- Research Article
- Abstract
- Introduction
- Proteomic Analysis of Stem Cells/Serum/Plasma Is Effective to Search For Cancer Diagnostic Markers
- Separation Technologies in Proteomic Analysis
- Mass Spectrometry
- Identification of Proteins/Peptides Using MS
- Characteristics of Proteomic Cancer Biomarkers and Diagnostic Availability
- Conclusion
- Acknowledgement
- References
New Approaches of proteomics in cancer and stem cells: Part I: from Basic Research to Clinical Application
Rakesh Sharma1,2, Shashi Prabha Singh2, Yuvraj Singh Negi1 and Robert Moffatt3
1Indian Institute of Technology, Roorkee, India
2Government Medical College, Saharanpur, India
3Florida State University, Tallahassee, USA
Submission: February 18, 2021; Published: November 09, 2022
*Corresponding Address: Rakesh Sharma, Indian Institute of Technology, Roorkee & Government Medical College, Saharanpur, India
How to cite this article: Rakesh S, Shashi P S, Yuvraj S N, Robert M. New Approaches of proteomics in cancer and stem cells: Part I: from Basic Research to Clinical Application. Canc Therapy & Oncol Int J. 2022; 22(4): 556094. DOI:10.19080/CTOIJ.2022.22.556094
- Research Article
- Abstract
- Introduction
- Proteomic Analysis of Stem Cells/Serum/Plasma Is Effective to Search For Cancer Diagnostic Markers
- Separation Technologies in Proteomic Analysis
- Mass Spectrometry
- Identification of Proteins/Peptides Using MS
- Characteristics of Proteomic Cancer Biomarkers and Diagnostic Availability
- Conclusion
- Acknowledgement
- References
Abstract
Proteomic analysis based on the separation and identification of proteins/peptides is a powerful tool for discovering new biomarkers. Two-dimensional differential gel electrophoresis (2D-DIGE), the ProteinChip array®, ClinProt®, and liquid chromatography are known as separation technologies. Although the detection and identification of proteins/peptides could be performed using mass spectrometry (MS), the types of MS used are diverse and the characteristics depend on the ionization method (e.g., matrix-assisted laser desorption/ionization [MALDI], surface-enhanced laser desorption/ionization [SELDI], and electrospray ionization [ESI]) and mass analyzer (e.g., time-of-flight [TOF], quadrupole [Q], and ion trap [IT]). The diverse proteomic systems that couple separation technologies with MS have been successfully applied to search for cancer biomarkers in biological fluid such as serum. Until now, many cancer biomarkers have been reported to be identified from serum and plasma. In order to identify diagnostic biomarkers of hepatocellular carcinoma (HCC) and adult T-cell leukemia (ATL), which are both human viral malignancies, we have been analyzing sera from patients using the ProteinChip array®-SELDI system and MALDI-TOF-TOF-MS. In this review, we present an overview of proteomic technologies used to search for biomarkers and cancer biomarkers discovered by these technologies, including our data on HCC and ATL. Cancer diagnosis using these proteomic biomarkers is also described.
Keywords: Cancer; Stem Cells; Hepatocellular Carcinoma; Mass Spectrometry; MALDI; SELDI; Electrospray ionization; Proteomics
- Research Article
- Abstract
- Introduction
- Proteomic Analysis of Stem Cells/Serum/Plasma Is Effective to Search For Cancer Diagnostic Markers
- Separation Technologies in Proteomic Analysis
- Mass Spectrometry
- Identification of Proteins/Peptides Using MS
- Characteristics of Proteomic Cancer Biomarkers and Diagnostic Availability
- Conclusion
- Acknowledgement
- References
Introduction
Cancer remains the major devastating disease throughout the world. Currently, it is estimated that cancer kills over 6 million people per year worldwide, with over 10 million new cases diagnosed every year. Despite advances in diagnostic imaging technologies, surgical management, and therapeutic modalities, the long-term survival is poor in most cancers. For example, the five-year survival rate is only 14% in lung cancer and 4% in pancreatic cancer [1,2]. The majority of cancers are detected in their advanced stages, and some have distant metastases, which are thought to be the causes of the current treatment ineffectiveness.
It is widely accepted that early diagnosis and intervention are the best way to cure cancer patients [3,4]. Cancer biomarkers provide diagnostic, prognostic, and therapeutic information about a particular cancer and show their ever-increasing importance in the early diagnosis of cancer [5,6]. Over the past several decades, enormous efforts have been made to screen and characterize useful cancer biomarkers. Several biomarkers approved clinically are available for early diagnosis and/or the successful monitoring of treatment and relapses and have contributed significantly to reduce mortality rates and increase overall survival for cancers such as prostate cancer [5,7]; however, unfortunately, most biomarkers are not satisfactory because of their limited specificity and/or sensitivity [8–10]. Therefore, there is an urgent need to discover better potential biomarkers for cancer diagnosis in clinical practice.
Global analyses have opened new avenues for cancer biomarker discovery [11–13]. Genomics, transcriptomics, proteomics, and metobolomics are studies of all genes, transcripts, proteins, and metabolites, respectively. From various specimens, such as cells, tissues, and body fluids, biomarkers detected differentially between cancer patients and healthy individuals have been investigated. In particular, serum/plasma is a less-invasive specimen that is suited to cancer diagnosis. Because serum/plasma has a high protein content (60–80 mg/ ml), analyzing all included proteins using proteomic analysis is efficient for diagnostic biomarker discovery.
Proteomic analysis is widely used to discover cancer biomarkers and is based on protein separation and identification technologies [14–17]. To search for diagnostic biomarkers, proteins/peptides in specimen are first separated with separation technologies. These expression profiles are then compared between cancer and control groups. Next, proteins/peptides differentially expressed under disease conditions can be identified using mass spectrometry (MS). Using proteomic analysis of serum/plasma, many cancer biomarkers have been reported to be identified [15,17]. Using proteomic technologies, we have also analyzed sera from patients with hepatocellular carcinoma (HCC) and adult T-cell leukemia (ATL), which are both human viral malignancies [18, 19]. In this review, we present an overview of the proteomic technologies used to search for biomarkers and the cancer biomarkers discovered by these technologies, including our data on HCC and ATL. Cancer diagnosis using these proteomic biomarkers is also described (Table 1).

- Research Article
- Abstract
- Introduction
- Proteomic Analysis of Stem Cells/Serum/Plasma Is Effective to Search For Cancer Diagnostic Markers
- Separation Technologies in Proteomic Analysis
- Mass Spectrometry
- Identification of Proteins/Peptides Using MS
- Characteristics of Proteomic Cancer Biomarkers and Diagnostic Availability
- Conclusion
- Acknowledgement
- References
Proteomic Analysis of Stem Cells/Serum/Plasma Is Effective to Search For Cancer Diagnostic Markers
There has been increasing interest in global analyses for the diagnosis of cancer [11–13]. Diagnostic biomarkers are currently being investigated throughout the world using genomic, transcriptomic, proteomic, and metabolomic analyses. In general, a biomarker whose expression is increased or decreased specifically under disease conditions is required to develop a new diagnostic method. Although specimen matrices such as serum/plasma, urine, and biopsy tissues are usually analyzed, the method needs to be chosen appropriately depending on the developmental purposes, cancer types, and specimen.
Characteristics of Global Analyses of Stem cells and Diagnostic Availability of Biomarkers
Table 1 shows the characteristics of global stem cell analyses. Genomic analysis is based on DNA sequencing of genes. By examining the whole genome sequence, we can obtain information about altered genetic and epigenetic events. Single nucleotide polymorphisms (SNPs) are known as genetic events and thought to be associated with disease susceptibility and individual responses to drug treatment [20]. While many SNPs are silent with no direct effect on proteins as gene products, some affect the coding or regulatory (usually promoter) regions of a gene and change the function and expression of translated proteins. There is a report that SNPs of the liver intestine-cadherin gene (CDH17) trigger aberrant splicing and are associated with the developmental risk of HCC [21]. As epigenetic events, DNA methylation has been suggested to be associated with carcinogenesis [22]. Characteristically, DNA methylation does not change genetic information; however, it alters the readability of DNA and results in gene inactivation. In particular, hypermethylation of CpG islands in promoter sequences is associated with the silencing of tumor suppressor genes and tumor-related genes by subsequent down-regulated expression of mRNA transcripts. Suppressors of cytokine signaling 1 and 3 (SOCS-1 and SOCS-3), which play important roles in cell growth and immune reaction, have been reported to be inactivated by hypermethylation in several cancers [23]. From these findings, it is suggested that SNPs and DNA methylation could be used clinically as genetic and epigenetic biomarkers for cancer diagnosis, respectively. Considering clinical applications, SNP and methylation arrays, by which it becomes easy to detect cancer-specific changes using extensive probe sets, might be useful. In addition, the translated proteins whose expression, amino acid sequences, and function are altered by SNPs and methylation may also be available for diagnosis as protein biomarkers.
Transcriptomic analysis is based on DNA microarray and is used to investigate extensive gene expression at the mRNA level [24,25]. Generally, mRNAs extracted from cancer stem cells and tissues are reverse transcribed to cDNAs and then hybridized with complementary oligonucleotide DNA probes. The expression levels of each mRNA can be measured by detecting hybridized cDNA. Because DNA probes for several tens of thousands of transcripts are fixed on the basal plate over a couple of centimeters, the expressions of all genes can be visualized simultaneously in just one experiment. Indeed, using Genechip arrays® (Affymetrix, Santa Clara, CA), the expression of about 50,000 transcripts can be examined. As part of the investigations into the ATLdevelopmental mechanism, Morishita’s group, in collaboration with us, applied DNA microarray to biomarker search [26]. The comparison of ATL cells derived from ATL patients with normal T-cells from healthy individuals showed that tumor suppressor in lung cancer 1 (TSLC1), which is a cell adhesion molecule, is overexpressed in ATL cells. While TSLC1 is a gene that associates with tissue invasion, its gene product (protein) may be also useful for ATL diagnosis. Cancer-specific genes found by DNA microarray analysis and these products could be used as diagnostic biomarkers [27–29].
Proteomic analysis based on the separation and identification of proteins/peptides is widely used to discover biomarkers, investigate cancer development, and to clarify the action mechanism of drugs [14–17]. To search for diagnostic biomarkers, proteins/peptides in a specimen are first separated with diverse separation technologies and the expression profiles are compared between disease and control groups (Chapter 2). Next, proteins/ peptides differentially expressed in disease status can be identified using MS (Chapters 3 & 4). In clinical practice, novel diagnostic methods are especially expected to be developed using identified biomarkers (Chapter 5).
chaptr 2345
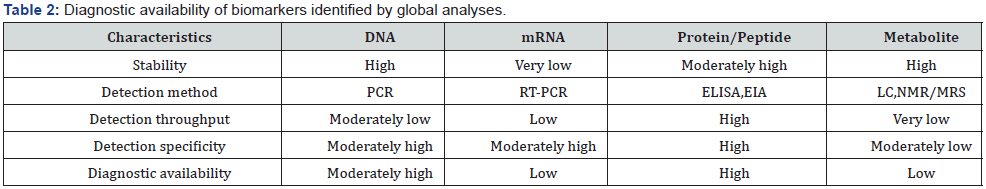
Metabolomics is the global study of all small molecular metabolites existing in stem cells and body fluids [30, 31]. Although metabolomic analysis consists of separation and identification processes as in proteomics, the target of analysis is small molecular compounds but not proteins/peptides [32]. For metabolomic separation, gas chromatography (GC) [33], LC [34], and capillary electrophoresis (CE) [35] are mainly used. Compounds are identified with MS [33–35] and nuclear magnetic resonance spectroscopy (NMR) [36,37], which are coupled together with separation technologies. Metabolomic analysis contributes to the understanding of how metabolites and their concentrations change under physiological conditions. At the same time, application of this analysis to the biomedical field may lead to the discovery of novel drug candidates and diagnostic protein biomarkers in stem cells and urine [38,39]. Recently, from more than 2,000 mass spectra of serum and urine metabolites with the expression of several peaks has been reported to be increased in renal cell carcinoma [40] (Table 2).
Advantages of Proteomic Analysis in the Search for Diagnostic Biomarkers in Cancer Stem cells
Genomic, transcriptomic, proteomic, metabolomic analyses provide information about disease-specific genes (SNPs and methylation sites), mRNAs, proteins, and small molecular metabolites in isolated stem cells. Considering that these biomarkers are actually used for diagnosis in clinical practice, developing measurement systems for them is necessary. The important requirements for diagnostic availability are simplicity of the detection procedure, high throughput, specificity of biomarker detection, and good reproducibility.
Table 2 shows the diagnostic availability of biomarkers identified by global analyses. At present, protein/peptide biomarkers appear to best meet these requirements. In particular, the use of specific antibodies makes it possible to develop measurement systems such as enzyme-linked immunosorbent assay (ELISA), which is used widely in clinical practice (Chapter 5).
In contrast to proteomic biomarkers, genomic and transcriptomic biomarkers are difficult to make available for widely used diagnostic systems at present. Genomic biomarkers such as SNPs and DNA methylation are detected by troublesome procedures, including DNA extraction from specimens and the amplification of target genes by polymerase chain reaction (PCR). Although its performance has been growing quickly recently, the throughput is still low for diagnostic application. Recently, a rapid SNP detection system named smart amplification process version 2 (SMAP 2) has reportedly been developed [41]; however, it is not common yet in clinical practice. Similarly, transcriptomic biomarkers are also poorly utilized for diagnosis. Although mRNA is detected by reverse transcriptase (RT)-PCR or Northern blotting, the method is tricky owing to mRNA flux. Collectively, it will take some time before the specific detection of genomic and transcriptomic biomarkers becomes common as a clinical laboratory test; however, if differential expression and SNPs can be detected at the protein level, ELISA using specific antibodies will become a widely used diagnostic system.
Metabolomic biomarkers are small molecular metabolites. They are expected to be applied in the diagnostic system; however, metabolomic analysis is still under development, in contrast to other global analyses. Although the identification of small molecular compounds is fundamental to the analysis, the database of mass and NMR spectra of metabolites is not yet complete. In addition, reference compounds for confirming identification are also lacking. Also, MS and NMR need extensive skills for their operation. Further, specific detection methods must be established with respect to each target compound; therefore, the development of a widely used diagnostic system using metabolic biomarkers is difficult. Recently, antibodies against small molecular compounds have been reported [42], thereby leading to the development of methods, although the specificity of antibodies may be low owing to few antigen determinants in the chemical structures of compounds. As described above, measurement systems using protein biomarker appear to be effective for the development of diagnostic methods. Consequently, it is reasonable that proteomic analysis using MS is focused on search for diagnostic biomarkers.
Specimen
Early diagnosis is very important to reduce cancer mortality. Protein biomarkers in cancer diagnosis can be obtained mainly from serum/plasma, urine, and pathologic tissues. Serum/plasma is a less-invasive specimen and is suitable for the early diagnosis. Because it constantly perfuses tissues, a certain abnormality in the living body is believed to be reflected in the expression and processing of contained proteins; for example, increased serum levels of prostate-specific antigen (PSA) and cancer antigen (CA) 125 are used to detect cancer in the prostate and ovary, respectively [43,44]. Urine is non-invasive and useful for biomarker search in addition to serum/plasma. While urine biomarkers are efficient, especially for urinary diseases such as bladder cancer [45], these concentrations in urine undergo many changes depending on diet, intake of water and drugs, and diseases. This is the difficulty in urine analysis. Pathologic tissues are frequently used in the definitive diagnosis of solid cancer such as HCC and kidney carcinoma; however, the pathologic diagnosis is highly invasive and unsuited to early diagnosis. In case of stem cell therapy of cancers, proteomics of stem cell samples is performed. From these characteristics of specimens, it is suggested that proteomic analysis of serum/plasma and stem cells is effective in searching for cancer diagnostic biomarkers and stem cell therapy.
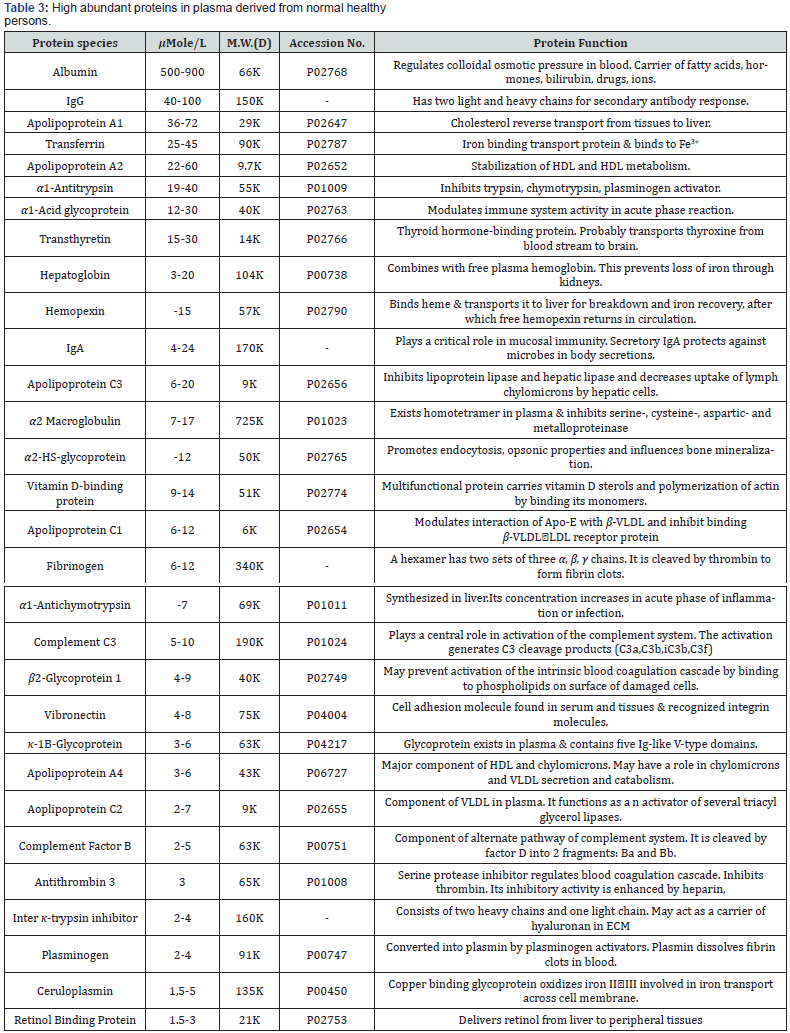
High protein content (i.e., 60–80 mg/ml) in serum/plasma and stem cells is attracting increasing interest in proteomic analysis. The expectation is that the characterization of thousands of individual proteins/peptides in serum/plasma/stem cells will enable the discovery of a number of reliable disease biomarkers; however, there is a difficulty in the proteomic analysis of serum/ plasma or stem cells. Over 99% protein content in serum/ plasma/stem cells is dominated by 30 proteins, such as albumin, immunoglobulins, transferrin, haptoglobulin, complement C3, lipoproteins and pleuripotent specific proteins [46,47] (Table 3). Serum/plasma proteins are present across an extraordinary dynamic concentration range that is likely to span more than 10 orders of magnitude. This dynamic range, namely the existence of highly abundant proteins, makes the detection of low abundant proteins (the remaining 1%) extremely challenging. There are two remedies for this difficulty. The first is the removal of highly abundant proteins by affinity columns using antibodies against them. The second is analysis aimed at peptides but not proteins. Regardless of this difficulty, considering that biomarkers discovered from serum/plasma are directly available for lessinvasive diagnosis, serum/plasma is an attractive specimen as the source of diagnostic biomarkers. However, stem cell proteomic biomarkers are less known or inconclusive (Table 3).
- Research Article
- Abstract
- Introduction
- Proteomic Analysis of Stem Cells/Serum/Plasma Is Effective to Search For Cancer Diagnostic Markers
- Separation Technologies in Proteomic Analysis
- Mass Spectrometry
- Identification of Proteins/Peptides Using MS
- Characteristics of Proteomic Cancer Biomarkers and Diagnostic Availability
- Conclusion
- Acknowledgement
- References
Separation Technologies in Proteomic Analysis
Proteomic analysis is based on the separation and identification of proteins/peptides in stem cells as a powerful tool for discovering new biomarkers. To search for diagnostic biomarkers from stem cells and serum/plasma, detection of proteins/peptides expressed in cancer is essential. Two-dimensional differential gel electrophoresis (2D-DIGE), ProteinChip array®, ClinProt®, and LC are typical separation technologies for biomarker discovery (Figure 1).
2D-DIGE
The gold standard for protein separation in proteomic analysis is 2D-DIGE (Figure 1A) [48,49]. First-dimensional separation is achieved by isoelectric focusing, which separates proteins on the basis of their charge. Namely, protein samples are applied to immobilized pH gradient (IPG) strips. The IPG strip is typically polyacrylamide gel in which the pH gradient (e.g., pH3–10) is immobilized. When the focusing voltage is applied to strips, the movement of a protein in a pH gradient is determined solely by its position relative to its isoelectric point (Figure 1a). If the position of the protein in the gradient is at a pH that is greater than its isoelectric point (pI), its net charge is negative, due to the net effect of dissociated carboxylate anions (negative charge) and uncharged amines in the protein; therefore, the protein migrates towards the positive electrode. If the position of the protein in the gradient is at a pH that is less than its pI, its net charge is positive, due to the net effect of uncharged carboxylic acid and protonated amines (positive charge). The protein then migrates towards the negative electrode. Any migration continues until the protein migrates to the position in the pH gradient that has no net charge. On this principle, the protein is focused at its pI on the IPG strip.
Isoelectric focusing is coupled with second-dimensional separation, which exploits sodium dodecylsulfate-polyacrylamide gel electrophoresis (SDS-PAGE) to separate proteins in accordance with molecular weight (Figure 1Ab). Under optimal conditions, the expression pattern of several thousands of individual proteins can be visualized simultaneously on a single 2D gel. The resulting gel shows the proteins separated according to pI across the x-axis of the 2D display and according to molecular weight across the y-axis (Figure 1).
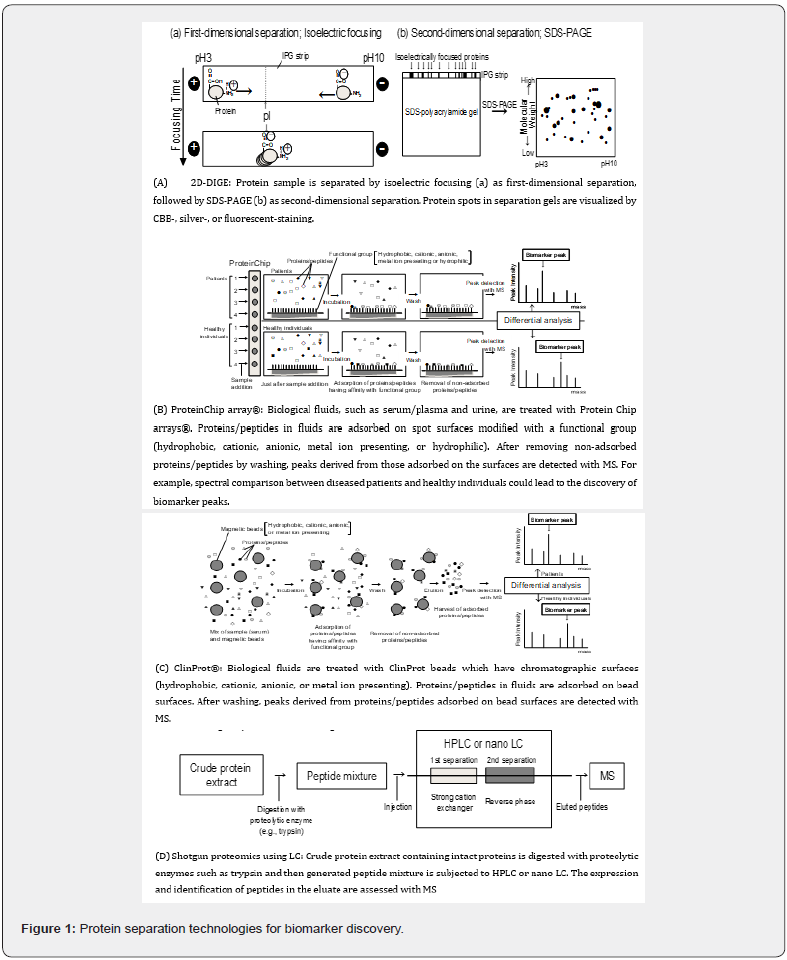

Protein profiles can be compared using sophisticated software packages to detect protein spots that are differentially expressed between samples. In general, protein spots are visualized by Coomassie brilliant blue (CBB) or silver staining [49]; however, CBB staining has low sensitivity and requires about 100 μg proteins per gel for spot detection, suggesting that this staining is not suited to proteomic analysis of valuable specimens. Although detection by silver staining can be performed with a small number of proteins (about 10 μg), irregular staining is often seen, leading to low reproducibility and quantification, which can be resolved using fluorescent dyes [50,51]. Fluorescent 2D-DIGE has high sensitivity and enables multiple protein extracts to be separated on the same 2D gel by labeling proteins using dyes with different excitation and emission wavelength pairs. Furthermore, because proteins are fluorescently labeled before electrophoresis, it is not necessary to consider the staining irregularity of gels. Recently, fluorescent 2D-DIGE has become popular for proteomic analysis [51].
ProteinChip Array®
ProteinChip array® (Bio-Rad Laboratories, Inc., CA, USA) has been successfully applied to diagnose various malignancies, such as ovarian cancer and gastric cancer [52,53]. As shown in Figure 1B, ProteinChip array® is a convenient separation tool that utilizes chromatographic properties and its surfaces can be modified with various functional groups (hydrophobic, cationic, anionic, metal ion presenting and hydrophilic). In this method, biological fluids, such as serum/plasma and urine, are directly applied to spot surfaces. After a series of binding and washing steps, proteins/ peptides and any contaminants that do not bind to the surface are removed. Protein mixtures retained on the surface are then analyzed with MS. In the binding step, the change of parameters (pH, salt concentration, and organic solvent concentration) in buffer solution alters the affinity between proteins/peptides and the surface, leading to the detection of a variety of proteins/ peptides. Also, pretreatment of the specimen with protein denaturants (e.g., urea) and detergents (e.g., CHAPS) affects the variation. Because differential analysis can be performed with a small amount of serum/plasma (a few microliters), ProteinChip array® is useful in biomarker searches using valuable specimens; however, the limitations of analysis include high cost and the fact that MS used in analysis is exclusive.
ClinProt®
ClinProt® (Bruker Daltonics, Bremen, Germany) is a purification method based on affinity beads (Figure 1C) [54, 55]. This method uses chromatographic surfaces (hydrophobic, cationic, anionic, and metal ion presenting) on an outer layer of magnetic beads. Certain subsets of proteins/peptides, which have affinity to the surface, can be selectively purified, allowing unbound impurities to be removed by washing with buffers. Proteins/peptides bound to magnetic beads are then eluted and directly analyzed by MS. The technical performance of affinity bead purification is similar to that of ELISA, and it can be used to process many samples in parallel. In addition, the cost of analysis is low, and data can be obtained with diverse types of MS (Figure 2).

Shotgun Proteomics Using LC
LC is a separation technology used commonly in the isolation of various biological molecules, including proteins/peptides from stem cells. The LC eluate is directly analyzed using MS in proteomic analysis [56,57] (Figure 1D). Because intact proteins cannot be identified owing to the measurement limits of MS, crude protein extract is digested in advance with proteolytic enzymes, such as trypsin, with the aim of directly identifying the LC eluate. The generated peptides, derived from a wide variety of proteins in extract, are subjected to LC and then sequences of peptides in the eluate are continuously determined using MS. Similarly, a method that identifies digested peptides in complex mixtures using a combination of LC and MS is referred to as “shotgun proteomics”. The name is derived from DNA shotgun sequencing, which is named for its analogy to the quasi-random firing pattern of a shotgun. Considering that LC eluate is directly analyzed with MS, in which involatile buffering agents and salts inhibit analysis, elution using a volatile buffer is necessary for peptide separation; therefore, elution with a water-acetonitrile gradient using reversed phase chromatography is mainly used to separate peptides prior to entering the MS. Furthermore, to enhance separation capacity, multidimensional LC (MDLC), which combines two or more forms of LC in shotgun analysis, has thrived over the past few years [58]. One of the most widely used combinations for MDLC is strong-cation exchange and reversed phase. In addition to the type of column, the flow rate is also important for separation. High performance LC (HPLC) and nano LC are generally used for peptide separation. In particular, nano LC is efficient for separating valuable specimens. In contrast to HPLC, its flow rate is extremely slow (typical flow rate: 20 nl/ min) [49]. This slow rate enables samples to be separated in a condensed state, indicating that analysis requires only a small amount of specimen.
- Research Article
- Abstract
- Introduction
- Proteomic Analysis of Stem Cells/Serum/Plasma Is Effective to Search For Cancer Diagnostic Markers
- Separation Technologies in Proteomic Analysis
- Mass Spectrometry
- Identification of Proteins/Peptides Using MS
- Characteristics of Proteomic Cancer Biomarkers and Diagnostic Availability
- Conclusion
- Acknowledgement
- References
Mass Spectrometry
Proteins/peptides separated using diverse technologies can be identified using MS, which is now the gold standard for identification. In this chapter, MS used in proteomic analysis is described.
Fundamentals of MS
A block diagram of a basic MS is shown in Figure 2. The sample inlet, ion source, mass analyzer, detector, vacuum system, control system, and data system are the major components [49]. Control system monitors and controls the other components. The mass analyzer, detector and parts of the ion source are maintained under vacuum. MS equipment is mainly defined by details of the ion source and mass analyzer. Analytes introduced from the sample inlet are ionized by various ionization methods and then ions are separated by a mass analyzer. Masses and quantities of their ions are shown as the mass-to-charge ratio (m/z) and peak intensity, respectively.

Ionization methods for biological macromolecules, such as proteins/peptides, were developed in the 1980s. Matrix-assisted laser/ionization (MALDI) is a soft ionization technique used in the analysis of proteins/peptides, which are fragile when ionized by conventional methods [59,60]. As shown in Figure 3A, proteins/ peptides are dissolved in a solution of a UV-absorbing compound, referred to as the “matrix”, and placed on sample stage. For peptide ionization, α-cyano-4-hydroxycinnamic acid (CHCA) is widely used as the matrix. Also, protein is ionized using sinnapic acid (SA) and 2,5-dihydroxy acetophenone (DHAP). As the solvent dries, the matrix compound crystallizes, and proteins/peptides molecules are included in matrix crystals. Pulses of UV laser light are used to vaporize the matrix and the included proteins/ peptides are carried into the gas phase. Because most matrixes are acidic compounds and dilute acid (e.g., trifluoroacetic acid; TFA) is added to the solvent, ionization occurs by protonation in the acidic environment. While monomolecular protons are normally added to a protein/peptide, multicharged ions are occasionally generated. These generated ions are then introduced into a mass analyzer. Surface-enhanced laser desorption/ionization (SELDI) is another ionization method that modifies MALDI and is a specialized method for ProteinChip array® (Chapter 2.2).
Along with MALDI, the ionization method used widely in proteomic analysis is electrospray ionization (ESI) [61,62]. Although the mechanism underlying ESI has not been fully clarified, its well-regarded model is illustrated in Figure 3B. In ESI, an acidic solution containing proteins/peptides is sprayed directly into the MS inlet through a small-diameter needle. A high, positive voltage is applied to this needle and droplets of the solution are sputtered from the tip. Protons from acidic conditions give the droplets a positive charge, causing then to move from the needle towards the negatively charged instrument (relative to the needle). During this movement, the size of the droplets is reduced by evaporation, a process that can be aided by a flow of nitrogen gas and heat. Protonated proteins/peptides desorbed from droplets to the gas phase are introduced into the mass analyzer. Characteristically, a significant portion of ions produced by ESI is multiply charged. Because proteins/peptides dissolved in liquid can be analyzed directly, ESI is coupled together with LC and the eluate obtained by solvent gradient is analyzed in waves (Figures 3 & 4).

Mass Types of MS Used in Proteomic Analysis
Mass analyzers separate ions according to their mass by applying electric and magnetic fields. Although the unit for their mass is the mass-to-charge ratio (m/z), care should be taken with the interchangeable use of “mass” and “m/z” because these values are not the same for any ion that is multiply charged. For example, a peptide ion with a mass of 1,000 Da that is doubly charged by the addition of two protons would be observed as an ion at 501 m/z. It is critical to remember this distinction, particularly when ESI, by which multicharged ions are mainly generated, is being used (Figure 3B). Time-of-flight (TOF), quadrupole (Q), and ion trap (IT) mass analyzers are used in MS for protein/peptides analysis [49].
The operating principles of the TOF mass analyzer are simple. As shown in Figure 4Aa, ions are first given a fixed amount of kinetic energy by acceleration in an electric field generated by a high voltage, typically +20 kV to +30 kV (for positive ions such as peptides). Following acceleration, ions enter a field-free region where they travel at velocities based on a law. Namely, their velocities are inversely proportional to their m/z; therefore, ions with low m/z travel more rapidly than ions with high m/z. The time required for the ion to travel the length of the field-free region is measured and used to calculate the m/z of the ion. To obtain a higher resolution, while longer flight tubes or higher accelerating voltages are effective, an ion reflector, also referred to as a “reflectron”, is built into high resolution MS. As shown in Figure 4Ab, the reflectron is an ion mirror created by an electric field and reverses the flight path of the ion. In theory, ions with the same mass travel in a flight tube with the same velocity; however, in fact, there is accidental error in the velocity. For example, ions with higher velocity than the theoretical value penetrate the flight tube further. In the reflectron, higher-velocity ions take longer to reverse the flight path. This process reduces the error in the velocity. Also, another effect of a reflectron is the increased flight distance, which also leads to better resolution.
The Q mass analyzer consists of four parallel metal rods (quadrupole) through which ions pass (Figure 4B). When opposite and neighbor rods are subjected to electric potentials having the same and different polarities, respectively, an electrostatic field of specific strength and frequency is created inside the quadrupole. The path of ions is influenced by the fluctuating electric field. Only ions of pre-selected mass make it through the inside diameter of the quadrupole and strike the detector (circular ion in Figure 4B). All other ions are deflected out of the quadrupole and never reach the detector (triangular ion in Figure 4B). Further, the Q mass analyzer can scan pre-selected masses at rates on the order of 1000 m/z per sec so that the acquisition of a mass spectrum between 200 m/z and 2000 m/z takes 1.8 sec. This high-speed scan is suitable for acquiring data on LC, from which analytes are continuously eluted.
The IT mass analyzer works on the same physical principle as the Q mass analyzer. The term “ion trap” is derived from the fact that electrostatic fields are applied so that ions of all m/z are initially trapped in the trapping region (Figure 4C). Trapped ions are ejected out of the trapping region and into the detector by changing the electrostatic fields dependent on the m/z of ions. An important characteristic of IT is that the detection sensitivity is extremely good, because of the efficient transmission and utilization of ions formed by an ion source. The overall dimensions of the trap are such that the distance from the trapping region to the detector is only a few centimeters. In the course of the trapping and ejection process, all ions injected into the trap can ultimately be sent to the detector; therefore, the compact size of ion traps and the detection of all ions injected lead to good sensitivity.
Types of MS Used in Proteomic Analysis
As described above, although the types of ion source and mass analyzer are diverse, MS is composed of their variousIdentification of Proteins/Peptides Using MS combinations; for example, MALDI-TOF-MS, SELDI-TOF-MS, ESIQ- MS, and ESI-IT-MS are used in proteomic analysis. Furthermore, to perform the measurement with good mass accuracy and high resolution, tandem MS, in which multiple mass analyzers are connected in series, is used widely at present. Tandem MS, such as MALDI-TOF-TOF-MS (Figure 5A), ESI-Q-TOF-MS (Figure 5B), and ESI-IT-TOF-MS (Figure 5C), is indispensable for the identification of proteins/peptides.
- Research Article
- Abstract
- Introduction
- Proteomic Analysis of Stem Cells/Serum/Plasma Is Effective to Search For Cancer Diagnostic Markers
- Separation Technologies in Proteomic Analysis
- Mass Spectrometry
- Identification of Proteins/Peptides Using MS
- Characteristics of Proteomic Cancer Biomarkers and Diagnostic Availability
- Conclusion
- Acknowledgement
- References
Identification of Proteins/Peptides Using MS
The identification of proteins/peptides is an essential step in the field of proteomics. Examining protein expression patterns alone is, of course, important for diagnosis in various diseases; however, without identifying the proteins/peptides associated with diseases, it is not possible to delve into their fundamental causes or to develop clinical applications, such as ELISA, that require antibodies against biomarkers. The identification of proteins/peptides using MS is the key step in the development of clinical applications using proteomic biomarkers.
Protein Identification by Peptide Mass Fingerprinting
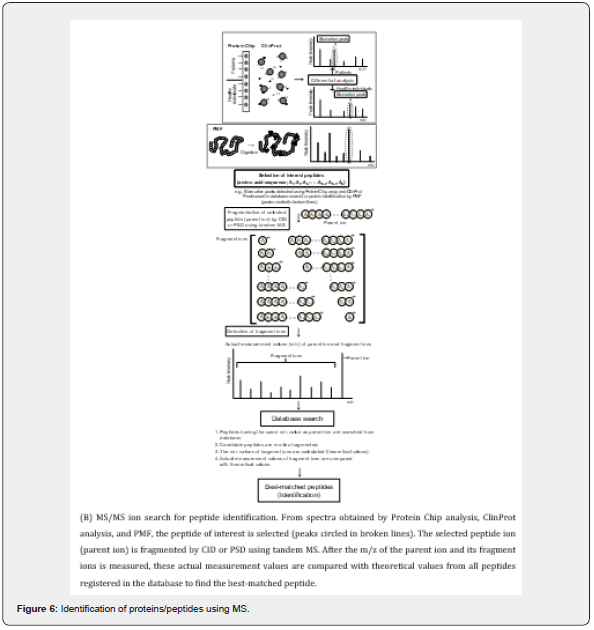
Peptide mass fingerprinting (PMF) is an analytical technique for protein identification [63–67] (Figure 6A). In PMF, after protein separation from stem cells, the stem cell proteins of interest are first digested with proteolytic enzymes such as trypsin. The m/z of generated peptides from digest is accurately measured with MS. These actual measurement values are then in silico compared with theoretical values of known proteins registered in a MUSCOT® database. As data on known proteins, registered proteins are theoretically digested into peptides with enzymes and the absolute masses of these reference peptides are calculated. By comparing the actual measurement values of the interest protein with theoretical values of the enormous numbers of proteins registered in the database statistically, the bestmatched protein is searched. Databases of the National Center for Biotechnology Information (NCBI) and Swiss-Prot are popular for identification. NCBI includes most of the public domain sequence databases and the Swiss-Prot database has a large amount of annotation together with the sequences to allow for easier understanding of the functionality of the identified proteins [68]. The advantage of PMF is that only the m/z of generated peptides has to be examined. De novo peptide sequencing, which is timeconsuming, is not necessary. A disadvantage is that the interest protein has to be registered in a database. Additionally, most PMF algorithms assume that the peptides come from a single protein [69]; therefore, the typical PMF samples are proteins isolated by 2D-DIGE and MDLC that have high separation capacity.

Peptide Identification by MS/MS Ion Search
Mass analysis is essentially a separation of ions according to their m/z. Because tandem MS has multiple mass analyzers that are connected in series, two stages of mass analysis (MS/ MS analysis) can be carried out in a single experiment. An ion with a specific m/z is selected in the first stage of mass analysis and structural information about selected ion is obtained in the second stage. In the case of peptide ions, the information is the amino acid sequence of the peptide. At present, peptides with a molecular weight of less than ~3,000 can be routinely identified by MS/MS ion search using tandem MS, such as MALDI-TOF-TOFMS, ESI-Q-TOF-MS, and ESI-IT-TOF-MS (Figure 5). MS/MS ion search is available to identify peptides detected by differential analyses using ProteinChip array®, ClinProt®, and shotgun proteomics, and to confirm the sequences of peptides used in protein identification by PMF.
MS/MS ion search is based on the fragmentation reaction of peptides and by database search [49,70,71] (Figure 6B). Namely, peptides ionized by ionization methods, including MALDI and ESI, have excess internal energy. After a peptide of interest (parent ion) is selected in the first mass analyzer, its peptide bonds are randomly cleaved by internal energy. While the fragmentation reaction occurs in ions with excess energy, most ions formed by MALDI and ESI have insufficient internal energy and are stable to drive any fragment reactions. Thus, what is required for MS/ MS analysis of peptides is a method to energize a stable parention after it has been mass-selected and to induce fragmentation reaction. The method used most often is collisionally induced dissociation (CID), in which a mass-selected ion is transmitted into a high-pressure region of the tandem MS where it undergoes a number of collisions with molecules of filler gas such as argon. As the ion collides with these gas molecules, a portion of the kinetic energy of the ion is converted into internal energy to make the ion unstable and drive the fragmentation reaction. In the Q-TOF mass analyzer (Figure 5B), after an ion is selected in the first stage (Q mass analyzer), it is fragmented in the collision cell. Also, in the IT-TOF mass analyzer (Figure 5C), the trapping region of IT serves as the cell. In contrast, in MALDI-TOF-TOF-MS (Figure 5A), CID is not often used for fragmentation. The selected ion decays into fragments without CID after leaving the acceleration region of the ion source; therefore, this fragmentation reaction is referred to as “post-source decay (PSD). In PSD, the reaction occurs mainly in the second stage of the mass analyzer (TOF2 in Figure 5A). The fragment ions produced are m/z analyzed in the second stage. Both the intact peptide mass and fragment ion masses are used for peptide identification through database searching (Figure 6B).


Cancer Biomarker and Diagnosis
Over the past several decades, enormous efforts have been made to screen and characterize useful cancer biomarkers. As clinically important biomarkers, PSA, carcinoembryonic antigen (CEA), α-fetoprotein (AFP), CA125, CA15-3, and CA19-9 have been identified [8–10]. Although they are commonly employed in clinical diagnosis, they are not satisfactory because of their limited sensitivity and/or specificity; for example, CA125 is widely used to detect ovarian cancer in post-menopausal women and is the best clinical marker currently available. Although CA125 levels are elevated in ~80% of patients with advanced stage ovarian cancer, serum levels are increased in only 50–60% of patients with the early stage [8,72–74]. Early diagnosis of cancers, including ovarian cancer, remains difficult. There is therefore an urgent need to discover better potential biomarkers for cancer diagnosis in clinical practice (Figures 5 & 6) (Table 4).

- Research Article
- Abstract
- Introduction
- Proteomic Analysis of Stem Cells/Serum/Plasma Is Effective to Search For Cancer Diagnostic Markers
- Separation Technologies in Proteomic Analysis
- Mass Spectrometry
- Identification of Proteins/Peptides Using MS
- Characteristics of Proteomic Cancer Biomarkers and Diagnostic Availability
- Conclusion
- Acknowledgement
- References
Characteristics of Proteomic Cancer Biomarkers and Diagnostic Availability
As described in Chapters 2–4, diverse types of protein separation technology and MS are proficiently combined in proteomic analysis. Although a variety of combined systems have been used in analysis, the 2D-DIGE–MS system and ProteinChip array®–SELDI-TOF-MS system are at present the main standards to search for new diagnostic biomarkers in many cancers [14–17]. Many studies have reported proteomic biomarkers in various cancers [75–105] (Figure 7).
Table 4 shows cancer biomarkers that have been identified by proteomic analysis of serum/plasma. Biomarkers whose expression is increased and/or decreased specifically in a particular cancer type have a great deal of potential in the diagnostic field; however, as the expression of α-antitrypsin has been reported to be increased in colorectal cancer [78], HCC [86], and ovarian cancer [96], the expression of most biomarkers is increased and/or decreased in multiple types of cancer. While proteins/peptides that are expressed differentially in various cancers constitute a large portion of cancer biomarkers, a few biomarkers are possibly specific to a particular cancer.
Neutrophil-activating peptide 2 (NAP-2) and Ras-related protein Rab-7B are expected to be specific biomarkers of HCC and lung cancer, respectively [87,91]. NAP-2 is the major form of CXCL7, which is a member of the chemokine family involved in regulating immunity angiogenesis, stem-cell trafficking, and mediating organ-specific metastases of cancers [106]. CXCL7 is translated as a propeptide and then cleaved to several smaller forms, including NAP-2. Although CXCL7 has been reported to be a marker of advanced myelodysplastic syndromes (MDS), hematologic stem cell malignancies in elderly patients [107], no expression of NAP-2 is observed in tissues of other types of cancer. Furthermore, serum levels of NAP-2 are correlated with the clinicopathological features and prognosis of HCC [108]. Rab7B, which is highly expressed in lung cancer, is a member of the Ras oncogene family and a small Rab GTPase that regulates vesicular traffic from early to late endosomal stages of the endocytic pathway. Rab7B is expressed in the heart, placenta, lung, skeletal muscle, and peripheral blood leukocytes [109]; however, no increase of its serum levels has been reported in cancers other than lung cancer. Consequently, these findings suggest that NAP-2 and Rab7B may be specific biomarkers of HCC and lung cancer, respectively, although many clinical studies must be performed using sera from patients with a variety of diseases.
The expression of serum amyloid A (SAA) is increased in a various cancer (Table 4). Although proteomic analysis of serum/plasma identified SAA in HCC [89], lung cancer [91], nasopharyngeal carcinoma [94], ovarian cancer [99], and renal cancer [105], its increased expression has also been reported in other cancers such as kidney, colorectal, and prostate cancers, as well as in leukemias and lymphomas [110]. In addition, SAA is an apolipoprotein secreted in an acute phase of inflammation [111]; therefore, its increased expression may represent the epiphenomenon of cancers. While SAA is unlikely to be of much clinical use for diagnosing the onset of cancer, it may be important to monitor the stage of cancer progression and relapse. In nasopharyngeal cancer, SAA has been reported to be useful for monitoring its relapse [112].
Although most proteomic biomarkers are not specific to a particular cancer, a combination of those with known diagnostic methods has been reported to improve the probability of cancer diagnosis [86,101]. Using CA125 alone as the best clinical marker of ovarian cancer, patients with early-stage cancer are diagnosed with a receiver operating characteristic area under the curve (ROC AUC) of 0.613. In contrast, when four proteomic biomarkers (transthyretin, hemoglobin subunit β, apolipoprotein A1, and transferrin) are combined with CA125, the diagnosis is performed with a ROC AUC of 0.955 [101]. The four biomarkers are highly abundant proteins and are not likely to be synthesized at the focal site. Thus, these biomarkers, even if they are not released directly by the tumor, could be used in combination with other markers, such as CA125, to improve the diagnostic probability of earlystage ovarian cancer. In another instance, AFP is a biomarker for HCC diagnosis. By 2D-DIGE of sera from HCC patients followed by protein identification using MALDI-TOF-TOF-MS, heat-shock protein 27 (HSP27) was identified as an HCC biomarker [86] (Table 4). HSP27 is a stress-inducible cytosolic protein that is ubiquitously present in many normal tissues and has been reported to be over-expressed in many types of cancer, such as breast cancer and astrocytic brain tumor [113,114]. HSP27 is not a specific cancer biomarker; however, it could be detected in HCC patients with negative AFP [86]. This suggests that HSP27 could be complementary to AFP in the diagnosis of HCC. Furthermore, combination diagnosis of HSP27 and AFP improves the diagnostic probability of HCC patients with small tumors (< 5 cm). By the same token, leucine-rich α2 glycoprotein (LRG) has been reported to serve as a complementary biomarker of CA19.9, which is a tumor marker commonly used for the diagnosis of pancreatic cancer [102].
Early Diagnosis of HCC Using ProteinChip Array®
In cancer diagnosis, early detection before the onset of clinical symptoms is very important. Early diagnosis can lead to curative treatment, significantly improving prognosis. Diagnosing patients who present without symptoms, however, is difficult, because the tumor is still small and there are few lesions. Using the ProteinChip array®–SELDI-TOF-MS system, early diagnostic methods have been reported in several cancers, including our data on HCC [18,115–117].
Approximately 170 million people worldwide are infected with hepatitis C virus (HCV), which when persistent can progress to HCC. Interferon (IFN) or combined IFN and ribavirin are currently the only effective treatments for chronic hepatitis C, leading to a decrease of HCC occurrence [118,119]; however, some patients do not receive IFN treatment or fail to recover from HCV even with IFN treatment. In addition, a subset of individuals remains unaware that they are infected with HCV. Namely, in these patients, HCC may present only in the advanced stage. The prognosis of patients presenting with symptoms related to HCC is extremely poor. One requirement is the early detection of HCC before the onset of clinical symptoms.
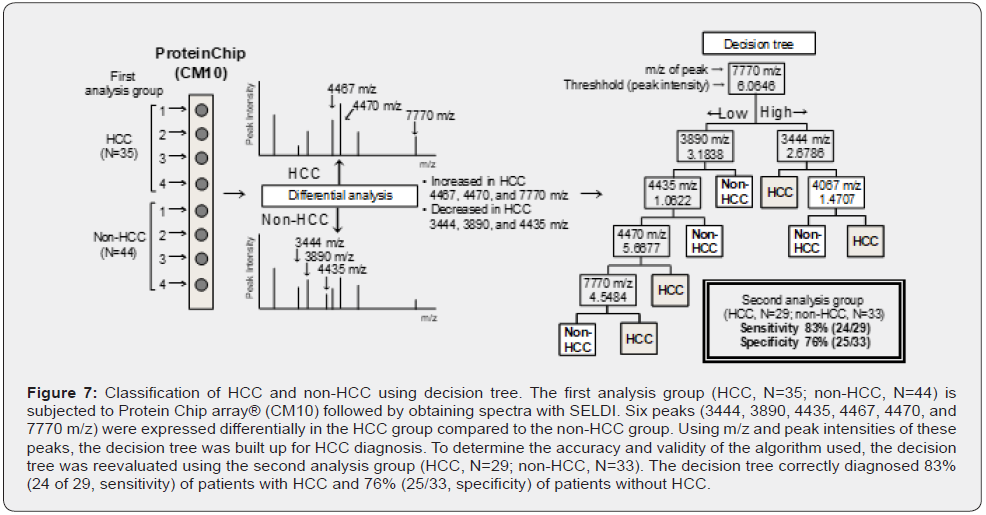
We have been studying the developmental mechanism of HCC [120–122]. As part of the investigation, we sought to identify novel diagnostic markers of HCC using the ProteinChip array®–SELDITOF- MS system and to develop a diagnostic system using these markers. The decision tree for HCC diagnosis was established using sera from patients with and without HCC (Figure 7) [18].
The first analysis group (HCC, N=35; non-HCC, N=44) was subjected to ProteinChip array® (CM10) followed by obtaining spectra with SELDI. Six peaks (3444, 3890, 4435, 4467, 4470, and 7770 m/z) that were expressed differentially in the HCC group compared to the non-HCC group were used for diagnosing HCC. The decision tree generated using m/z and peak intensities correctly classified 97% of HCC samples. Further, to determine the accuracy and validity of the algorithm used, the decision tree was reevaluated using the second analysis group (HCC, N=29; non- HCC, N=33). The decision tree correctly diagnosed 83% (24 of 29, sensitivity) of patients with HCC and 76% (25/33, specificity) of patients without HCC. Importantly, the accuracy of decision-tree diagnosis for HCC was higher than that of other known tumor markers such as AFP (sensitivity, 41%; specificity, 67%) (Figure 8).
The most fundamental requirement for serum-based marker detection is the identification of carcinoma in an early stage when treatment has the greatest impact on prognosis. Decisiontree diagnosis was performed using serum samples taken from 7 patients 1 year before HCC development. Six of the 7 (86%) patients who later developed HCC were diagnosed with HCC using the decision tree. One year before development, HCC was still undetectable by the current diagnostic methods. It is suggested that this decision tree is useful for the early diagnosis of HCC.
Proteomic Strategy for Discovering Specific Cancer Biomarkers
As shown in Table 4, many cancer biomarkers have been identified by serum/plasma proteomic analysis. While specific biomarkers such as NAP-2 and Rab7B are discovered fairly infrequently [87,91], most of these are unfortunately not specific to a particular cancer. In addition, the proteomic biomarker is mostly highly abundant proteins (Table 3). Certainly, non-specific and/or highly expressed biomarkers are useful to complement conventional diagnosis methods [86,101] and for monitoring cancer relapse [112]; however, because they are not necessarily released directly from tumor tissues, their diagnosis methods may have little scientific basis. Discovering specific cancer biomarkers that are based on scientific evidence are important for the development of novel diagnostic methods.
Proteomic analysis of low abundant protein (less than 1% of serum/plasma protein content) in serum/plasma may lead to discovering specific biomarkers. Although shotgun proteomics using LC (Figure 1D) and tandem MS (Figure 5B, C) has been established for serum/plasma analysis [123,124], cancer biomarkers useful for early diagnosis are still unknown. Proteomic approaches based on other perspectives might also be needed. Many researchers have hypothesized that the best cancer biomarkers were likely to be secreted proteins [125]. In addition, many classical cancer biomarkers, such as CEA and CA15.3, are bound to the cell membrane, but their extracellular domains could be found, through shedding, in the circulatory system. On the basis of this hypothesis, Roessler et al. aimed to identify novel serum biomarkers of colorectal cancer [126]. First, 16 matched colorectal cancer and adjacent normal tissue samples were subjected to 2D-DIGE followed by protein identification using MS. Five proteins whose expression is increased markedly in colorectal cancer were identified as transforming growth factor-β-induced protein ig-h3 (βIG-H3), nicotinamide N-methyltransferase (NNMT), nucleoside diphosphate kinase A (nm23-H1), purine nucleoside phosphorylase (PNPH), and mannose-6-phosphate receptor binding protein 1 (M6P1). Validation using ELISA showed that elevated levels of NNMT were found in sera from patients with colorectal cancer. Furthermore, the accuracy of NNMT diagnosis (ROC AUC, 0.84) was higher than that of the established tumor marker CEA (ROC AUC, 0.78). Similarly, proteomic analysis of tumor tissues occasionally leads to discovering serum/plasma biomarkers. Using the same strategy, phosphoglycerate kinase 1 (PGK1) has reportedly been identified as a serum biomarker of pancreatic ductal adenocarcinoma [127].
High abundant proteins are possibly suitable to search for specific cancer biomarkers. As shown in Table 4, high abundant proteins expressed differentially in various cancers appear capable of being categorized in three groups: 1) components of complement system and regulated proteins (C3, C4, and factor B etc.), 2) apolipoproteins (apolipoproteins A1, A2, and C1 etc.), 3) iron-metabolic proteins (transferrin, haptoglobin, and hemopexin etc.). Furthermore, as seen in C3a [75,78,85] and fibrinopetide A [81,82,90], which are fragment peptides derived from complement C3 and fibrinogen, respectively, protein cleavage products also serve as cancer biomarkers. Changes in the expressions and/or processing patterns of highly abundant proteins in cancer may be caused by specific molecules in pathogens. We are focusing on the relationships between the complement system and ATL, a fatal leukemia caused by infection with human T lymphotropic virus type-1 (HTLV-1) [19].
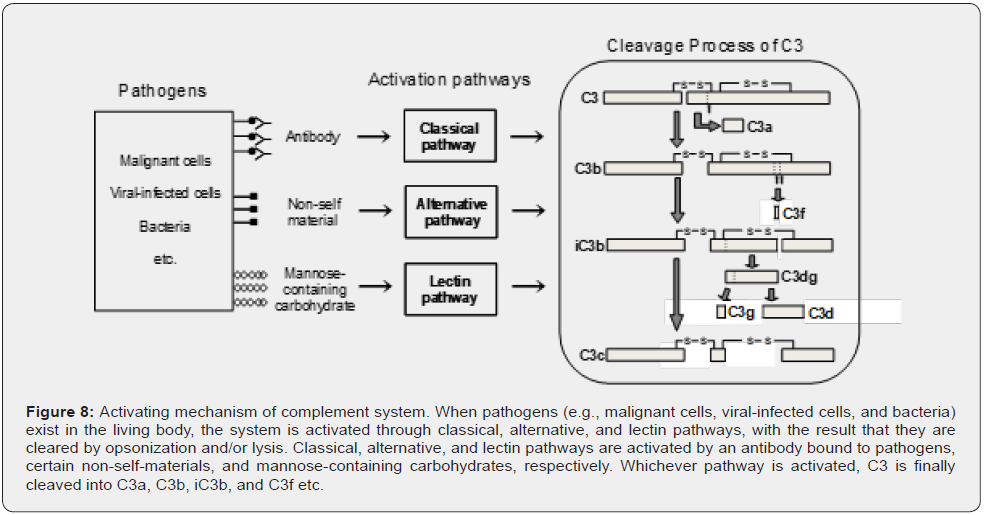
The complement system is a major mediator of innate immune defense [128–130]. The system consists of a group of over 30 proteins, such as C3, C4, and factor B. When pathogens (e.g., malignant cells, viral-infected cells, and bacteria) exist in the living body, the system is activated, with the result that they are cleared by opsonization and/or lysis. The complement system has three distinct pathways (classical, alternative, and lectin pathways) (Figure 8). Classical, alternative, and lectin pathways are activated by an antibody bound to pathogens, certain nonself- materials, and mannose-containing carbohydrates expressed on the pathogen surface, respectively. Whichever pathway is activated, C3 is finally cleaved into C3a, C3b, iC3b, and C3f etc. The increase of C3 cleavage products in cancer patients is a reflection of this activation.
We identified C3f as the serum biomarker of ATL by differential analysis using the ProteinChip array®–SELDI-TOF-MS system followed by MS/MS ion search using MALDI-TOF-TOF-MS [19]. Further, analysis of markers specific to the activation pathways showed that the serum concentration of the marker of the lectin pathway was significantly higher in ATL patients. These results suggest that the complement system is activated through the lectin pathway in ATL. Furthermore, this implies the presence of a trigger that activates the lectin pathway. As one possibility, it is considered that aberrant glycoproteins and glycolipids, including mannose-containing carbohydrates, are expressed on the surface of ATL cells. C3f is not specific to ATL, because its expression is increased in various cancers [19]. The trigger is, however, possibly suitable for diagnosis as an ATL-specific biomarker. In addition to identifying proteins/peptides by proteomic analysis of serum/plasma, clarifying a molecule associated with the signal transduction pathway such as the complement system may be necessary to discovery specific cancer biomarkers.
From Biomarker Discovery to Clinical Application
The most important issue in developing diagnosis methods is reproducibility in clinical practice as well as in the laboratory. In particular, in the early diagnosis of cancer, because tumors are microscopic, insensible changes in the living body have to be captured. In addition, previous studies, including our data, indicated that measurement systems using multiple biomarkers are effective in early diagnosis [18,115–117]; therefore, measurement systems are required that reproducibly quantify multiple biomarkers simultaneously using just one experiment.
The ProteinChip array®–SELDI-TOF-MS system enables the detection of multiple proteins/peptides in just one experiment. Diagnosis can be performed by the decision tree using m/z and peak intensities without the identification of proteins/peptides. While many studies have reported diagnostic methods using decision trees, irreproducibility is a problem [131]. Although our experience also shows this, the irreproducibility could be attributed to analytical steps from the time the serum is collected until the spectrum is acquired. Unfortunately, no universal standard protocol exists for the collection, storage, and transport of serum samples. Also, there is no universal quality control of the decision tree and spectra obtained with MS. In particular, the spectral patterns acquired by SELDI-TOF-MS seem to be affected by the aging degradation of the ionization laser. Whether this is true, the cause of the irreproducibility is uncertain. If peaks useful for diagnosis are found, the better choice is to identify proteins/ peptides derived from these peaks followed by the development of other measurement systems with good reproducibility and quantitation (Figure 9).
ELISA is a biochemical technique used mainly in immunology to detect the presence of an antibody or an antigen in a sample [132]. ELISA has been widely used as a diagnostic system in clinical practice; for example, sandwich ELISA, which has good sensitivity, quantitation and reproducibility, is performed as follows (Figure 9): 1) the plate is coated with a capture antibody; 2) sample is added and any antigen present binds to capture the antibody; 3) the detecting antibody is added and binds to the antigen; 4) enzyme-linked secondary antibody is added and binds to the detecting antibody; 5) substrate is added and is converted by the enzyme to the detectable form. Early diagnosis of ovarian cancer has reportedly been performed using ELISA for proteomic biomarkers (transthyretin, hemoglobin subunit β, apolipoprotein A1, and transferrin) [101]. Also, as a high-throughput and multiplex measurement system, antibody array has recently become of interest [133]. To apply proteomic biomarkers in a clinical setting, the development of measurement systems, such as ELISA and antibody array, may be a key step.

Cancer MALDI imaging and HR-MAS NMR biopsy Diagnosis
MALDI is a technique to identify peptides from MS peak spectrum as described in section 4. Same technique is now extended in imaging these peptides in 3 dimensions to generate a 3-D tumor reconstruct. Cancer tissues are routinely processed for MALDI imaging to identify specific proteins. However, method suffers from technical as well as operator biases because of very rigorous expercise to pin point MS peaks from thousands of chemical species visible in spectra. MALDI imaging technique puts a new dimension of protein location in tumors and visualization. Recently many integration methods have been applied to pin point location of tumor specific proteins on images to make use of them in diagnosis and tumor staging. However, the use of imaging MALDI is limited as research tool possibly a future diagnostic modality in place of serum proteomics in oncology and cancer therapeutics.
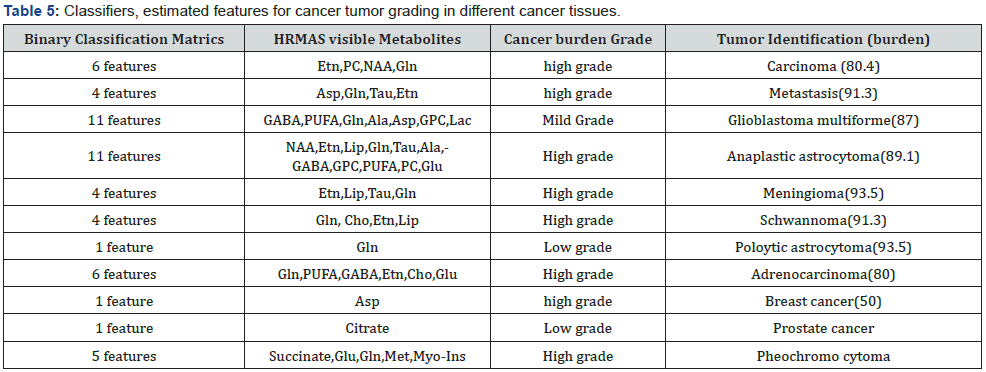
Total through-bond spectroscopy (TOBSY) of ex vivo HRMAS MR spectra of tumor biopsies from glioblastoma multiforme, Schwannoma, Pilocytic astrocytoma, adrenocarcinoma and breast metastasis showed distinct alanine, choline, GABA, glutamine/glutamate, glycerophosphocholine, lipids, myoinositol, phosphocholine, phosphoethanolamine, polyunsaturated fatty acids, taurine NMR resonance assignments as cancer classifiers as shown in Figure 9 and Table 5. These initial results strongly suggest the potentials of biopsy or intact tissue metabolic characterization [72] (Figure 9) (Table 5).
- Research Article
- Abstract
- Introduction
- Proteomic Analysis of Stem Cells/Serum/Plasma Is Effective to Search For Cancer Diagnostic Markers
- Separation Technologies in Proteomic Analysis
- Mass Spectrometry
- Identification of Proteins/Peptides Using MS
- Characteristics of Proteomic Cancer Biomarkers and Diagnostic Availability
- Conclusion
- Acknowledgement
- References
Conclusion
Proteomic analysis based on protein/peptide separation and identification using MS is an essential technology for biomarker discovery. Although most proteomic biomarkers may be not specific for a particular cancer, they are certainly of help in the early diagnosis of several cancers. There is considerable difficulty in discovering specific biomarkers; however, a variety of strategies for discovering specific biomarkers may make it possible. We hope that proteomic cancer biomarkers will be widely used in the future for early diagnosis in clinical practice.
- Research Article
- Abstract
- Introduction
- Proteomic Analysis of Stem Cells/Serum/Plasma Is Effective to Search For Cancer Diagnostic Markers
- Separation Technologies in Proteomic Analysis
- Mass Spectrometry
- Identification of Proteins/Peptides Using MS
- Characteristics of Proteomic Cancer Biomarkers and Diagnostic Availability
- Conclusion
- Acknowledgement
- References
Acknowledgement
We thank Dr Doris Terry, Bioproteomics Lab, Florida State University, Tallahassee for doing proteomics experiments of MCF- 7 induced cancer tumor tissues from rats.
- Research Article
- Abstract
- Introduction
- Proteomic Analysis of Stem Cells/Serum/Plasma Is Effective to Search For Cancer Diagnostic Markers
- Separation Technologies in Proteomic Analysis
- Mass Spectrometry
- Identification of Proteins/Peptides Using MS
- Characteristics of Proteomic Cancer Biomarkers and Diagnostic Availability
- Conclusion
- Acknowledgement
- References
References
- Yeo TP, Hruban RH, Leach SD, Wilentz RE, Sohn TA, et al. (2002) Pancreatic cancer. Curr Probl Cancer 26(4): 176-275.
- Chen G, Gharib TG, Wang H, Huang CC, Kuick R, et al. (2003) Protein profiles associated with survival in lung adenocarcinoma. Proc Natl Acad Sci USA 100(23): 13537-13542.
- Etzioni R, Urban N, Ramsey S, McIntosh M, Schwartz S, et al. (2003) The case for early detection. Nat Rev Cancer 3(4): 243-252.
- Yokota T, Ishiyama S, Saito T, Teshima S, Narushima Y, et al. (2004) Lymph node metastasis as a significant prognostic factor in gastric cancer: a multiple logistic regression analysis. Scand J Gastroenterol 39(4): 380-384.
- Ludwig JA, Weinstein JN (2005) Biomarkers in cancer staging, prognosis, and treatment selection. Nat Rev Cancer 5(11): 845-856.
- Lam T, Nabi G (2007) Potential of urinary biomarkers in early bladder cancer diagnosis. Expert Rev Anticancer Ther 7(8): 1105-1115.
- Sidransky D (2002) Emerging molecular markers of cancer. Nat Rev Cancer 2(3): 210-219.
- Menon U, Jacobs I (2000) Recent developments in ovarian cancer screening. Curr Opin Obstet Gynecol 12(1): 39-42.
- Chatterjee SK, Zetter BR (2005) Cancer biomarkers: knowing the present and predicting the future. Future Oncol 1(1): 37-50.
- Polanski M, Anderson NL (2006) A list of candidate cancer biomarkers for targeted proteomics. Biomark Insights 2: 1-48.
- Willard HF, Angrist M, Ginsburg GS (2005) Genomic medicine: genetic variation and its impact on the future of health care. Phil Trans R Soc 360(1460): 1543-1550.
- Stathopoulou FT (2005) Genome and proteome in oncology. New York: Nova Science Publishers Inc.
- Kim YS, Maruvada P (2008) Frontiers in metabolomics for cancer research: proceedings of a National Cancer Institute workshop. Metabolomics 4: 105-113.
- Wulfkuhle JD, Liotta LA, Petricoin EF (2003) Proteomic applications for the early detection of cancer. Nat Rev Cancer 3(4): 267-275.
- Diamandis EP (2004) Mass spectrometry as a diagnostic and a cancer biomarker discovery tool: opportunities and potential limitations. Mol. Cell. Proteomics 3(4): 367-378.
- Righetti PG, Castagna A, Antonioli P, Cecconi D, Campostrini N, et al. (2005) Proteomic approaches for studying chemoresistance in cancer. Expert Rev Proteomics 2(2): 215-228.
- Chen CH (2008) Review of a current role of mass spectrometry for proteome research. Anal Chim Acta 624(1): 16-36.
- Kanmura S, Uto H, Kusumoto K, Ishida Y, Hasuike S, et al. (2007) Early diagnostic potential for hepatocellular carcinoma using the SELDI ProteinChip system. Hepatology 45(4): 948-956.
- Ishida Y, Yamashita K, Sasaki H, Takajou I, Kubuki Y, et al. (2008) Activation of complement system in adult T-cell leukemia (ATL) occurs mainly through lectin pathway; a serum proteomic approach using mass spectrometry. Cancer lett 271(1): 167-177.
- Shastry BS (2002) SNP alleles in human disease and evolution. J Hum Genet 47(11): 561-566.
- Wang XQ, Luk JM, Garcia-Barcelo M, Miao X, Leung PP, et al. (2006) Liver intestine-cadherin (CDH17) haplotype is associated with increased risk of hepatocellular carcinoma. Clin Cancer Res 12(7): 5248-5252.
- Esteller M, Corn PG, Baylin SB, Herman JG (2001) A gene hypermethylation profile of human cancer. Cancer Res 61(8): 3225-3229.
- Tischoff I, Tannapfe A (2008) DNA methylation in hepatocellular carcinoma. World J Gastroenterol 14: 1741-1748.
- Bowtell D, Sambrook J (2002) DNA microarrays: a molecular cloning manual. Cold Spring Harbor Laboratory Press, New York.
- Lee NH, Saeed AI (2007) Microarrays: an overview. Methods Mol Biol 353: 265-300.
- Sasaki H, Nishikata I, Shiraga T, Akamatsu E, Fukami T, et al. (2005) Overexpression of a cell adhesion molecule, TSLC1, as a possible molecular marker for acute-type adult T-cell leukemia. Blood 105(3): 1204-1213.
- Simon R (2003) Diagnostic and prognostic prediction using gene expression profiles in high-dimensional microarray data. Br J Cancer 89(9): 1599-1604.
- Brennan DJ, Kelly C, Rexhepaj E, Dervan PA, Duffy MJ, et al. (2007) Contribution of DNA and tissue microarray technology to the identification and validation of biomarkers and personalised medicine in breast cancer. Cancer Genomics Proteomics 4(3): 121-134.
- Sotiriou C, Piccart MJ (2007) Taking gene-expression profiling to the clinic: when will molecular signatures become relevant to patient care? Nat Rev Cancer 7(7): 545-553.
- Harrigan GG, Goodacre R (2003) Metabolic profiling: its role in biomarker discovery and gene function analysis. Boston: Kluwer Academic Publishers.
- Tomita M, Nishioka T (2005) Metabolomics: the frontier systems biology. New York, Springer.
- Griffin JL, Nicholls AW (2006) Metabolomics as a functional genomic tool for understanding lipid dysfunction in diabetes, obesity and related disorders. Parmacogenomics 7(7): 1095-1107.
- Fiehn O, Kopka J, Dörmann P, Altmann T, Trethewey RN (2000) Metabolite profiling for plant functional genomics. Nat Biotechnol 18(11): 1157-1161.
- Plumb R, Granger J, Stumpf C, Wilson ID, Evans JA, et al. (2003) Metabonomic analysis of mouse urine by liquid-chromatography-time of flight mass spectrometry (LC-TOFMS): detection of strain, diurnal and gender differences. Analyst 128: 819-823.
- Soga T, Baran R, Suematsu M, Ueno Y, Ikeda S, et al. (2006) Differential metabolomics reveals ophthalmic acid as an oxidative stress biomarker indicating hepatic glutathione consumption. J Biol Chem 281(24): 16768-16776.
- Spraul M, Freund AS, Nast RE, Withers RS, Maas WE, et al. (2003) Advancing NMR sensitivity for LC-NMR-MS using a cryoflow probe: application to the analysis of acetaminophen metabolites in urine. Anal Chem 75(6): 1536-1541.
- Yang Z (2006) Online hyphenated liquid chromatography-nuclear magnetic resonance spectroscopy-mass spectrometry for drug metabolite and nature product analysis. J Pharm Biomed Anal 40(3): 516-527.
- Schnackenberg LK, Beger RD (2006) Monitoring the health to disease continuum with global metabolic profiling and systems biology. Pharmacogenomics 7(7): 1077-1086.
- Kaddurah-Daouk R, Kristal BS, Weinshilboum RM (2008) Metabolomics: a global biochemical approach to drug response and disease. Annu Rev Pharmacol Toxicol 48: 653-683.
- Kind T, Tolstikov V, Fiehn O, Weiss RH (2007) A comprehensive urinary metabolomic approach for identifying kidney cancer. Anal Biochem 363(2): 185-195.
- Mitani Y, Lezhava A, Kawai Y, Kikuchi T, Oguchi-Katayama A (2007) Rapid SNP diagnostics using asymmetric isothermal amplification and a new mismatch-suppression technology. Nat Methods 4(3): 257-262.
- Kuzuhara T, Kise D, Shirakawa Y, Sasada K, Suganuma M, et al. (2008) Generation of mouse monoclonal antibody against (-)-epigallocatechin gallate. Biol Pharm Bull 31(5): 816-819.
- Grossklaus DJ, Smith JA, Shappell SB, Coffey CS, Chang SS, et al. (2002) The free/total prostate-specific antigen ratio (%fPSA) is the best predictor of tumor involvement in the radical prostatectomy specimen among men with an elevated PSA. Urol Oncol 7(5): 195-198.
- Whitehouse C, Solomon E (2003) Current status of the molecular characterization of the ovarian cancer antigen CA125 and implications for its use in clinical screening. Gynecol Oncol 88: S152-S157.
- Irmak S, Tilki D, Heukeshoven J, Oliveira-Ferrer L, Friedrich M, et al. (2005) Stage-dependent increase of orosomucoid and zinc-alpha2-glycoprotein in urinary bladder cancer. Proteomics 5(16): 4296-4304.
- Tirumalai RS, Chan KC, Prieto DA, Issaq HJ, Conrads TP, et al. (2003) Characterization of the low molecular weight human serum proteome. Mol Cell Proteomics 2(10): 1096-1103.
- Hortin GL (2006) The MALDI-TOF mass spectrometric view of the plasma proteome and peptidome. Clin Chem 52(7): 1223-1237.
- Link AJ (1999) 2-D proteome analysis protocols. Methods Mol Biol 112, Humana Press: Totowa.
- Kinter M, Sherman NE (2000) Protein sequencing and identification using tandem mass spectrometry. Wiley Interscience, New York, USA.
- Tonge R, Shaw J, Middleton B, Rowlinson R, Rayner S, et al. (2001) Validation and development of fluorescence two-dimensional differential gel electrophoresis proteomics technology. Proteomics 1(3): 377-396.
- Marouga R, David S, Hawkins E (2005) The development of the DIGE system: 2D fluorescence difference gel analysis technology. Anal Bioanal Chem 382(3): 669-678.
- Tang N, Tornatore P, Weinberger SR (2004) Current developments in SELDI affinity technology. Mass Spectrom Rev 23(1): 34-44.
- Seibert V, Ebert MP, Buschmann T (2005) Advances in clinical cancer proteomics: SELDI-TOF-mass spectrometry and biomarker discovery. Brief Funct Genomic Proteomic 4(1): 16-26.
- Ketterlinus R, Hsieh SY, Teng SH, Lee H, Pusch W (2005) Fishing for biomarkers: analyzing mass spectrometry data with the new ClinProTools software. Biotechniques 38(36): 37-40.
- Solassol J, Jacot W, Lhermitte L, Boulle N, Maudelonde T, et al. (2006) Clinical proteomics and mass spectrometry profiling for cancer detection. Expert Rev Proteomics 3(3): 311-320.
- McDonald WH, Yates JR 3rd (2002) Shotgun proteomics and biomarker discovery. Dis Markers 18(2): 99-105.
- Wu CC, MacCoss MJ (2002) Shotgun proteomics: tools for the analysis of complex biological systems. Curr Opin Mol Ther 4(3): 242-250.
- Motoyama A, Yates JR 3rd (2008) Multidimensional LC separations in shotgun proteomics. Anal Chem 80(19): 7183-7193.
- Tanaka K, Waki H, Ido Y, Akita S, Yoshida, et al. (1988) Protein and polymer analyses up to m/z 100 000 by laser ionization time-of-flight mass spectrometry. Rapid Commun Mass Spectrom 2(8): 151-153.
- Karas M, Hillenkamp F (1988) Laser desorption ionization of proteins with molecular masses exceeding 10,000 daltons. Anal Chem 60(20): 2299-3201.
- Whitehouse CM, Dreyer RN, Yamashita M, Fenn JB (1985) Electrospray interface for liquid chromatographs and mass spectrometers. Anal Chem 57(3): 675-679.
- Fen JB, Mann M, Meng CK, Wong SF, Whitehouse CM (1989) Electrospray ionization for mass spectrometry of large biomolecules. Science 246(4926): 64-71.
- Pappin DJ, Hojrup P, Bleasby AJ (1993) Rapid identification of proteins by peptide-mass fingerprinting. Curr Biol 3(6): 327-332.
- Henzel WJ, Billeci TM, Stults JT, Wong SC, Grimley C, et al. (1993) Identifying proteins from two-dimensional gels by molecular mass searching of peptide fragments in protein sequence databases. Proc Natl Acad Sci 90(11): 5011-5015.
- Mann M, Hojrup P, Roepstorff P (1993) Use of mass spectrometric molecular weight information to identify proteins in sequence databases. Biol Mass Spectrom 22(6): 338-345.
- James P, Quadroni M, Carafoli E, Gonnet G (1993) Protein identification by mass profile fingerprinting. Biochem Biophys Res Commun 195(1): 58-64.
- Yates JR 3rd, Speicher S, Griffin PR, Hunkapiller T (1993) Peptide mass maps: a highly informative approach to protein identification. Anal Biochem 214(2): 397-408.
- Siuzdak G (2003) The expanding role of mass spectrometry in biotechnology. MCC Press, San Diego, CA, USA.
- Shevchenko A, Jensen ON, Podtelejnikov AV, Sagliocco F, Wilm M, et al. (1996) Linking genome and proteome by mass spectrometry: large-scale identification of yeast proteins from two dimensional gels. Natl Acad Sci 93(25): 14440-14445.
- Johnson RS, Davis MT, Taylor JA, Patterson SD (2005) Informatics for protein identification by mass spectrometry. Methods 35(3): 223-236.
- Shadforth I, Crowther D, Bessant C (2005) Protein and peptide identification algorithms using MS for use in high throughput, automated pipelines. Proteomics 5(16): 4082-4095.
- Bast RC Jr, Klug TL, St John E, Jenison E, Niloff JM, et al. (1983) A radioimmunoassay using a monoclonal antibody to monitor the course of epithelial ovarian cancer. N Engl J Med 309(15): 883-887.
- Bast RC Jr, Urban N, Shridhar V, Smith D, Zhang Z et al. (2002) Early detection of ovarian cancer: promise and reality. Cancer Treat Res 107: 61-97.
- Cohen LS, Escobar PF, Scharm C, Glimco B, Fishman DA (2001) Three-dimensional power Doppler ultrasound improves the diagnostic accuracy for ovarian cancer prediction. Gynecol Oncol 82(1): 40-48.
- Li J, Orlandi R, White CN, Rosenzweig J, Zhao J, et al. (2005) Independent validation of candidate breast cancer serum biomarkers identified by mass spectrometry. Clin Chem 51(12): 2229-2235.
- Goncalves A, Esterni B, Bertucci F, Sauvan R, Chabannon C, et al. (2006) Postoperative serum proteomic profiles may predict metastatic relapse in high-risk primary breast cancer patients receiving adjuvant chemotherapy. Oncogene 25(7): 981-989.
- Liu L, Wang J, Liu B, Dai S, Wang X, et al. (2008) Serum levels of variants of transthyretin down-regulation in cholangiocarcinoma. J Cell Biochem 104(3): 745-755.
- Ward DG, Suggett N, Cheng Y, Wei W, Johnson H, et al. (2006) Identification of serum biomarkers for colon cancer by proteomic analysis. Br J Cancer 94(12): 1898-1905.
- Albrethsen J, Bogebo R, Gammeltoft S, Olsen J, Winther B, et al. (2005) Upregulated expression of human neutrophil peptides 1, 2, and 3 (HNP 1-3) in colon cancer serum and tumours: a biomarker study. BMC Cancer 5: 8.
- Engwegen JYMN, Helgason HH, Cats A, Harris N, Bonfrer JMG, et al. (2006) Identification of serum proteins discriminating colorectal cancer patients and healthy controls using surface-enhanced laser desorption ionisation-time of flight mass spectrometry. World J Gastroenterol 12(10): 1536-1544.
- Su Y, Shen J, Qian H, Ma H, Ji J, et al. (2006) Diagnosis of gastric cancer using decision tree classification of mass spectral data. Cancer Sci 98(1): 37-43.
- Ebert MP, Niemeyer D, Deininger SO, Wex T, Knippig C, et al. (2006) Identification and confirmation of increased fibrinopeptide a serum protein levels in gastric cancer sera by magnet bead assisted MALDI-TOF mass spectrometry. J Proteome Res 5(9): 2152-2158.
- Liu W, Liu B, Xin L, Zhang Y, Chen X, et al. (2007) Down-regulated expression of complement factor I: a potential suppressive protein for gastric cancer identified by serum proteome analysis. Clin Chim Acta 377(1-2): 119-126.
- Ward DG, Cheng Y, N’Kontchou G, Thar TT, Barget N, et al. (2006) Preclinical and post-treatment changes in the HCC-associated serum proteome. Br J Cancer 95(10): 1379-1383.
- Lee IN, Chen CH, Sheu JC, Lee HS, Huang GT, et al. (2006) Identification of complement C3a as a candidate biomarker in human chronic hepatitis C and HCV-related hepatocellular carcinoma using a proteomics approach. Proteomics 6(9): 2865-2873.
- Feng JT, Liu YK, Song HY, Dai Z, Qin LX, et al. (2005) Heat-shock protein 27: a potential biomarker for hepatocellular carcinoma identified by serum proteome analysis. Proteomics 5(17): 4581-4588.
- He M, Qin J, Zhai R, Wei X, Wang Q, et al. (2008) Detection and identificaiton of NAP-2 as a biomarker in hpatitis B-related hepatocellular carcinoma by proteomic approach. Proteome Sci 6(10).
- Zinkin NT, Grall F, Bhaskar K, Otu HH, Spentzos D, et al. (2008) Serum proteomics and biomarkers in hepatocellular carcinoma and chronic liver disease. Clin Cancer Res 14(2): 470-477.
- He QY, Zhu R, Lei T, Ng MYM, Luk JM, et al. (2008) Toward the proteomic identification of biomarkers for the prediction of HBV related hepatocellular carcinoma. J Cell Biochem 103(3): 740-752.
- Orvisky E, Drake SK, Martin BM, Abdel-Hamid M, Ressom HW, et al. (2006) Enrichment of low molecular weight fraction of serum for MS analysis of peptides associated with hepatocellular carcinoma. Proteomics 6(9): 2895-2902.
- Dowling P, O’Driscoll L, Meleady P, Henry M, Roy S, et al. (2007) 2-D difference gel electrophoresis of the lung squamous cell carcinoma versus normal sera demonstrates consistent alterations in the levels of ten specific proteins. Electrophoresis 28(23): 4302-4310.
- Okano T, Kondo T, Kakisaka T, Fujii K, Yamada M, et al. (2006) Plasma proteomics of lung cancer by a linkage of multi-dimensional liquid chromatography and two-dimensional difference gel electrophoresis. Proteomics 6(13): 3938-3948.
- Doustjalali SR, Yusof R, Govindasamy GK, Bustam AZ, Pillay B, et al. (2006) Patients with nasopharyngeal carcinoma demonstrate enhanced serum and tissue ceruloplasmin expression. J Med Invest 53(1-2): 20-28.
- Liao Q, Zhao L, Chen X, Deng Y, Ding Y (2008) Serum proteome analysis for profiling protein markers associated with carcinogenesis and lymph node metastasis in nasopharyngeal carcinoma. Clin Exp Metastasis 25(4): 465-476.
- Cheng AJ, Chen LC, Chien KY, Chen YJ, Chang JT et al. (2005) Oral cancer plasma tumor marker identified with bead-based affinity-fractionated proteomic technology. Clin Chem 51(12): 2236-2244.
- Jackson D, Craven RA, Hutson RC, Graze I, Lueth P, et al. (2007) Proteomic profiling identifies afamin as a potential biomarker for ovarian cancer. Clin Cancer Res 13(24): 7370-7379.
- Ahmed N, Barker G, Oliva KT, Hoffmann P, Riley C, et al. (2004) Proteomic-based identification of haptoglobin-1 precursor as a novel circulating biomarker of ovarian cancer. Br J Cancer 91: 129-140.
- Ahmed N, Oliva KT, Barker G, Hoffmann P, Reeve S, et al. (2005) Proteomic tracking of serum protein isoforms as screening biomarkers of ovarian cancer. Proteomics 5(17): 4625-4636.
- Moshkovskii SA, Serebryakova MV, Kuteykin-Teplyakov KB, Tikhonova OV, Goufman EI, et al. (2005) Ovarian cancer marker of 11.7 kDa detected by proteomics is a serum amyloid A1. Proteomics 3790-3797.
- Woong-Shick A, Sung-Pil P, Su-Mi B, Joon-Mo L, Sung-Eun N, Identification of hemoglobin-α and -β subunits as potential serum biomarkers for the diagnosis and prognosis of ovarian cancer. Cancer Sci 96(3): 197-201.
- Kozak KR, Su F, Whitelegge JP, Faull K, Reddy S, et al. (2005) Farias-Eisner, R. Characterization of serum biomarkers for detection of early-stage ovarian cancer. Proteomics 5(17): 4589-4596.
- Kakisaka T, Kondo T, Okano T, Fujii K, Honda K, et al. (2007) Plasma proteomics of pancreatic cancer patients by multi-dimensional liquid chromatography and two-dimensional difference gel electrophoresis (2D-DIGE): up-regulation of leucine-rich alpha-2-glycoprotein in pancreatic cancer. J Chromatogr B 852(1-2): 257-267.
- Bloomston M, Zhou JX, Rosemurgy AS, Frankel W, Muro-Cacho CA, et al. (2006) Fibrinogen γ overexpression in pancreatic cancer identified by large-scale proteomic analysis of serum samples. Cancer Res 66(5): 2592-2599.
- Malik G, Ward MD, Gupta SK, Trosset MW, Grizzle WE, et al. (2005) Serum levels of an isoform of apolipoprotein A2 as a potential marker for prostate cancer. Clin Cancer Res 11(3): 1073-1085.
- Tolson J, Bogumil R, Brunst E, Beck H, Elsner R, et al. (2004) Serum protein profiling by SELDI mass spectrometry: detection of multiple variants of serum amyloid alpha in renal cancer patients. Lab Invest 84(7): 845-856.
- Akiba J, Yano H, Ogasawara S, Higaki K, Kojiro M (2001) Expression and function of interleukin-8 in human hepatocellular carcinoma. Int J Oncol 18(2): 257-264.
- Aivado M, Spentzos D, Germing U, Alterovitz G, Meng XY, et al. (2007) Serum proteome profiling detects myelodysplastic syndromes and identifies CXC chemokine ligands 4 and 7 as markers for advanced disease. Proc Natl Acad Sci 104(4): 1307-1312.
- Ren Y, Poon RT, Tsui HT, Chen WH, Li Z, et al. (2003) Interleukin-8 serum levels in patients with hepatocellular carcinoma: correlations with clinicopathological features and prognosis. Clin Cancer Res 9: 5996-6001.
- Yang M, Chen T, Han C, Li N, Wan T, et al. (2004) Rab7b, a novel lysosome-associated small GTPase, is involved in monocytic differentiation of human acute promyelocytic leukemia cells. Biochem Biophys Res Commun 318(3): 792-799.
- Benson MD, Cohen AS (1979) Serum amyloid A protein in amyloidosis, rheumatic and neoplastic diseases. Arthritis Rheum 22(1): 36-42.
- Uhlar CM, Whitehead AS (1999) Serum amyloid A, the major vertebrate acute-phase reactant. Eur J Biochem 265(2): 501-523.
- Cho WC, Yip TT, Yip C, Yip V, Thulasiraman V, et al. (2004) Identification of serum amyloid a protein as a potentially useful biomarker to monitor relapse of nasopharyngeal cancer by serum proteomic profiling. Clin Cancer Res 10(1): 43-52.
- Storm FK, Gilchrist KW Warner TF, Mahvi DM (1995) Distribution of HSP-27 and HER-2/neu in in situ and invasive ductal breast carcinomas. Ann Surg Oncol 2(1): 43-48.
- Assimakopoulou M, Sotiropoulou-Bonikou G, Maraziotis T, Varakis I (1997) Prognostic significance of HSP-27 in astrocytic brain tumors: an immunohistochemical study. Anticancer Res 17(4A): 2677-2682.
- Yu Y, Chen S, Wang LS, Chen WL, Guo WJ, et al. (2015) Prediction of pancreatic cancer by serum biomarkers using surface-enhanced laser desorption/ionization-based decision tree classification. Oncology 68(1): 79-86.
- Wei YS, Zheng YH, Liang WB, Zhang JZ, Yang ZH, et al. (2008) Identification of serum biomarkers for nasopharyngeal carcinoma by proteomic analysis. Cancer 112(3): 544-551.
- Wang J, Zhang X, Ge X, Guo H, Xiong G, et al. (2008) Proteomic studies of early-stage and advanced ovarian cancer patients. Gynecol Oncol 111(1): 111-119.
- Yoshida H, Shiratori Y, Moriyama M, Arakawa Y, Ide T, et al. (1999) Interferon therapy reduces the risk for hepatocellular carcinoma: national surveillance program of cirrhotic and noncirrhotic patients with chronic hepatitis C in Japan. IHIT Study Group. Inhibition of Hepatocarcinogenesis by Interferon Therapy. Ann Intern Med 131(3): 174-181.
- Heathcote EJ (2004) Prevention of hepatitis C virus-related hepatocellular carcinoma. Gastroenterology 127: S294-302.
- Uto H, Hayashi K, Kusumoto K, Hasuike S, Nagata K, et al. (2006) Spontaneous elimination of hepatitis C virus RNA in individuals with persistent infection in a hyperendemic area of Japan. Hepatol Res 34(1): 28-34.
- Kusumoto K, Uto H, Hayashi K, Takahama Y, Nakao H, et al. (2006) Interleukin-10 or tumor necrosis factor-α polymorphisms and the natural course of hepatitis C virus infection in a hyperendemic area of Japan. Cytokine 34(1-2): 24-31.
- Uto H, Kurogi J, Takahama Y, Kusumoto K, Hayashi K, et al. (2007) Alanine aminotransferase flare-up in hepatitis C virus carriers with persistently normal alanine aminotransferase levels in a hyperendemic area of Japan. Gastroenterol 42(8): 673-680.
- Tang HY, Ali-Khan N, Echan LA, Levenkova N, Rux JJ, et al. (2005) A novel four-dimensional strategy combining protein and peptide separation methods enables detection of low-abundance proteins in human plasma and serum proteomes. Proteomics 5(13): 3329-3342.
- Fujii K, Nakano T, Kawamura T, Usui F, Bando Y, et al. (2004) Multidimensional protein profiling technology and its application to human plasma proteome. J Proteome Res 3(4): 712-718.
- Welsh JB, Sapinoso LM, Kern SG, Brown DA, Liu T, et al. (2003) Large-scale delineation of secreted protein biomarkers overexpressed in cancer tissue and serum. Proc Natl Acad Sci 100(6): 3410-3415.
- Roessler M, Rollinger W, Palme S, Hagmann ML, Berndt P, et al. (2005) Identification of nicotinamide N-methyltransferase as a novel serum tumor marker for colorectal cancer. Clin Cancer Res 11(18): 6550-6557.
- Hwang TL, Liang Y, Chien KY, Yu JS (2006) Overexpression and elevated serum levels of phosphoglycerate kinase 1 in pancreatic ductal adenocarcinoma. Proteomics 6(7): 2259-2272.
- Law SK, Dodds AW (1997) The internal thioester and the covalent binding properties of the complement proteins C3 and C4. Protein Sci 6(2): 263-274.
- Rother K, Till GO, Hansch GM (1998) The complement system. second edn., Springer press, Berlin, Germany.
- Sim RB, Laich A (2000) Innate immunity: serine proteases of the complement system. Biochem Soc Trans 28(5): 545-550.
- Veenstra TD, Prieto DA, Conrads TP (2004) Proteomic patterns for early cancer detection. Drug Discov Today 9(20): 889-997.
- Crowther JR (2001) The ELISA guidebook: methods in molecular biology. Totawa, Humana Press, USA.
- Nielsen UB, Geierstanger BH (2004) Multiplexed sandwich assays in microarray format. J Immunol Methods 290(1-2): 107-120.






























