondria and decreased NAA indicates neuronal integrity loss in mitochondrial diseases. The NAA/Cr ratio is different in cerebellum in 93% and in cortical gray matter regions in 87%, appearing normal on MRI [125]. The NAA/Cr ratio signal changes in patients with detectable Lac peaks. In some studies, the apparent decrease in the NAA/Cr signal was reversible [126,127]. As with elevated Lac, decreased NAA is not specific for mitochondrial or even metabolic disease [128,129]. The choline (Cho) signal on MRS includes free choline, phosphorylcholine, and phosphatidylcholine in brain myelin and cell membranes [130,131]. Cho elevation reflects membrane turnover and demyelination in patients with mitochondrial disease. The Cho/Cr ratio in 40% of patients showed reduced NAA/Cr and 53% of patients showed reduced NAA/Cr ratio with elevated Lac [132]. Succinate is present in brain at low concentration 0.5 mmol/kg wet weight [140]. Succinate peak is not visible on MRS [133] due to overlapping with glutamate and glutamine [134]. Significant increases in 2.4 ppm succinate peak by multiple voxel MRS indicated complex II deficiency with reduced NAA/Cr in white matter and cortical gray matter and increased Lac signals within white matter [135]. This highlights the importance of utilizing multivoxel acquisition.
The diagnostic value of glucose or fructose loading provocative clinical tests is poor. Other exercise bicycle and trademill decreased exercise tolerance, lactate production and slow lactate clearance tests indicate mitochondrial myopathy [136]. The venous oxygen saturation in skeletal muscle results in poor extraction of oxygen from blood in mitochondrial disease [137,138]. Invasive skin biopsy investigations in many children OXPHOS defects in skeletal muscle might show change in respiratory chain enzyme activities in cultured skin fibroblasts [139]. Detectable respiratory chain enzyme defects were reported in liver [140] or cardiac muscle [141] and skeletal muscle activities. Skeletal muscle biopsy analysis is used for mitochondrial histochemical, immunohistochemical, and even ultrastructural study to evaluate for morphologic evidence of primary OXPHOS disease by respiratory chain enzyme activity analysis, integrated OXPHOS capacity, and extraction of DNA for genetic testing. The mitochondrial proliferation or regged-red fiber deposits by Trichrome Gomori stain within myofibers in sarcolemma space suggests mitochondrial atropy OXPHOS disorder. The mitochondrial enzyme activities of NADH dehydrogenase, succinate dehydrogenase (SDH), and cytochrome c oxidase histochemistry staining indicated complex II disease encoded by nuclear genes and subsarcolemmal mitochondrial accumulation [141]. Change in normal brown fibers to COXdeficient blue fibers evaluated the complex IV mitochondrial and nuclear genomes [142]. However, RRF are rarely seen in childhood but frequent sarcolemma accumulation as mild manifestation of mitochondrial proliferation in pediatric muscle biopsy [143].

Moreover, age-related muscular dystrophies, myotonic dystrophy, inflammatory myopathies, glycogenoses, and congenital myopathies also show COX-deficient non-specific muscle OXPHOS disorders including neurogenic atrophy, internal nuclei, abnormal variation in fiber size, and accumulations of glycogen or lipid [144,145]. Staining for glycogen and lipid is the gold standard to evaluate glycogen or lipid storage disorders. Rhabdomyolysis and dystrophic changes are rare in mitochondrial OXPHOS disorders. The presence of RRF with strong subsarcolemmal SDH activity and low COX activity are due to mtDNA deletions (i.e., Kearns- Sayre Syndrome or progressive external ophthalmoplegia (PEO)) or tRNA mutations (i.e., myoclonic epilepsy and ragged red fibers (MERRF)) indicating impaired mitochondrial protein synthesis [143]. The reason for COX deficiency is speculated when wildtype mtDNA levels fall below the threshold necessary for COX protein subunit expression. As a result, only few COX-deficient fibers or none may be present while high percentages of mtDNA mutation evenly distributed mutant and wild-type mtDNA throughout the fiber in MELAS due to an A3243G tRNALeu gene mutation in which RRF are often COX-positive. An increase in vascular smooth muscle SDH activity frequently is also seen in MELAS [146]. The mosaic and segmental pattern of COX activity indicates heteroplasmic mtDNA disorder while decreased COX activity throughout the length of the muscle fibers sparing spindles indicates the mutation in a nuclear gene encoding of SURF1 in homoplasmic tRNA mutations of fatal and benign forms of infantile COX-deficient myopathy [147]. Mutations in the mtDNA protein-encoding genes are occasionally associated with RRF but COX-positive fibers in cytochrome b, COX IIII, and some mutations in complex I (ND1-6, ND4L) genes.
In contrast, mutations associated with NARP/MILS (T8993G/C mutation in the ATPase 6 mtDNA gene) and ND mutations indicate LHON specific changes on muscle biopsy. Mitochondrial myopathy is OXPHOS disorder with absent nDNA mutations in nuclearencoded complex I and II subunits. However, CoQ10 deficiency, benign infantile COX-deficient myopathy are exceptions [148]. Electron microscopy ultrastructural changes as increased mitochondrial number and size distorted or absent cristae, and osmophilic or paracrystalline inclusions are well documented in patients with mitochondrial myopathic and neuropathic diseases [149]. Ultrastructural changes of mitochondria have been described in muscle biopsies of patients with no RRF or COX deficient fibers [144].
Biochemical analysis of skeletal muscle distinguishes classical syndromic presentations (i.e., Kearns-Sayre syndrome, Pearson syndrome, Leigh syndrome, MELAS, MERRF, or LHON). Over 1,000 genes are involved in the mitochondrial function [150]. For OXPHOS biochemical investigation, spectrophotometric assays of enzyme activity in isolated in vitro tissue mitochondria or cultured cells predict the diagnosis of an OXPHOS disorder [151]. Spectrophotometric data gives valuable information of respiratory complexes following either detergent- or freeze-thaw disruption of the inner mitochondrial membrane [152,153]. However, in vitro conditions do not correspond to the physiological, cytosolic environment of mitochondria. Spectrophotometric assays of the different electron transport chain (ETC) enzyme complexes measure the complex I (NADH-ubiquinone oxidoreductase), complex II (succinate-ubiquinone oxidoreductase), complex III (decylubiquinone-cytochrome c reductase) or complex IV (cytochrome c oxidase), or they can be studied together as complex I+III (NADH-cytochrome c reductase) or complex II+III (succinatecytochrome c reductase) [154]. Direct assays of ATP synthase (complex V) hydrolytic activity are available by spectrophotometric analysis. Citrate synthase enzyme distinguishes primary electron transport chain mitochondrial OXPHOS defects from secondary deficiencies [147,153,155]. However, these assays use artificial substrates in disrupted mitochondrial preparations different from intact cellular mitochondrial respiration in a stereochemical environment of multi-enzyme complexes and supercomplexes. Isolated one complex defect may point out a mutation of either an mtDNA-encoded or nDNA encoded subunit or assemblyfactor of that complex (i.e., SURF1 mutation in COX deficiency). Partial enzyme deficiencies involving complexes I, III and IV may not distinguish patients with a generalized translation defect due to either mtDNA deletions or tRNA mutations [156,157]. Complex I activity is decreased in patients with MERRF, MELAS, COX deficiency and mitochondrial polymerase deficiency [156]. The combined measurements of I+III and II+III predict treatable coenzyme Q10 deficiency disorders or CoQ10 binding/electron exchange disorders by simple coenzyme Q10 quantification in muscle [158,159]. Earlier, polarographic oxygen consumption by Clark electrode in various substrate mixture (e.g., malate + pyruvate or malate + glutamate to donate NADH, succinate to donate FADH2, or TMPD+ ascorbate to donate electrons directly to cytochrome C) indicated mitochondrial respiratory burst or oxidation rates [155].
In complex I deficiency patients, impaired respiration with NADH–producing substrates was seen while normal oxidation rates with other substrates. Patients with complex II deficiency showed reduced oxygen consumption with FADH2 producing substrates [155]. Low respiratory rates with both NADHand FADH2-producing substrates but normal oxidation of durohydroquinone and TMPD+ascorbate suggested deficiency of coenzyme Q10 otherwise with low oxidation of durohydroquinone suggested a deficiency of complex III. Diminished oxidation rates in the presence of all substrates tested for deficiency of complex IV. Polarographic studies may indicate a possible pyruvate dehydrogenase complex (PDHC) deficiency by measuring a reduced oxidation of pyruvate in the presence of normal oxidation of glutamate [160].
Respiratory chain function can be measured by isotopic 14CO2, and ATP produced from radioactively labeled substrates (e.g., [1-14C] pyruvate, [U-14C] malate and [1,4-14C] succinate) in the absence or presence of inhibitors. Citrate synthase activity is a biomarker of mitochondrial function. Various ratios can be determined to pinpoint the deficiency to a single respiratory chain complex [161]. Muscle mitochondria in muscle fibers placed in saponin housed in polarographic chamber can quantify oxygen consumption rates in presence of several substrates [162] Fresh muscle tissue is ideal for measuring oxygen consumption with polarography, substrate oxidation, or ATP production rates. Alternatively, high resolution respirometry of muscle fibers to measure oxygen consumption in isolated mitochondria or a fresh, needle biopsy of muscle or liver from 2 milligrams to evaluate all complexes’ activities in OXPHOS system [163,164]. The isolated mitochondria under in vivo like conditions identify mitochondrial defects by polarographic ETC assays such as identification of coenzyme Q10 synthetic defects [165,166], abnormal PDHC deficiency, tricarboxylic acid cycle enzymes, beta-oxidation, coenzymes, complex V, and transmembrane carriers [51], deficiency of pyruvate and phosphate transport carriers in fresh muscle mitochondria [167,168]. Coenzyme Q10 is an electron carrier to shuttle electrons to complex III from I, II, electron transfer factor (ETF), or other electron donors. It recycles tocopherol and ascorbate and has proxidant functions [169]. Impaired coenzyme Q10 dependent respiratory chain activity suggests primary coenzyme Q10 deficiency manifested as four phenotypes:
i. An encephalomyopathy with exercise intolerance, myopathy, myoglobinuria, seizures, and ataxia.
ii. Severe infantile encephalopathy with renal tubulopathy.
iii. Myopathic form with exercise intolerance, myopathy, and rhabdomyolysis.
iv. Encephalopathy with ataxia, seizures, and basal ganglia disease [170,171].
The impaired coenzyme Q10-dependent respiratory chain activity and plasma Q10 levels may indicate secondary coenzyme Q10 deficiency in muscles of Kearns-Sayre syndrome patients [172,173]. The concentration of coenzyme Q10 in heart muscle is highest at 114 ug/gram [174]. The coenzyme Q10 is lipoprotein bound reflects the cholesterol normative values established as a function of age [175]. Skeletal muscle coenzyme Q10 concentrations are two-fold high in healthy children [176].
Defects in mitochondrial oxidative phosphorylation found in both hypertrophic and dilated cardiomyopathy [177,178]. Dilated cardiomyopathy reflects pump failure arising from hypertrophic cardiomyopathies. Chamber dilation increases wall tension, cardiac output and blood pressure through equilibrium of Laplace (pressure-volume) and Frank-Starling (wall tension force) principles. Multisystemic dysfunction with isolated cardiomyopathy [155,179] arise due to mitochondrial dysfunction primary or secondary to a failing heart [180,181] in cardiac aging and cardiac disease [182,183]. Therefore, cardiac muscle biopsy may evaluate the respiratory chain OXPHOS defects secondary to primary mechanical or coronary vascular insufficiency [178]. In a severely ill neonate, infant or young child, direct gene testing by mutations in SCO2, COX-10, and COX-15 can give rise to cardiac hypertrophy.
Liver dysfunction is clinically hepatic or hepatocerebral mitochondrial disease. It is evaluated by blood DNA testing for POLG, dGUOK, or MPV17, SCO1 and SUCLA2 mutations indicating ketoacidosis (SCO1) or myopathy (SCO1 and SUCLA2) (see Table 1). More extensive evaluation is indicated by simultaneous biopsy and DNA studies at earliest. Needle biopsy yields sufficient tissue for histology, electron microscopy, and miniaturized respiratory chain polarography and biochemistry to confirm abnormal liver histologies of mitochondrial disease [179,180]; COX deficiency in respiratory chain biochemical testing; multiple ETC deficiencies with mtDNA depletion associated with POLG, dGUOK, MPV17, or SUCLA2 mutations; quantitative Southern analysis or real-time quantitative PCR to detect mtDNA depletion; normal mtDNA content on liver biopsies of POLG disease and Alpers syndrome [181] as routine liver biopsy tests.

A skin biopsy is very simple office procedure done with local anesthesia using a disposable 3- or 4-mm punch biopsy without stitches to study mitochondrial disease biochemistry [182]. Cultured skin fibroblasts can be stored indefinitely as a renewable source of DNA but not useful in mitochondria phenotyping or respiratory chain defect expression [183,184] due to altered heteroplasmy and a high tissue regeneration rate of fibroblasts [183]. In cultured skin fibroblasts, biochemical analysis includes electron transport chain enzyme analysis via spectrophotometry and radiolabeled isotope fatty acid oxidation, measurements of lactate and pyruvate [184], and ATP production and consumption in fibroblasts [185]. Fibroblasts are best for protein studies by western blot and blue native polyacrylamide gel electrophoresis (BN-PAGE) analysis [186]. BN-PAGE analyzes multisubunit mitochondrial electron transport protein complexes to identify complex abnormalities SURF1 defects [187] or tissue-specific mitochondrial translation defects [188]. The 2-dimensional BNPAGE/ SDS-PAGE was performed in combination with Western analysis using antibodies directed proteins [189,190].
Mitochondrial DNA Analysis
Mitochondrial DNA analysis is done for mitochondrial DNA mutation types. The types of mitochondrial DNA abnormalities include single basepair point mutations, several basepair insertions or deletions, large scale deletions on the order of 100s to 1000s of basepairs, or relative depletion of total mitochondrial DNA content. Routine PCR with restriction fragment length polymorphism (RFLP) analysis screens specific mutations with allele specific oligonucleotide (ASO) analysis to screen for multiple known mutations simultaneously. Other methods are Single-strand conformation polymorphism (SSCP) [191], heteroduplex screening assays (such as temporal temperature gradient gel electrophoresis (TTGE) [192-194], temperature gradient gel electrophoresis (TGGE) [195], denaturant gradient gel electrophoresis (DGGE) [196,197], denaturing high performance liquid chromatography (dHPLC) [198], and DNA sequencing. DNA sequencing is the gold standard for mutation detection in nuclear genes. For mtDNA sequencing, interference by nuclear pseudogenes exclusion is done by use of rho- cells for new PCR-based assays. Point mutations or deletions may not be present in all mtDNA genomes within a cell or tissue, but rather be heteroplasmic with wildtype to varying degrees. Quantification of mutant heteroplasmy is performed by densitometer scanning utilizing RFLP or realtime ARMS qPCR [199]. Southern blot analysis is the method most often used for the detection of largescale mtDNA deletions and duplications, although these can be detected by long range polymerase chain reaction (PCR) analysis. Finally, depletion testing utilizes either Southern densitometry scanning or real time qPCR to quantify mtDNA copy number in reference to a nuclear gene [199].
Nuclear DNA Analysis
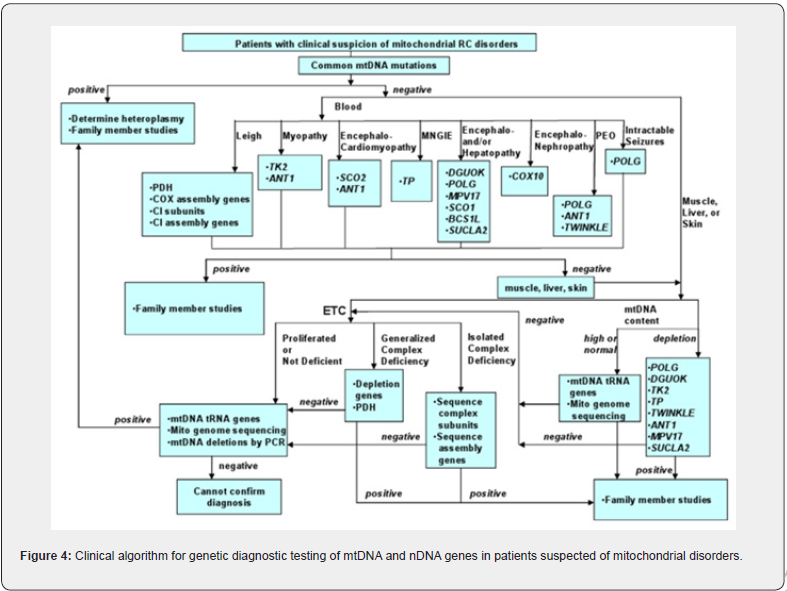
Nuclear DNA Analysis measures 75%-90% pediatric primary mitochondrial disease resulting from nDNA mutations. (Figure 2)
Complex I structural subunits [151], B17.2L & NDUFAF1 [151], SDHA [152], SDHC [153], SDHD [153],
BCS1L [154,155], SURF1 [156], SCO1 [157], SCO2 [158], COX10 [159], COX15 [160,161], ETHE1 [162], LRPPRC (LSFC) [138], ATP12 [163],
PDSS1 [164], PDSS2 [165], COQ2 [164], APTX [166], ADCK3 [167], ETFDH [168], TSFM [169], TUFM [170], EGF1 [170], MRPS16 [171],
PUS1 [172], DARS2 [173], PDHA1 [14], POLG1 [107], TK2 [174,175], DGUOK [176], MPV17 [137], SUCLA2 [177], SUCLG1 [177], RRM2B
[178], TWINKLE [179], ANT1 [180], ABC7 [181], FXN [182], OPA1 [183], DDP1 [184], SPG7 [185,186], TAZ [187].
bSensorineural hearing loss localized to the central brainstem or peripheral cochlear in pediatric and adult patients, can be seen in virtually any mitochondrial disease.
A sample algorithm of DNA-based analyses in mitochondrial disease is shown in Figure 2. Specific nuclear genes in mitochondrial disease in each patient show a detailed compilation of biochemical and clinical findings associated with pathogenic mutations in nuclear DNA genes currently implicated in primary mitochondrial diseases.
Regular clinical signs and symptoms in mitochondrial disorders are developmental and degenerative. Alpers or Kearns-Sayre syndrome presents the full clinical spectrum of mitochondrial disease. Establishing a genetic diagnosis permits accurate etiologic, prognostic, recurrence risk, and management counseling. In the future, it is hoped that this will also permit accurate subgrouping among the heterogeneous category of mitochondrial disease patients to develop disease-focused interventions. DNA-based new emerging Techniques are mtDNAM. Sequencing Microarray such as Affymetrix oligonucleotide resequencing microarray (MitoChip 2.0) to yield sequence data on the complete coding region and 15,452 of the 15,569 bp mtDNA [200] without amplifying pseudogenes by first amplifying mtDNA in 3 long overlapping fragments do not present in the nDNA. It detects heteroplasmy down to about 2-5% [201]. ESI-TOF Mass Spectrometry screens the PCR-amplified fragments of mtDNA by electrospray ionization time-of-flight mass spectrometry (ESI-TOF MS), Fourier transform ion cyclotron resonance mass spectrometry (FTICR-MS) [202] and ion-pair reverse phase HPLC time-of-flight mass spectrometry (ICEMS) [203]. ESI-TOF-MS and ICEMS are economical and well-suited for rapid sequence screening and the quantitation of heteroplasmy in the range of 1-2%.
Protein-based Immunocytochemistry is based on monoclonal antibodies for diagnosis of selected OXPHOS defects [204] using the expression of the mitochondrial protein defect in cultured skin fibroblast pyruvate dehydrogenase E1α defects [190]. However, background autofluorescence complicates the use of immunofluorescent probes in muscle biopsies. Selected use of OXPHOS antibodies in Western immunoblot analysis [204] and immunocapture of assembled complexes [205] have proven powerful analytical techniques. Proteomics and Systems Biology in purified mitochondria from different tissues combined with gel and chromatography-enhanced mass spectrometry methods, offer tissue-specific mitochondrial proteome characterization [206] and examination of tissue-specific heterogeneities [207]. A powerful bioinformatics tool “Maestro” identified 1,451 mitochondrial proteins out of 33,860 proteins in the Ensembl human protein data set [208]. Application of “Maestro” identified MPV17 as a cause of mtDNA depletion syndromes [209] and LRPPRC as the cause of Leigh syndrome [210].
31P and 13C NMR Spectroscopy measures the dynamic capacity of mitochondria during changing physiologic conditions in real-time determines. In vivo 31Phosphorus magnetic resonance spectroscopy (31P-MRS) monitors baseline, exercise, and recovery measurements of ATP and phosphocreatine to reveal very subtle derangements in mitochondrial function [211]. 13C-MRS is another emerging technology [212]. Optics, Biophotonics, and Nanotechnologies are emerging. Bio cavity Laser Spectroscopy uses the bio cavity laser, a semiconductor nanolaser about the size of a dime. Cells or mitochondria that flow through the bio cavity laser trigger the emission of photons of laser light. Measurement of the red shift of this emitted laser light provides information of the biophysical composition of healthy and diseased mitochondria. The bio cavity laser can measure the overall biophysical state of the cell and mitochondrial dysfunction in cancer [213] and primary OXPHOS disease [214].
Nanomedicine—Mitochondria have the unique electrophysical property of concentrating or attracting amphipathic, chargedelocalized, cationic molecules and nanoparticles. This allows for the rational design of a variety of cargo-delivery systems. Indeed, mitochondriotropic tags incorporated into paclitaxelbearing nanoliposomes inhibited tumor cell growth in a mouse model, whereas free paclitaxel had no effect [215]. The selective delivery of DNA to mitochondria in cell culture is state of art [216]. Mitochondria Light Interactions might stimulate incipient mitochondria cells [217]. Traces of important light interactions of mitochondria with specific wavelengths in the red and near infrared ranges (ca 600-1200 nm) are known to stimulate expansion of cardiac stem cells in culture [218] and accelerated healing of chemotherapy-associated mucositis [219]. Low level laser treatment using an 808 nm light source over mitochondria has shown promise in a blinded, prospective clinical trial in acute stroke patients [220].
In nutshell, an in-depth evaluation of mitochondrial disease is possible and indeed warranted to obtain a clear diagnosis for individual patients. This is necessary to minimize the cost of duplicate investigations, correct misinformation to families and health care providers, and provide accurate prognostic and recurrence risk counseling. Recognized mitochondrial diagnostic centers working in close collaboration with referring clinicians can successfully navigate the complexities of the diagnostic evaluation and work towards the development of quality assessment programs in the laboratory-based investigations of mitochondrial disease.
Conclusion
Mitochondria are unique energy generating and storage cellular organelles. Mitochondrial density varies in hundreds to thousands per cell. Every mitochondrion house five to ten copies of mitochondrial DNA to encode almost 13-16 bio signal mitochondrial proteins as subunits of the mitochondrial respiratory chain. Mitochondrial genetics evolved over years to recognize several nanoplatforms to confirm mtDNA-related disorders, mostly inherited maternal diseases. Mitochondria are the primary source of redox ROS radicals and ATP molecules. These bio signals control many programmed cell-death events or apoptosis. Mitochondria plays a major role in ATP synthesis, regulation of cellular oxidative state, and Ca2+ homeostasis to control cell activation bio signaling. In the case of mitochondrial dysfunctions, several clinically heterogeneous diseases are developed in a single or multiple organs. The disturbances in mitochondrial metabolism and oxidant formation alter the biology of aging and age-associated metabolic diseases, such as diabetes, cardiovascular diseases, and multiple disorders of the central and peripheral nervous system. Now, mitochondrial oxidative stress nanoplatforms may facilitate the early detection of ROS towards prevention and/or decreased mitochondrial dysfunction to restrict the generation of ROS, and mitochondrial abnormalities. Such monitoring may have benefits in lowering the disease symptoms and increasing health and lifespan of humans.
Acknowledgement
This work was supported by AMET University Innovations and Solutions Inc Global funds.
References
- Frederick RL, McCaffery JM, Cunningham KW, Okamoto K, Shaw JM (2004) Yeast Miro GTPase, Gem1p, Regulates Mitochondrial Morphology via a Novel Pathway. J Cell Biol 167(1): 87-98.
- Werner E, Werb Z (2002) Integrins engage mitochondrial function for signal transduction by a mechanism dependent on Rho GTPases. J Cell Biol 158(2): 357-368.
- Rice SE, Gelfand VI (2006) Paradigm lost: milton connects kinesin heavy chain to miro on mitochondria. J Cell Biol 173: 459-461.
- Saotome M, Safiulina D, Szabadkai G, Das S, Fransson A, et al. (2008) Bidirectional Ca2+- dependent control of mitochondrial dynamics by the Miro GTPase. Proc Natl Acad Sci 105(52): 20728-20733.
- Boveris A, Oshino N, Chance B (1972) The cellular production of hydrogen peroxide. Biochem J 128(3): 617-630.
- Lucas DT, Szweda LI (1998) Cardiac reperfusion injury: aging, lipid peroxidation, and mitochondrial dysfunction. Proc Natl Acad Sci 95(2): 510-514.
- Green K, Brand MD, Murphy MP (2004) Prevention of mitochondrial oxidative damage as a therapeutic strategy in diabetes. Diabetes 53(Supp 1): S110-S118.
- Lopez-Campistrous A, Hao L, Xiang W, Ton D, Semchuk P, et al. (2008) Mitochondrial dysfunction in the hypertensive rat brain: respiratory complexes exhibit assembly defects in hypertension. Hypertension 51(2): 412-419.
- Lee YI, Hwang JM, Im JH, Kim NS, Kim DG, et al. (2004) Human hepatitis B virus-X protein alters mitochondrial function and physiology in human liver cells. J Biol Chem 279(15): 15460-15471.
- Aguilera-Aguirre L, Bacsi A, Saavedra-Molina A, Kurosky A, Sur S, et al. (2009) Mitochondrial dysfunction increases allergic airway inflammation. J Immunol 183(8): 5379-5387.
- Guo J, Lemire BD (2003) The ubiquinone-binding site of the Saccharomyces cerevisiae succinate-ubiquinone oxidoreductase is a source of superoxide. J Biol Chem 278(48): 47629-47635.
- Moghaddas S, Hoppel CL, Lesnefsky EJ (2003) Aging defects at the QO site of complex III augments oxyradical production in rat heart interfibrillar mitochondria. Arch Biochem Biophys 414(1): 59-66.
- Sen T, Sen N, Jana S, Khan FH, Chatterjee U, et al. (2007) Depolarization and cardiolipin depletion in aged rat brain mitochondria: relationship with oxidative stress and electron transport chain activity. Neurochem Int 50(5): 719-725.
- Paravicini TM, Touyz RM (2008) NADPH oxidases, reactive oxygen species, and hypertension: clinical implications and therapeutic possibilities. Diabetes Care 31(Suppl 2): S170-S180.
- Vazquez-Memije ME, Capin R, Tolosa A, El-Hafidi M (2008) Analysis of age-associated changes in mitochondrial free radical generation by rat testis. Mol Cell Biochem 307(1-2): 23-30.
- Halliwell B, Gutteridge JMC (1999) Free radicals in biology and medicine, (3rd edn), Oxford University Press, Oxford 1-25.
- Costa AD, Quinlan CL, Andrukhiv A, West IC, Jaburek M, et al. (2006) The direct physiological effects of mitoK (ATP) opening on heart mitochondria. Am J Physiol Heart Circ Physiol 290(1): H406-H415.
- Ebadi M, Sharma SK (2003) Peroxynitrite and mitochondrial dysfunction in the pathogenesis of Parkinson's disease, Antioxid Redox Signal 5(3): 319-335.
- Han Z, Chen YR, Meenakshisundaram G, Zweier JL, Alevriadou BR (2007) Shear-induced reactive nitrogen species inhibit mitochondrial respiratory complex activities in cultured vascular endothelial cells. Am J Physiol Cell Physiol 292(3): C1103-C1112.
- Sidoti-de Fraisse C, Rincheval V, Risler Y, Mignotte B, Vayssiere JL (1998) TNF-alpha activates at least two apoptotic signaling cascades. Oncogene 17(13): 1639-1651.
- Kalra N, Kumar V (2004) c-Fos is a mediator of the c-myc-induced apoptotic signaling in serum-deprived hepatoma cells via the p38 mitogen-activated protein kinase pathway. J Biol Chem 279(24): 25313-25319.
- Esteve P, Embade N, Perona R, Jimenez B, del Peso L, et al. (1998) Rho-regulated signals induce apoptosis in vitro and in vivo by a p53-independent, but Bcl2 dependent pathway. Oncogene 17(14): 1855-1869.
- Lores P, Morin L, Luna R, Gacon G (1997) Enhanced apoptosis in the thymus of transgenic mice expressing constitutively activated forms of human Rac2GTPase. Oncogene 15(5): 601-605.
- Bazenet CE, Mota MA, Rubin LL (1998) The small GTP-binding protein Cdc42 is required for nerve growth factor withdrawal-induced neuronal death. Proc Natl Acad Sci USA 95(7): 3984-3989.
- Fiorentini C, Matarrese P, Straface E, Falzano L, Donelli G, et al. (1998) Rho-dependent cell spreading activated by E. coli cytotoxic necrotizing factor 1 hinders apoptosis in epithelial cells. Cell Death Differ 5(11): 921-929.
- Burridge K, Wennerberg K (2004) Rho and Rac take center stage. Cell 116(2): 167-179.
- Kowluru A, Chen HQ, Tannous M (2003) Novel roles for the rho subfamily of GTP-binding proteins in succinate-induced insulin secretion from betaTC3 cells: further evidence in support of the succinate mechanism of insulin release. Endocr Res 29(3): 363-376.
- Fransson A, Ruusala A, Aspenstrom P (2003) Atypical Rho GTPases have roles in mitochondrial homeostasis and apoptosis. J Biol Chem 278(8): 6495-6502.
- Guo X, Macleod GT, Wellington A, Hu F, Panchumarthi S, et al. (2005) The GTPase dMiro is required for axonal transport of mitochondria to Drosophila synapses. Neuron 47(3): 379-393.
- Cai Q, Sheng ZH (2009) Moving or stopping mitochondria: Miro as a traffic cop by sensing calcium. Neuron 61(4): 493-496.
- Del Re DP, Miyamoto S, Brown JH (2007) RhoA/Rho kinase up-regulate Bax to activate a mitochondrial death pathway and induce cardiomyocyte apoptosis. J Biol Chem 282(11): 8069-8078.
- Yanazume T, Hasegawa K, Wada H, Morimoto T, Abe M, et al. (2002) Rho/ROCK pathway contributes to the activation of extracellular signal-regulated kinase/GATA-4 during myocardial cell hypertrophy. J Biol Chem 277(10): 8618-8625.
- Zhang Y, Gu, X, Yuan X (2007) Phenylalanine activates the mitochondria-mediated apoptosis through the RhoA/Rho-associated kinase pathway in cortical neurons. Eur J Neurosci 25(5): 1341-1348.
- Koyanagi M, Takahashi J, Arakawa Y, Doi D, Fukuda H, et al. (2008) Inhibition of the Rho/ROCK pathway reduces apoptosis during transplantation of embryonic stem cell-derived neural precursors. J Neurosci Res 86(2): 270-280.
- Chuang TH, Hahn KM, Lee JD, Danley DE, Bokoch, GM (1997) The small GTPase Cdc42 initiates an apoptotic signaling pathway in Jurkat T lymphocytes. Mol Biol Cell 8(9): 1687-1698.
- Embade N, Valeron PF, Aznar S, Lopez-Collazo E, Lacal JC (2000) Apoptosis induced by Rac GTPase correlates with induction of FasL and ceramides production. Mol Biol Cell 11(12): 4347-4358.
- Meske V, Albert F, Richter D, Schwarze J, Ohm TG (2003) Blockade of HMG-CoA reductase activity causes changes in microtubule-stabilizing protein tau via suppression of geranylgeranylpyrophosphate formation: implications for Alzheimer's disease. Eur J Neurosci 17(1): 93-102.
- Le SS, Loucks FA, Udo H, Richardson-Burns S, Phelps RA, et al. (2005) Inhibition of Rac GTPase triggers a c-Jun- and Bim-dependent mitochondrial apoptotic cascade in cerebellar granule neurons. J Neurochem 94(4): 1025-1039.
- Gulbins E, Coggeshall KM, Brenner B, Schlottmann K, Linderkamp O, et al. (1996) Fas-induced apoptosis is mediated by activation of a Ras and Rac protein-regulated signaling pathway. J Biol Chem 271(42): 26389-26394.
- Gallo KA, Johnson GL (2002) Mixed-lineage kinase control of JNK and p38 MAPK pathways. Nat Rev Mol Cell Biol 3(9): 663-672.
- Ozaki M, Deshpande SS, Angkeow P, Suzuki S, Irani K (2000) Rac1 regulates stress-induced, redox-dependent heat shock factor activation. J Biol Chem 275(45): 35377-35383.
- Freeman JL, Abo A, Lambeth JD (1996) Rac "insert region" is a novel effector region that is implicated in the activation of NADPH oxidase, but not PAK65. J Biol Chem 271(33): 19794-19801.
- Joneson T, Bar-Sagi D (1998) A Rac1 effector site controlling mitogenesis through superoxide production. J Biol Chem 273(29): 17991-17994.
- Kajimoto H, Hashimoto K, Bonnet SN, Haromy A, Harry G, et al. (2007) Oxygen activates the Rho/Rho-kinase pathway and induces RhoB and ROCK-1 expression in human and rabbit ductus arteriosus by increasing mitochondria-derived reactive oxygen species: a newly recognized mechanism for sustaining ductal constriction. Circulation 115(13): 1777-1788.
- Nimnual AS, Taylor LJ, Bar-Sagi, D (2003) Redox-dependent downregulation of Rho by Rac. Nat Cell Biol 5(3): 236-241.
- Jin L, Ying Z, Webb RC (2004) Activation of Rho/Rho kinase signaling pathway by reactive oxygen species in rat aorta. Am J Physiol Heart Circ Physiol 287(4): H1495-1500.
- Sofic E, Paulus W, Jellinger K, Riederer P, Youdim MB (1991) Selective increase of iron in substantia nigra zona compacta of parkinsonian brains. J Neurochem 56(3): 978-982.
- Faucheux BA, Nillesse N, Damier P, Spik G, Mouatt-Prigent A, et al. (1995) Expression of lactoferrin receptors is increased in the mesencephalon of patients with Parkinson disease. Proc Natl Acad Sci U S A 92(21): 9603-9607.
- Toyokuni S (1996) Iron-induced carcinogenesis: the role of redox regulation. Free Radic Biol Med 20(4): 553-566.
- Carmine TC, Evans P, Bruchelt G, Evans R, Handgretinger R, et al. (1995) Presence of iron catalytic for free radical reactions in patients undergoing chemotherapy: implications for therapeutic management. Cancer Lett 94(2): 219-226.
- Speechley M, Barton JC, Passmore L, Harrison H, Reboussin DM, et al. (2009) Potential Nonresponse Bias in a Clinical Examination After Initial Screening Using Iron Phenotyping and HFE Genotyping in the Hemochromatosis and Iron Overload Screening Study. Genet Test Mol Biomarkers 13(6): 721-728.
- Fernandez-Real JM, Vendrell J, Baiget M, Gimferrer E, Ricart W (1999) C282Y and H63D mutations of the hemochromatosis candidate gene in type 2 diabetes. Diabetes Care 22(3): 525-526.
- Gao X, Campian JL, Qian M, Sun XF, Eaton JW (2009) Mitochondrial DNA damage in iron overload. J Biol Chem 284(8): 4767-4775.
- Gordon N (2000) Friedreich's ataxia and iron metabolism. Brain Dev 22(8): 465-468.
- Graf E, Mahoney JR, Bryant RG, Eaton JW (1984) Iron-catalyzed hydroxyl radical formation. Stringent requirement for free iron coordination site. J Biol Chem 259(6): 3620-3624.
- Luo Y, Han Z, Chin SM, Linn S (1994) Three chemically distinct types of oxidants formed by iron-mediated Fenton reactions in the presence of DNA. Proc Natl Acad Sci U S A 91(26): 12438-12442.
- Kwok EKD (2006) Molecular biology of metal homeostasis and detoxification. From microbes to man., Springer, Goteborg, Sweden.
- Craig EA, Voisine C, Schilke B (1999) Mitochondrial iron metabolism in the yeast Saccharomyces cerevisiae. Biol Chem 380(10): 1167-1173.
- Radisky DC, Babcock MC, Kaplan J (1999) The yeast frataxin homologue mediates mitochondrial iron efflux. Evidence for a mitochondrial iron cycle. J Biol Chem 274(8): 4497-4499.
- Rotig A, De Lonlay P, Chretien D, Foury F, Koenig M, et al. (1997) Aconitase and mitochondrial iron-sulphur protein deficiency in Friedreich ataxia. Nat Genet 17(2): 215-217.
- Foury F, Cazzalini O (1997) Deletion of the yeast homologue of the human gene associated with Friedreich's ataxia elicits iron accumulation in mitochondria. FEBS Lett 411(2-3): 373-337.
- Wilson RB, Roof DM (1997) Respiratory deficiency due to loss of mitochondrial DNA in yeast lacking the frataxin homologue. Nat Genet 16(4): 352-357.
- Cortés-Rojo C, Calderón-Cortés E, Clemente-Guerrero M, Estrada-Villagómez M, Manzo-Avalos S, et al. (2009) Elucidation of the effects of lipoperoxidation on the mitochondrial electron transport chain using yeast mitochondria with manipulated fatty acid content. J Bioenerg Biomembr 41(1): 15-28.
- Augustin W, Wiswedel I, Noack H, Reinheckel T, Reichelt O (1997) Role of endogenous and exogenous antioxidants in the defense against functional damage and lipid peroxidation in rat liver mitochondria. Mol Cell Biochem 174(1-2): 199-205.
- Reinheckel T, Wiswedel I, Noack H, Augustin W (1995) Electrophoretic evidence for the impairment of complexes of the respiratory chain during iron/ascorbate induced peroxidation in isolated rat liver mitochondria. Biochim Biophys Acta 1239(1): 45-50.
- Rafique R, Schapira AH, Cooper JM (2001) Sensitivity of respiratory chain activities to lipid peroxidation: effect of vitamin E deficiency. Biochem J 357(Pt 3): 887-892.
- Forsmark-Andree P, Lee CP, Dallner G, Ernster L (1997) Lipid peroxidation and changes in the ubiquinone content and the respiratory chain enzymes of submitochondrial particles. Free Radic Biol Med 22(3): 391-400.
- Cortés-Rojo C, Calderón-Cortés E, Clemente-Guerrero M, Manzo-Avalos S, Uribe S, et al. (2007) Electron transport chain of Saccharomyces cerevisiae mitochondria is inhibited by H2O2 at succinate-cytochrome c oxidoreductase level without lipid peroxidation involvement. Free Radic Res 41(11): 1212-1223.
- Paradies G, Petrosillo G, Pistolese M, Ruggiero FM (2002) Reactive oxygen species affect mitochondrial electron transport complex I activity through oxidative cardiolipin damage. Gene 286(1): 135-141.
- Petrosillo G, Ruggiero FM, Di Venosa N, Paradies G (2003) Decreased complex III activity in mitochondria isolated from rat heart subjected to ischemia and reperfusion: role of reactive oxygen species and cardiolipin. FASEB J 17(6): 714-716.
- Paradies G, Petrosillo G, Pistolese M, Ruggiero FM (2001) Reactive oxygen species generated by the mitochondrial respiratory chain affect the complex III activity via cardiolipin peroxidation in beef-heart submitochondrial particles. Mitochondrion 1(2): 151-159.
- Paradies G, Petrosillo G, Pistolese M, Di Venosa N, Federici A, et al. (2004) Decrease in mitochondrial complex I activity in ischemic/reperfused rat heart: involvement of reactive oxygen species and cardiolipin. Circ Res 94(1): 53-59.
- Chicco AJ, Sparagna GC (2007) Role of cardiolipin alterations in mitochondrial dysfunction and disease. Am J Physiol Cell Physiol 292(1): C33-44.
- Petrosillo G, Matera M, Moro N, Ruggiero FM, Paradies G (2009) Mitochondrial complex I dysfunction in rat heart with aging: critical role of reactive oxygen species and cardiolipin. Free Radic Biol Med 46(1): 88-94.
- Lesnefsky EJ, Hoppel CL (2008) Cardiolipin as an oxidative target in cardiac mitochondria in the aged rat. Biochim Biophys Acta 1777(7-8): 1020-10207.
- Paradies G, Ruggiero FM, Petrosillo G, Quagliariello E (1997) Age-dependent decline in the cytochrome c oxidase activity in rat heart mitochondria: role of cardiolipin. FEBS Lett 406(1-2): 136-138.
- Paradies G, Petrosillo G, Ruggiero FM (1997) Cardiolipin-dependent decrease of cytochrome c oxidase activity in heart mitochondria from hypothyroid rats. Biochim Biophys Acta 1319(1): 5-8.
- Paradies G, Ruggiero FM, Petrosillo G, Quagliariello E (1998) Peroxidative damage to cardiac mitochondria: cytochrome oxidase and cardiolipin alterations. FEBS Lett 424(3): 155-158.
- Desai ID, Tappel AL (1963) Damage to Proteins by Peroxidized Lipids. J Lipid Res 4: 204-247.
- Burdon RH, Rice-Evans C (1994) Free radical damage and its control. Elsevier Amsterdam New York, USA.
- Cardoso SM, Pereira C, Oliveira R (1999) Mitochondrial function is differentially affected upon oxidative stress. Free Radic Biol Med 26(1-2): 3-13.
- Trumper L, Hoffmann B, Wiswedel I, Augustin W (1988) Impairment of the respiratory chain in the b-c1 region as early functional event during Fe2+/ascorbate induced peroxidation in rat liver mitochondria. Biomed Biochim Acta 47(12): 933-939.
- Masini A, Ceccarelli D, Giovannini F, Montosi G, Garuti C, et al. (2000) Iron-induced oxidant stress leads to irreversible mitochondrial dysfunctions and fibrosis in the liver of chronic iron-dosed gerbils. The effect of silybin. J Bioenerg Biomembr 32(2): 175-182.
- Tuller G, Nemec T, Hrastnik C, Daum G (1999) Lipid composition of subcellular membranes of an FY1679-derived haploid yeast wild-type strain grown on different carbon sources. Yeast 15(4): 1555-1564.
- Trumpower BL (1990) Cytochrome bc1 complexes of microorganisms. Microbiol Rev 54(2): 101-129.
- Tsai AL, Palmer G (1986) The role of phospholipids in the binding of antimycin to yeast Complex III. Biochim Biophys Acta 852(1): 100-105.
- Lange C, Nett JH, Trumpower BL, Hunte C (2001) Specific roles of protein-phospholipid interactions in the yeast cytochrome bc1 complex structure. Embo J 20(23): 6591-6600.
- Yang S, Ma HW, Yu L, Yu CA (2008) On the mechanism of quinol oxidation at the QP site in the cytochrome bc1 complex: studied using mutants lacking cytochrome bL or bH. J Biol Chem 283(42): 28767-28776.
- Winterbourn CC (1993) Superoxide as an intracellular radical sink. Free Radic Biol Med 14(1): 85-90.
- Forman HJ, Fukuto JM, Torres M (2004) Redox signaling: thiol chemistry defines which reactive oxygen and nitrogen species can act as second messengers. Am J Physiol Cell Physiol 287(2): C246-256.
- Lin TK, Hughes G, Muratovska A, Blaikie FH, Brookes PS, et al. (2002) Specific modification of mitochondrial protein thiols in response to oxidative stress: a proteomics approach. J Biol Chem 277(19): 17048-17056.
- Hurd TR, Requejo R, Filipovska A, Brown S, Prime TA, et al. (2008) Complex I within oxidatively stressed bovine heart mitochondria is glutathionylated on Cys-531 and Cys-704 of the 75-kDa subunit: potential role of CYS residues in decreasing oxidative damage. J Biol Chem 283(236): 24801-24815.
- Le-Quoc K, Le-Quoc D, Gaudemer Y (1981) Evidence for the existence of two classes of sulfhydryl groups essential for membrane-bound succinate dehydrogenase activity. Biochemistry 20(7): 1705-1710.
- Zhang Y, Marcillat O, Giulivi C, Ernster L, Davies KJ (1990) The oxidative inactivation of mitochondrial electron transport chain components and ATPase. J Biol Chem 265(27): 16330-16336.
- Vygodina TV, Pecoraro C, Mitchell D, Gennis R, Konstantinov AA (1998) Mechanism of inhibition of electron transfer by amino acid replacement K362M in a proton channel of Rhodobacter sphaeroides cytochrome c oxidase. Biochemistry 37(9): 3053-3061.
- Vygodina TV, Konstantinov AA (2007) Peroxidase activity of mitochondrial cytochrome c oxidase. Biochemistry (Mosc) 72(10): 1056-1064.
- Jha N, Jurma O, Lalli G, Liu Y, Pettus EH, et al. (2000) Glutathione depletion in PC12 results in selective inhibition of mitochondrial complex I activity. Implications for Parkinson's disease. J Biol Chem 275(34): 26096-26101.
- Shivakumar BR, Kolluri SV, Ravindranath V (1995) Glutathione and protein thiol homeostasis in brain during reperfusion after cerebral ischemia. J Pharmacol Exp Ther 274(3): 1167-1173.
- Balijepalli S, Boyd MR, Ravindranath V (1999) Inhibition of mitochondrial complex I by haloperidol: the role of thiol oxidation. Neuropharmacology 38(4): 567-577.
- Hurd TR, Prime TA, Harbour ME, Lilley KS, Murphy MP (2007) Detection of reactive oxygen species-sensitive thiol proteins by redox difference gel electrophoresis: implications for mitochondrial redox signaling. J Biol Chem 282(30): 22040-22051.
- Jacob C, Holme AL, Fry FH (2004) The sulfinic acid switch in proteins. Org Biomol Chem 2(14): 1953-1956.
- Nulton-Persson AC, Szweda LI (2001) Modulation of mitochondrial function by hydrogen peroxide. J Biol Chem 276 (26): 23357-23361.
- Haas RH, Parikh S, Falk MJ, Saneto RP, Wolf NI et al. (2008) In-Depth evaluation of suspected mitochondrial disease. The Mitochondrial Medicine Society committee on diagnosis. Mol Genet Metab 94(1): 16-37.
- Chow SL (2005) The significance of elevated CSF lactate. Arch Dis Child 90(11): 1188-1189.
- Finsterer J (2001) Cerebrospinal-fluid lactate in adult mitochondriopathy with and without encephalopathy. Acta Med Austriaca 28(5): 152-155.
- Cameron JM (2004) Deficiency of pyruvate dehydrogenase caused by novel and known mutations in the E1alpha subunit. Am J Med Genet A 131(1): 59-66.
- Maj MC, Cameron JM, Robinson BH (2006) Pyruvate dehydrogenase phosphatase deficiency: orphan disease or an under-diagnosed condition? Mol Cell Endocrinol 249(12): 1-9.
- Zschocke J, Hoffmann GF (2004) Vademecum Metabolicum 2nd Friedrichs Dorf Germany Milupa p. 32.
- Campos Y (1995) Mitochondrial DNA deletion in a patient with mitochondrial myopathy, lactic acidosis, and stroke-like episodes (MELAS) and Fanconi's syndrome. Pediatr Neurol 13(1): 69-72.
- Niaudet P (1994) Deletion of the mitochondrial DNA in a case of de Toni-Debre-Fanconi syndrome and Pearson syndrome. Pediatr Nephrol 8(2): 164-168.
- Chace DH, Kalas TA (2005) A biochemical perspective on the use of tandem mass spectrometry for newborn screening and clinical testing. Clin Biochem 38(4): 296-309.
- Barshop BA (2004) Metabolomic approaches to mitochondrial disease: correlation of urine organic acids. Mitochondrion 4(56): 521-527.
- Sperl W (2006) Deficiency of mitochondrial ATP synthase of nuclear genetic origin. Neuromuscul Disord 16(12): 821-829.
- Gibson KM (1991) Phenotypic heterogeneity in the syndromes of 3-methylglutaconic aciduria. J Pediatr 118(6): 885-890.
- Minkler PE, Hoppel CL (1993) Quantification of free carnitine, individual short- and medium-chain acylcarnitines, and total carnitine in plasma by high-performance liquid chromatography. Anal Biochem 212(2): 510-518.
- Van der Knaap MS, Jakobs C, Valk J (1996) Magnetic resonance imaging in lactic acidosis. J Inherit Metab Dis 1996;19(4): 535-547.
- Barkovich AJ (1993) Mitochondrial disorders: analysis of their clinical and imaging characteristics. AJNR Am J Neuroradiol 14(5): 1119-1137.
- Lin DD, Crawford TO, Barker PB (2003) Proton MR spectroscopy in the diagnostic evaluation of suspected mitochondrial disease. AJNR Am J Neuroradiol 24(1): 33-41.
- Dinopoulos A (2005) Brain MRI and proton MRS findings in infants and children with respiratory chain defects. Neuropediatrics 36(5): 290-301.
- Bianchi MC (2003) Proton MR spectroscopy of mitochondrial diseases: analysis of brain metabolic abnormalities and their possible diagnostic relevance. AJNR Am J Neuroradiol 24(10): 1958-1966.
- Kingsley PB, Shah TC, Woldenberg R (2006) Identification of diffuse and focal brain lesions by clinical magnetic resonance spectroscopy. NMR Biome 19(4): 435-62.
- Saneto RP (2001) MRS detection of CNS lactate peaks in primary mitochondrial cytopathies. Neurology 56 (suppl 3): A42.
- Clark JB (1998) N-acetyl aspartate: a marker for neuronal loss or mitochondrial dysfunction. Dev Neurosci 20(45): 271-276.
- Pavlakis SG (1998) Magnetic resonance spectroscopy: use in monitoring MELAS treatment. Arch Neurol 55(6): 849-852.
- De Stefano N, Matthews PM, Arnold DL (1995) Reversible decreases in N-acetylaspartate after acute brain injury. Magn Reson Med 34(5): 721-727.
- Martin E (2001) Absence of N-acetylaspartate in the human brain: impact on neurospectroscopy? Ann Neurol 49(4): 518-521.
- Tan J (1998) Lack of effect of oral choline supplement on the concentrations of choline metabolites in human brain. Magn Reson Med 39(6): 1005-1010.
- Miller BL (1996) In vivo 1H MRS choline: correlation with in vitro chemistry/histology. Life Sci 58(22): 1929-1935.
- Govindaraju V, Young K, Maudsley AA (2000) Proton NMR chemical shifts and coupling constants for brain metabolites. NMR Biomed 13(3): 129-153.
- Klunk WE (1996) Quantitative 1H and 31P MRS of PCA extracts of postmortem Alzheimer's disease brain. Neurobiol Aging 17(3): 349-357.
- Brockmann K (2002) Succinate in dystrophic white matter: a proton magnetic resonance spectroscopy finding characteristic for complex II deficiency. Ann Neurol 52(1): 38-46.
- Lin A (2005) Efficacy of proton magnetic resonance spectroscopy in neurological diagnosis and neurotherapeutic decision making. NeuroRx 2(2): 197-214.
- Haas RH, Barshop BA (1998) Diet change in the management of metabolic encephalomyopathies. Biofactors 7(3): 259-262.
- Tarnopolsky M (2004) Exercise testing as a diagnostic entity in mitochondrial myopathies. Mitochondrion 4(56): 529-542.
- Taivassalo T (2002) Venous oxygen levels during aerobic forearm exercise: An index of impaired oxidative metabolism in mitochondrial myopathy. Ann Neurol 51(1): 38-44.
- Jensen TD (2002) A forearm exercise screening test for mitochondrial myopathy. Neurology 58(10): 1533-1538.
- Thorburn DR, Chow CW, Kirby DM (2004) Respiratory chain enzyme analysis in muscle and liver. Mitochondrion 4(5-6): 363-375.
- Garcia-Cazorla A (2006) Mitochondrial respiratory chain deficiencies expressing the enzymatic deficiency in the hepatic tissue: a study of 31 patients. J Pediat 149(3): 401-405.
- Taylor RW (2003) A homoplasmic mitochondrial transfer ribonucleic acid mutation as a cause of maternally inherited hypertrophic cardiomyopathy. J Am Coll Cardiol 41(10): 1786-1796.
- Tanji K, Bonilla E (2007) Optical imaging techniques (histochemical, immunohistochemical, and in situ hybridization staining methods) to visualize mitochondria. Methods Cell Biol 80(26): 135-154.
- Scaglia F (2004) Clinical spectrum, morbidity, and mortality in 113 pediatric patients with mitochondrial disease. Pediatrics 114(4): 925-931.
- Rollins S (2001) Diagnostic yield muscle biopsy in patients with clinical evidence of mitochondrial cytopathy. Am J Clin Pathol 116(3): 326-330.
- Tulinius MH (1991) Mitochondrial encephalomyopathies in childhood. I. Biochemical and morphologic investigations. J Pediatr 119(2): 242-250.
- Hasegawa H (1991) Strongly succinate dehydrogenase-reactive blood vessels in muscles from patients with mitochondrial myopathy, encephalopathy, lactic acidosis, and stroke-like episodes. Ann Neurol 29(6): 601-605.
- Tanji K, Bonilla E (2000) Neuropathologic aspects of cytochrome C oxidase deficiency. Brain Pathol 10(3): 422-430.
- Oldfors A, Tulinius M (2003) Mitochondrial encephalomyopathies. J Neuropathol Exp Neurol 62(3): 217-227.
- Cotter D (2004) MitoProteome: mitochondrial protein sequence database and annotation system. Nucleic Acids Res 32(Database issue): D463-467.
- Thorburn DR (2004) Biochemical and molecular diagnosis of mitochondrial respiratory chain disorders. Biochim Biophys Acta 1659(23): 121-128.
- Birch-Machin MA (1994) An evaluation of the measurement of the activities of complexes I-IV in the respiratory chain of human skeletal muscle mitochondria. Biochem Med Metab Biol 51(1): 35-42.
- Chretien D (1994) Reference charts for respiratory chain activities in human tissues. Clin Chim Acta 228(1): 53-70.
- Rustin P (1994) Endomyocardial biopsies for early detection of mitochondrial disorders in hypertrophic cardiomyopathies. J Pediatr 124(2): 224-228.
- Bindoff LA (1991) Multiple defects of the mitochondrial respiratory chain in mitochondrial encephalopathy (MERRF): a clinical, biochemical and molecular study. J Neurol Sci 102(1): 17-24.
- Moraes CT (1992) The mitochondrial tRNA (Leu(UUR)) mutation in mitochondrial encephalomyopathy, lactic acidosis, and strokelike episodes (MELAS): genetic, biochemical, and morphological correlations in skeletal muscle. Am J Hum Genet 50(5): 934-949.
- Montero R (2005) Muscle coenzyme Q10 concentrations in patients with probable and definite diagnosis of respiratory chain disorders. Biofactors 25(1-4): 109-115.
- Quinzii CM, D S, Hirano M (2007) Human Coenzyme Q (10) Deficiency. Neurochem Res 32(4-5): 723-727.
- Tulinius M (2005) A family with pyruvate dehydrogenase complex deficiency due to a novel C>T substitution at nucleotide position 407 in exon 4 of the X-linked Epsilon1alpha gene. Eur J Pediatr 164(2): 99-103.
- Janssen AJ (2006) Measurement of the energy-generating capacity of human muscle mitochondria: diagnostic procedure and application to human pathology. Clin Chem 52(5): 860-871.
- Wenchich L (2003) Polarographic evaluation of mitochondrial enzymes activity in isolated mitochondria and in permeabilized human muscle cells with inherited mitochondrial defects. Physiol Res 52(6): 781-8.
- Sperl W (1997) High resolution respirometry of permeabilized skeletal muscle fibers in the diagnosis of neuromuscular disorders. Mol Cell Biochem 174(12): 71-78.
- Kuznetsov AV (2002) Evaluation of mitochondrial respiratory function in small biopsies of liver. Anal Biochem 305(2): 186-194.
- Boitier E (1998) A case of mitochondrial encephalomyopathy associated with a muscle coenzyme Q10 deficiency. J Neurol Sci 156(1): 41-46.
- Rotig A (2000) Quinone-responsive multiple respiratory-chain dysfunction due to widespread coenzyme Q10 deficiency. Lancet 356(9227): 391-395.
- Brivet M (2003) Impaired mitochondrial pyruvate importation in a patient and a fetus at risk. Mol Genet Metab 78(3): 186-192.
- Mayr JA (2007) Mitochondrial phosphate-carrier deficiency: a novel disorder of oxidative phosphorylation. Am J Hum Genet 80(3): 478-84.
- Linnane AW, Kios M, Vitetta L (2007) Coenzyme Q(10) - Its role as a prooxidant in the formation of superoxide anion/hydrogen peroxide and the regulation of the metabolome. Mitochondrion 7: S51-S61.
- Rotig A (2007) Infantile and pediatric quinone deficiency diseases. Mitochondrion 7: S112-S121.
- Quinzii C (2006) A Mutation in Para-Hydroxybenzoate-Polyprenyl Transferase (COQ2) Causes Primary Coenzyme Q10 Deficiency. Am J Hum Genet 78(2): 345-349.
- Ogasahara S (1986) Treatment of Kearns-Sayre syndrome with coenzyme Q10. Neurology 36(1): 45-53.
- Haas RH (2007) The evidence basis for coenzyme Q therapy in oxidative phosphorylation disease. Mitochondrion 7: S136-S145.
- Turunen M, Olsson J, Dallner G (2004) Metabolism and function of coenzyme Q. Biochim Biophys Acta 1660(12): 171-199.
- Duncan AJ (2005) Determination of coenzyme Q10 status in blood mononuclear cells, skeletal muscle, and plasma by HPLC with di-propoxy-coenzyme Q10 as an internal standard. Clin Chem 51(12): 2380-2382.
- Miles MV (2005) Age-related changes in plasma coenzyme Q10 concentrations and redox state in apparently healthy children and adults. Clin Chim Acta 347(1-2): 139-144.
- Pastore A (2005) Simultaneous determination of ubiquinol and ubiquinone in skeletal muscle of pediatric patients. Anal Biochem 342(2): 352-355.
- Lev D (2004) Clinical presentations of mitochondrial cardiomyopathies. Pediatr Cardiol 25(5): 443-450.
- Marin-Garcia J (1999) Mitochondrial dysfunction in skeletal muscle of children with cardiomyopathy. Pediatrics 103(2): 456-459.
- Marin-Garcia J, Goldenthal MJ, Filiano JJ (2002) Cardiomyopathy associated with neurologic disorders and mitochondrial phenotype. J Child Neurol 17(10): 759-765.
- Jarreta D, (2000) Mitochondrial function in heart muscle from patients with idiopathic dilated cardiomyopathy. Cardiovasc Res 45(4): 860-865.
- Stanley WC, Hoppel CL (2000) Mitochondrial dysfunction in heart failure: potential for therapeutic interventions? Cardiovasc Res 45(4): 805-806.
- Lesnefsky EJ (2001) Mitochondrial dysfunction in cardiac disease: ischemia-reperfusion, aging, and heart failure. J Mol Cell Cardiol 33(6): 1065-1089.
- Riva A (2005) Structural differences in two biochemically defined populations of cardiac mitochondria. Am J Physiol Heart Circ Physiol 289(2): H868-H872.
- Nguyen KV (2006) Molecular diagnosis of Alpers syndrome. J Hepatol 45(1): 108-116.
- Labarthe F (2005) Clinical, biochemical, and morphological features of hepatocerebral syndrome with mitochondrial DNA depletion due to deoxyguanosine kinase deficiency. J Hepatol 43(2): 333-341.
- Van den Heuvel LP, Smeitink JA, Rodenburg RJ (2004) Biochemical examination of fibroblasts in the diagnosis and research of oxidative phosphorylation (OXPHOS) defects. Mitochondrion 4(5-6): 395-401.
- Cameron JM (2004) Respiratory chain analysis of skin fibroblasts in mitochondrial disease. Mitochondrion 4(56): 387-394.
- Carrozzo R (2001) The T9176G mtDNA mutation severely affects ATP production and results in Leigh syndrome. Neurology 56(5): 687-690.
- McKenzie M (2007) Analysis of mitochondrial subunit assembly into respiratory chain complexes using Blue Native polyacrylamide gel electrophoresis. Anal Biochem 364(2): 128-137.
- Tiranti V (1999) Characterization of SURF-1 expression and Surf-1p function in normal and disease conditions. Hum Mol Genet 8(13): 2533-2540.
- Antonicka H (2006) The molecular basis for tissue specificity of the oxidative phosphorylation deficiencies in patients with mutations in the mitochondrial translation factor EFG1. Hum Mol Genet 15(11): 1835-1846.
- Capaldi RA (2004) Immunological approaches to the characterization and diagnosis of mitochondrial disease. Mitochondrion 4(5-6): 417-426.
- Margineantu DH (2002) Heterogeneous distribution of pyruvate dehydrogenase in the matrix of mitochondria. Mitochondrion 1(4): 327-338.
- Suomalainen A (1992) Use of single strand conformation polymorphism analysis to detect point mutations in human mitochondrial DNA. J Neurol Sci 111(2): 222-226.
- Wong LJ (2002) Comprehensive scanning of the entire mitochondrial genome for mutations. Clin Chem 48(11): 1901-1912.
- Wong LJ, Chen TJ, Tan DJ (2004) Detection of mitochondrial DNA mutations using temporal temperature gradient gel electrophoresis. Electrophoresis 25(15): 2602-2610.
- Chen TJ, Boles RG, Wong LJ (1999) Detection of mitochondrial DNA mutations by temporal temperature gradient gel electrophoresis. Clin Chem 45(8 Pt 1): 1162-1167.
- Wartell RM, Hosseini SH, Moran CP Jr (1990) Detecting base pair substitutions in DNA fragments by temperature-gradient gel electrophoresis. Nucleic Acids Res 18(9): 2699-2705.
- Sternberg D (1998) Exhaustive scanning approach to screen all the mitochondrial tRNA genes for mutations and its application to the investigation of 35 independent patients with mitochondrial disorders. Hum Mol Genet 7(1): 33-42.
- Michikawa Y (1997) Comprehensive, rapid and sensitive detection of sequence variants of human mitochondrial tRNA genes. Nucleic Acids Res 25(12): 2455-2463.
- Van Den Bosch BJ (2000) Mutation analysis of the entire mitochondrial genome using denaturing high performance liquid chromatography. Nucleic Acids Res 28(20): E89.
- Bai RK, Wong LJ (2004) Detection and quantification of heteroblastic mutant mitochondrial DNA by real-time amplification refractory mutation system quantitative PCR analysis: a single-step approach. Clin Chem 50(6): 996-1001.
- Zhou S (2006) An oligonucleotide microarray for high-throughput sequencing of the mitochondrial genome. J Mol Diagn 8(4): 476-482.
- Coon KD, Jon Valla, Szabolics Szelinger, Lonnie E Schneider, Tracy L Niedzielko, et al. (2006) Quantitation of heteroplasmy of mtDNA sequence variants identified in a population of AD patients and controls by array-based resequencing. Mitochondrion 6(4): 194-210.
- Jiang Y, Thomas A Hall, Steven A Hofstadler, Robert K Naviaux (2007) Mitochondrial DNA mutation detection by electrospray mass spectrometry. Clin Chem 53(2): 195-203.
- Oberacher H, Niederstätter H, Florian P, Walther P (2006) Profiling 627 mitochondrial nucleotides via the analysis of a 23-plex polymerase chain reaction by liquid chromatography-electrospray ionization time-of-flight mass spectrometry. Anal Chem 78(22): 7816-7827.
- Kramer KA, Devin O, Stacy J Hartman, Joe H, Bambi A, et al. (2005) Automated spectrophotometric analysis of mitochondrial respiratory chain complex enzyme activities in cultured skin fibroblasts. Clin Chem 51(11): 2110–2116.
- Keeney PM, Jing Xie, Roderick A Capaldi, James P Bennett Jr (2006) Parkinson's disease brain mitochondrial complex I has oxidatively damaged subunits and is functionally impaired and misassembled. J Neurosci 26(19): 5256-5264.
- Forner F, Leonard J Foster, Stefano C, Giorgio V, Matthias M (2006) Quantitative proteomic comparison of rat mitochondria from muscle, heart, and liver. Mol Cell Proteomics 5(4): 608-619.
- Johnson DT, Robert Harris A, Paul Blair V, Robert BalabanS (2007) Functional consequences of mitochondrial proteome heterogeneity. Am J Physiol Cell Physiol 292(2): C698–C707.
- Calvo S, Mohit J, Xiaohui X, Sunil Sheth A, Betty C, et al. (2006) Systematic identification of human mitochondrial disease genes through integrative genomics. Nat Genet 38(5): 576-582.
- Spinazzola A, Carlo V, Erika Fernandez-Vizarra, Franco C, Pio D'Adamo, et al. (2006) MPV17 encodes an inner mitochondrial membrane protein and is mutated in infantile hepatic mitochondrial DNA depletion. Nat Genet 38(5): 570-575.
- Mootha VK, Pierre Lepage, Kathleen Miller, Jakob Bunkenborg, Michael Reich, et al. (2003) Identification of a gene causing human cytochrome c oxidase deficiency by integrative genomics. Proc Natl Acad Sci U S A 100(2): 605-610.
- Saft C, Jochen Z, Jürgen A, Müller K, Katrin L, et al. (2005) Mitochondrial impairment in patients and asymptomatic mutation carriers of Huntington's disease. Mov Disord 20(6): 674-679.
- Ross B, Alexander L, Kent H, Pratip B, Brian S (2003) Clinical experience with 13C MRS in vivo. NMR Biomed 16(6-7): 358-369.
- Gourley PL, Judy KH, Anthony E McDonald, Guild Copeland R, Keith EB, et al. (2005) Mitochondrial correlation microscopy and nanolaser spectroscopy - new tools for biophotonic detection of cancer in single cells. Technol Cancer Res Treat 4(6): 585-592.
- Gourley CR, Naviaux RK (2005) Optical phenotyping of human mitochondria in a biocavity laser. IEEE J Selected Topics Quantum Electronics. 11(4): 818–826.
- Weissig V, Sarathi VB, Shing-Ming C, Gerard GM D’Souza (2006) Liposomes and liposome-like vesicles for drug and DNA delivery to mitochondria. J Liposome Res.16(3): 249-264.
- D'Souza GG, Boddapati SV, Weissig V (2007) Gene therapy of the other genome: the challenges of treating mitochondrial DNA defects. Pharm Res 24(2): 228-238.
- Cavalier-Smith T (2006) Origin of mitochondria by intracellular enslavement of a photosynthetic purple bacterium. Proc Biol Sci 273(1596): 1943-1952.
- Tuby H, Maltz L, Oron U (2007) Low-level laser irradiation (LLLI) promotes proliferation of mesenchymal and cardiac stem cells in culture. Lasers Surg Med 39(4): 373-378.
- Desmet KD, David AP, Jesse JC, Janis TE, Margaret TT Wong-Riley, et al. (2006) Clinical and experimental applications of NIR-LED photobiomodulation. Photomed Laser Surg 24(2): 121-128.
- Lampl Y, Justin AZ, Marc Fisher, Robert Lew, Lennart Welin, et al. (2007) Infrared laser therapy for ischemic stroke: a new treatment strategy: results of the NeuroThera Effectiveness and Safety Trial-1 (NEST-1). Stroke 38(6): 1843-1849.






























