Promoter Hypermethylation of Apoptotic genes in Oral Squamous Cell Carcinoma
Neha Pal1, Shalini Rathore1, Priya Kaushik1, Swati Tyagi1, Anshuman Kumar2 and Abhimanyu Kumar Jha1*
1Department of Biotechnology, Faculty of Life Sciences, Institute of Applied Medicines and Research, Ghaziabad, Uttar Pradesh, India
2Director, Surgical Oncology, Dharamshila Narayana Super speciality Hospital, Delhi, India
Submission: April 23, 2019; Published: May 23, 2019
*Corresponding Address: Abhimanyu Kumar Jha, Department of Biotechnology, Faculty of Life Sciences, Institute of Applied Medicines and Research, Ghaziabad, Uttar Pradesh, India
How to cite this article: Neha Pal, Shalini Rathore, Priya Kaushik, Abhimanyu Kumar Jha, et al. Promoter Hypermethylation of Apoptotic genes in Oral Squamous Cell Carcinoma. Canc Therapy & Oncol Int J. 2019; 13(5): 555872. DOI:10.19080/CTOIJ.2019.13.555872
Abstract
Cancer is one of those diseases whose treatment requires more rapid and potent response and epigenetic modifications have emerged one amongst the cancer hallmarks, commutating the concept of malignant pathologies as being singlely genetic-based conditions. Oral Cancer is the type of malignancy that is found in any area of mouth and nearby regions. The epigenetic landscape is accountable for traditional development however additionally for the non uniformity among tissues in terms of gene expression patterns. Dysregulation in these mechanisms has been related to illness stage, and raised attention is currently granted to cancer so as to require advantage of those modifications in terms of novel therapeutic methods or diagnosis/prognosis tools. Epigenetics has revealed a diagnostic tool that is tumour markers which is also known as biomarkers. These biomarkers are either methylated at their CpG islands called as DNA hypermethylation or another modification could be possibly observed that would help us to identify the neoplasm for its medication because it is usually symptomless at its beginning stages. This review encompasses the different apoptotic genes that have been found as a biomarker for diagnosing the OSCC (Oral Squamous Cell Carcinoma) and how they are related to epigenetics. This can hold an immense importance in the treatment of cancer and could help in discovering novel therapeutic ways in its treatment.
Keywords: Oral Cancer; Epigenetics; DNA Hypermethylation
Introduction
Cancer stands among the foremost advanced human diseases with multiple levels of regulation. Despite the numerous progress that has been created in basic and clinical analysis, the incidence associated mortality rates related to malignant pathologies are still at high levels with an expected raised rate within the next fifteen years Irimie et al. [1]. The sixth most typical malignancy that has been found is Head and Neck cancer, wherever carcinoma represents the foremost frequent subtype inside this spectrum Irimie et al. [2], Hema et al. [3]. Also, Oral Squamous Cell Carcinoma (OSCC) accounts for roughly ninetieth of the oral subtype, being far and away the foremost common sort of malignancy inside the mouth Parkin et al. [4], Stewart et al. [5]. Apoptosis is that one pathway that could lead the elimination of oncogenic cells. Apoptosis can be explained as a process in which natural death of cells is initiated to maintain the cellular homeostasis. A variety of changes are done in the structure of cells by apoptotic proteins that eliminate the cell out of body. In this mechanism events occurs which result into formation of cascades that encounters the already anteceding cells leading them to demise, also known as “programmed cell death”, Elmore [6], Hervouet et al. [7]. The initiation andmaturation of oral carcinoma caused by total of genetic changes combined with environmental risk factors (special mention to alcohol and tobacco use however various viral infectious agents and persistent inflammations) cause changes within the pursuit of cancer causing genes i.e., oncogenes and tumour suppressor genes Perez et al. [8], Irimie et al. [9].
The extent of complexness behind the unusual expression and performance of these genes is extraordinarily high, wherever epigenetic mechanisms are standing as another section of regulation besides the changes within the DNA sequence, that embrace mutations, deletions, and amplifications Lingen et al. [10]. But it has been observed that this mechanism suffers with problems or gets defected in various human diseases such as in cancer. As cancer is one of the major concerns of today’s world and it has been observed that the uncontrolled growth of cells resulting tumour escapes the apoptosis process. The various genes that plays role in apoptosis are silenced by a process named DNA hypermethylation. There are also various other epigenetic modifications that have their crucial role in oral cancer such as DNA hypomethylation, histone modification, micro RNA’s and many other.
Apoptosis
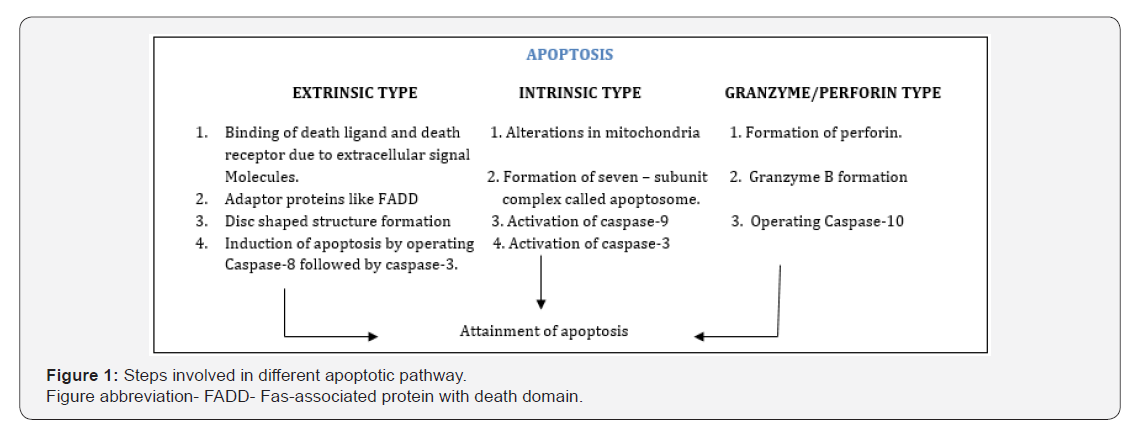
Apoptosis is highly organized process that involves list of events for eliminating the cell out of body. The major two different types of mechanisms are as following:
i. Extrinsic pathway or death receptor pathway
ii. Intrinsic pathway or mitochondrial pathway
The procedure begins by triggering the caspase-3 followedby DNA fragmentation, degradation of cytoskeletal and nuclear proteins, formation of apoptotic bodies, expression of ligands for phagocytic cell receptors and finally uptake by phagocytic cells in extrinsic, intrinsic and granzyme B pathways. Grainzyme pathway does not depend upon degradation of capsase as it initiates parallel Martinvalet et al. [11], deflections in apoptotic process has been observed due to tumourgensis (cancer) due to which various pursuits can’t carried out properly as deactivation of propapoptotic genes and translation of antiapoptotic genes occurs in high quantity promoting uncontrollable growth of oncologic cells Evan et al. [12] (Figure 1).
Oral Cancer
Oral cancer is a form of cancer that is found in the middle of vermillion border of lips and the junction of the hard and soft palates or the posterior of the tongue. It is a type of major malignancy found over the world. Although it was reported that oral squamous cell carcinoma (OSCC) is the most found subclassified malignancy within the oral cancer. The major causes of oral cancer can be either alcohol consumption or tobacco use or it could be atrocious inflammation in the oral mucousal cavity, there have been findings that suggests that global hypomethylation is connected to smoking Datta et al. [13], Esteller [14], Blot et al. [15] while drinking has been associated with CpG hypermethylation according to clinical studies effectuated on patient samples and also in murine models for oral malignancies. It has been estimated that globally the rate of death due to oral squamous cell carcinoma (OSCC) is more than 1,00000 and 20 per 100000 people of India are over blown by oral cancer which indicates about 30% of all types of cancer Sankaranarayanan et al. [16], Parkin et al. [17]. According to the Globocan (2018), 572,034 (3.2%) new cases for oesophageal are found out of 18,078,957 i.e. total 508,585 (5.3%) deaths in 2018 out of total 9,555,027.
Correlation between Oral cancer and Wnt Signalling
Wnt-1 was identified as proto-oncogene protein present on gene locus 12q13.12. Earlier mouse mammary tumour virus(MMTV) triggered the mammary oncogene which was then appeared as Wnt-1. The decline in pursuit of GSK-3 β and rise in β catenin pool, regulating cadherin catenin complex creation and cell-to-cell adhesion, and aiding cells from apoptosis by operating tyrosine kinase receptors Brown et al. [18]. Due to the rise in transcription elements from head and neck OSCC line, the Wnt1 genes participate sometimes. The discovery of anti- Wnt1 antibodies declined the cyclin D1 and β-catenin proteins formation in OSCC line by decreasing the Wnt1/frizzleddependent transcription elements (TCF/LEF) Muzio et al. [19], Rhee et al. [20]. Initiation of apoptosis occurs due to choked Wnt-1 signalling. The mechanism which involves adherence molecules, elements participating in transcription and efficacious oncogenes helps to decipher the relation between oral cancer and Wnt1 signalling pathway.
Epigenetics
Epigenetics can be explained as changes in chromosome and genes that affects gene activity and expression. Epigenetics in oral cancer has been proving as a revolutionary step in diagnosis and treatment of cancer. It includes DNA methylation, histone modification and post-transcriptional gene down-regulation by microRNAs Santoro et al. [21]. These changes are not like genetic changes; they alter the DNA and histone by various mechanisms. Out of all these DNA methylations have been a hot topic for scientists in OSCC, which could suitably prove as a biomarker for the tumour detection.
DNA Methylation
In this process addition of methyl groups are done on specific sites known as CpG islands that contain plenty of CpG. This could be done by using enzyme DNA Methyltransferase. Several forms of DNMT’s are present such as DNMT1, DNMT2, and DNMT3. The methyl group is transferred to the carbon on 5th position of cytosine in CpG motifs which silences the gene transcription. DNA Methylation is majorly of two types - one is DNA hypermethylation that is performed by DNMT 1, DNMT 3a and DNMT 3b. Another is global DNA hypomethylation. DNA hypomethylation has been enlisted in promoting the growth of cancer cells by various mode of actions like generation of chromosomal instability, resuscitation of transposable elements, and dissipation of imprinting Esteller [22].
Histone Modification
Histone proteins are susceptible to totally different modifications that include ubiquitylation, sumoylation, methylation, acylation and additionally phosphorylation. These modifications occur majorly at the N-terminal tails, influencing gene transcription and crucial signalling pathways Kouzarides[23]. Histone modification is alternate to DNA methylation as it can explicitly not only mute the gene expression, but it also stimulates the transcription.
Hypermethylated Apoptotic Genes
CDH1
CDH1 which is also known as E-cadherin is found on chromosome 16q22.1 and supports cell–cell adhesion by forming adherent junction in epithelial cells for initiating and preserving intercellular adhesion, cell polarity, intercellular signalling and structure of tissue. There have many reports that emphasize on a fact that CDH1 is being expressed in several types of cancer. And if the CDH1 is not present, it is adamantly indicating that there might be changes in cell’s function and motility. It was also observed that antagonistic gesture of OSCC has its connection with the lesser expression of CDH1 (E-cadherin). It was identified that the chances of tumour avalanche which is a known property of cancerous cells “metastasis” could happen if mutation occurs or if gene expresses less. Promoter methylation of CDH1 is found to be 17–85% in oral-cavity tumours Supic et al. [24], Chang et al. [25], Kordi et al. [26], Shaw et al. [27] (Table 1).
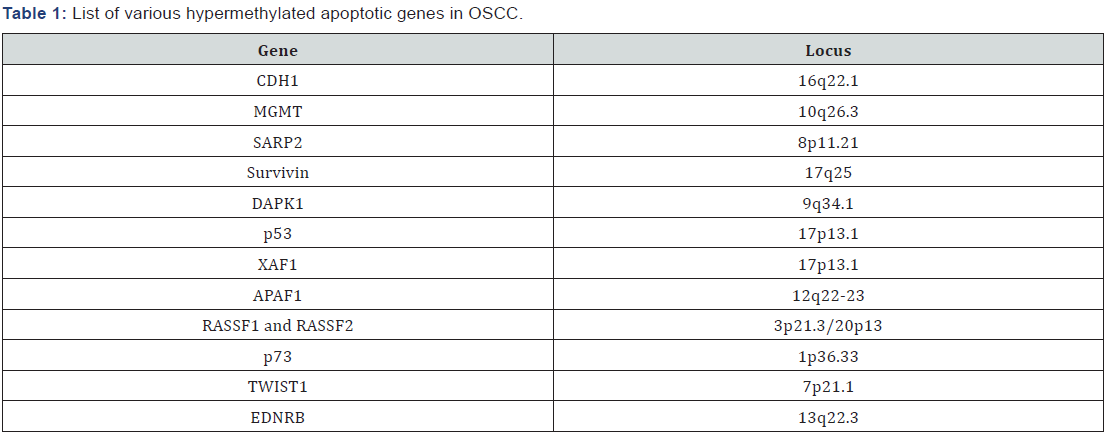
CDH1: E- cadherin 1; MGMT: O6- methylguanine- DNAmethyltransferase; SARP2: Secreted apoptosis- related macromolecule 2; DAPK1: Death associated protein kinase 1; p53 (Tp53): Tumor suppressor protein having 53 kiloDalton weight; XAF1: XIAP ( X-linked inhibitor of apoptosis protein ) associated factor 1; APAF1: Apoptotic protease activating factor 1; RASSF1 and RASSF2: Ras association domain family proteins; p73: Tumor suppressor protein having 73 Kilodalton weight; TWIST1: Twist- related protein 1; EDNRB- Endothelin Receptor Type B.
MGMT
MGMT stands for O6 - methylguanine – DNA – methyltransferase present on gene locus 10q26.3. It is a DNA repair enzyme that abolishes O6- guanine- DNA adducts generated by alkylating agents MGMT is a detoxifying agent of DNA adducts. It is important in preventing alkylation and, thus,could be predictive of chemosensitivity. It has been examined that 44 of many tumour types, MGMT was methylated in up to 38% of primary colon cancers, head and neck cancers. It has been also observed from few sources that 25–52% of primary oral squamous cell carcinomas had alkyl MGMT. The promoter hypermethylation is the reason due to which MGMT drops its activity and malignancy gets activated. Epigenetic silencing of MGMT has been associated with OSCCs where tobacco exposure and betel quid chewing are suspected to be etiological factors Huang et al. [28].
SARP2
Secreted Apoptosis-Related macromolecule (SARP) family’s square measure thought-about to counteract the oncogenicWnt signalling pathway and inactivation of this factor could aid cancer development and progression. SARP proteins, additionally named sFRP (for Secreted Frizzled-Related Protein), exert differential effects on osteoblastic differentiation of mouse mesenchymal cells and cellular programmed cell death of mouse osteoblasts in vitro Cho et al. [29]. SARP2 methylation was determined in each benign and malignant tumour and no applied mathematics link can be found between SARP2 methylation and illness progression.
Survivin
Survivin is another important factor that modifies apoptotic processes in oral cancer and in other different types of cancer found in our body by modulating cell cycle, gratifying the overpowering of checkpoints. It has been recorded that the transcription of this gene rises in various tumour samples like in those that have been taken from the patients suffering from oral cancer and it has been found that it was also connected with clinical as well as pathological features; in addition this gene’s expression is absent in normal tissue Muzio et al. [30], Khan et al. [31]. Single nucleotide polymorphism was also correlated with the increased expression of survivn gene in the basic cancer research. Although there have been facts that suggest that hypomethylation Gasche et al. [32], Chen et al. [33], Hsue et al. [34] is found in this gene contradictory to normal samples where hypermethylation averts the overexpression.
DAPK1
DAPK1 stands for death associated protein kinase 1 involved in OSCC as one of the hypermethylated apoptotic gene that could serve as a biomarker in future. The location is found on gene locus 9q34.1. It is responsible for operating serine/threonine kinase pathway that initiates apoptosis by calcium/calmodulin (i.e. proapoptotic) Kulkarni et al. [35], Kato et al. [36], Ha et al. [37]. It was found from few reports that 18-27% of promoter hypermethylation was observed in DAP Hasegawa et al. [38], Sanchez et al. [39], Rosas et al. [40]. Both Intrinsic and Extrinsic type of apoptosis can be performed by DAPK. In various cancers the hypermethylation of DAPK purely indicates its correlation with disease. Methylation is responsible for the reduced gene expression of DAPK gene and the blocked apoptosis process indicates that the initiation of tumour begins and other events at molecular level also described Mittag et al. [41].
P53
P53 (TP53) is a tumour suppressor protein located on 17p13.1. This gene plays a very vital role in several processes such as cellular differentiation, cell cycle progression, DNA repair and apoptosis. If any unfavourable changes occur either due to any external or internal factors it results into rise of p53 expression that ultimately chokes cellular functions, permitting DNA repair. It has been observed that in most types of cancers, the mutation in p53 is found involving OSCC. The methylation status of p53 is approx. 25% to 69% Levine et al. [42], Boyle etal. [43], Caamano et al. [44], Baral [45]. Epigenetic changes also led the reduced activity of p53. Therefore, epigenetic silencing of p53 protein (non mutated) due to HPV virus, mainly involving HPV 16 and 18 found in OSCC like in few laryngeal cancers.
RASSF1 and RASSF2
The Ras Association domain family proteins included in the Ras/PI3K/AKT pathways. RASSF1 is located on 3p21.3 and RASSF2 is located on 20p13. It was displayed by radiotherapy treatment in patients (50%) that Ras/PI3K/AKT mechanisms were triggered due to promoter methylation related to RASSF1A/RASSF2A gene silencing Huang et al. [46]. It has been a promising challenge for researcher’s that could help in discovery of biomarkers by identifying the methylation patterns in the tumour invasion (metastasis). 12-38% of methylation is found in RASSF1 and methylation status of RASSF2 is around 39% in no less than one RASSF2 gene Supic et al. [24], Taioli et al. [47], Huang et al. [46].
Conclusion
These studies have emerged as an effective tool for prognosis and diagnosis of such neoplasms. DNA Methylation involves a novel marker i.e. tumour marker (biomarker) [48-65]. There have been many genes that have been investigated as a biomarker few of which are described in this review. The methylation prevents the cancerous cells from apoptosis that causes invasion of these cells due to metastasis property possessed by cancerous cells. Therefore, if possibly reversing of such epigenetic modifications occurs promoting them to apoptosis then it would also help us to treat the cancer. These biomarkers are apoptotic proteins (genes) which get inactive due to methylation. In future these new therapies if adapted to treat patients by merging this present remedy seems to be more approachable and reactivation of apoptotic mechanism then would prove as very useful in the treatment of cancer.
References
- Irimie AI, Ciocan C, Gulei D, Mehterov N, Atanasov AG, et al. (2018) Current Insights into Oral Cancer Epigenetics. Int J Mol Sci 19(3).
- Irimie AI, Braicu C, Cojocneanu-Petric R, Berindan-Neagoe I, Campian RS (2015) Novel technologies for oral squamous carcinoma biomarkers in diagnostics and prognostics. Acta Odontol Scand 73(3): 161-168.
- Hema KN, Smitha T, Sheethal HS, Mirnalini SA (2017) Epigenetics in oral squamous cell carcinoma. J Oral Maxillofac Pathol 21(2): 252–259.
- Parkin DM, Bray F, Ferlay J, Pisani P (2005) Global cancer statistics, 2002. CA Cancer J Clin 55(2): 74-108.
- Stewart BW, Greim H, Shuker D, Kauppinen T (2003) Defence of iarc monographs. Lancet 361(9365): 1300.
- Elmore S (2007) Apoptosis: A review of programmed cell death. Toxicol Pathol 35(4): 495–516.
- Hervouet E, Cheray M, Vallette FM, Cartron PF (2013) DNA methylation and apoptosis resistance in cancer cells. Cell 2(3): 545-573.
- Perez-Sayans M, Somoza-Martin J M, Barros-Angueira F, Reboiras-Lopez, Gandara Rey JM, et al. (2009) Genetic and molecular alterations associated with oral squamous cell cancer (review). Oncol Rep 22(6): 1277–1282.
- Irimie AI, Braicu C, Pileczki V, Petrushev B, Soritau O, et al. (2016) Knocking down of p53 triggers apoptosis and autophagy, concomitantly with inhibition of migration on ssc-4 oral squamous carcinoma cells. Mol Cell Biochem 419(1-2): 75–82.
- Lingen MW, Pinto A, Mendes RA, Franchini R, Czerninski R, et al. (2011) Genetics/epigenetics of oral premalignancy: Current status and future research. Oral Dis 17(1): 7–22.
- Martinvalet D, Zhu P, Lieberman J (2005) Granzyme A induces caspase- independent mitochondrial damage, a required first step for apoptosis. Immunity 22(3): 355-370.
- Evan G, Vousden K (2001) Proliferation, cell cycle and apoptosis in cancer. Nature 411(6835): 342-348.
- Datta K, Saha RK, Chakrabarti RN (1997) A simple risk estimates study for oral cavity cancer: practical approach in Indian context. Journal of the Indian Medical Association 95(3): 70–71.
- Esteller M (2003) Cancer epigenetics: DNA methylation and chromatin alterations in human cancer. Adv Exp MedBiology 532: 39–49.
- Blot WJ, McLaughlin JK, Winn DM, Austin DF, Greenberg RS, et al. (1998) Smoking and drinking in relation to oral and pharyngeal cancer. Cancer res 48(11): 3282-3287.
- R Sankaranarayanan, K Ramadas, G Thomas, Muwonge R, Thara S, et al. (2005) Effect of screening on oral cancer mortality in Kerala, India: a clusterrandomised controlled trial. The Lancet 365(9475): 1927–1933.
- Parkin DM, Pisani P, Ferlay J (1990) Estimates of the worldwide incidence of 25 major cancers in 1990. Int J Cancer 80(6): 827–841.
- Brown A, Wildin R, Prendergast T, Varmus H (1986) A retrovirus vector expressing the putative mammary oncogene INT-L causes partial transformation of a mammary epithelial cell line. Cell 46(7): 1001-1009.
- Muzio LL (2001) A possible role for the Wnt-1 pathway in oral carcinogenesis. Crit Rev Oral Biol Med 12(2): 152-165.
- Rhee CS, Sen M, Lu D, Wu C, Leoni L (2002) Wnt and frizzled receptors as potential targets for immunotherapy in head and neck squamous cell carcinomas. Oncogene 21(43): 6598-6605.
- Santoro A, Pannone G, Papagerakis S, Serpico R, Guida A, et al. (2012) Epigenetic Profiling of Oral Cancer. Intech open p. 32.
- Esteller M (2008) Epigenetics in Cancer. N Engl J Med 358: 1148-1159.
- Kouzarides T (2007) Chromatin modifications and their function. Cell 128(4): 693–705.
- Supic G, Kozomara R, Jovic N, Zeljic K, Magic Z (2011) Prognostic significance of tumor-related genes hypermethylation detected in cancer-free surgical margins of oral squamous cell carcinomas. Oral Oncol 47(8): 702–708.
- Chang HW, Chow V, Lam KY, Wei W I, Yuen A (2002) Loss of E-cadherin expression resulting from promoter hypermethylation in oral tongue carcinoma and its prognostic significance. Cancer 94(2): 386–392.
- Kordi-Tamandani DM, Moazeni-Roodi AK, Rigi-Ladiz MA, Hashemi M, Birjandian E, et al. (2010) Promoter hypermethylation and expression profile of MGMT and CDH1 genes in oral cavity cancer. Arch Oral Biol 55(10): 809–814.
- Shaw RJ, Liloglou T, Rogers SN, Brown JS, Vaughan ED, et al. (2006) Promoter methylation of P16, RARβ, E-cadherin, cyclin A1 and cytoglobin in oral cancer: quantitative evaluation using pyrosequencing. Br J Cancer 94(4): 561–568.
- Huang SH, Lee HS, Mar K, Ji DD, Huang MS, Hsia KT (2010) Loss expression of O6‑methylguanine DNA methyltransferase by promoter hypermethylation and its relationship to betel quid chewing in oral squamous cell carcinoma. Oral Surg Oral Med Oral Pathol Oral Radiol Endod 109(6): 883‑889.
- Cho SW, Her SJ, Sun HJ, Choi OK, Yang JY, et al. (2008) Differential effects of secreted frizzled-related proteins (sFRPs) on osteoblastic differentiation of mouse mesenchymal cells and apoptosis of osteoblasts. Biochem. Biophys Res Commun 367(2): 399–405.
- Lo Muzio, L, Pannone G, Staibano S, Mignogna MD, Rubini C, et al. (2003) Survivin expression in oral squamous cell carcinoma. Br J Cancer 89: 2244–2248.
- Khan Z, Tiwari RP, Mulherkar R, Sah NK, Prasad GB, et al. (2009) Detection of survivin and p53 in human oral cancer: Correlation with clinicopathologic findings. Head Neck 31(8): 1039–1048.
- Gasche JA, Goel A (2012) Epigenetic mechanisms in oral carcinogenesis. Future Oncol 8(11): 1407–1425.
- Chen YK, Hsue SS, Lin LM (2005) Survivin expression is regulated by an epigenetic mechanism for dmba-induced hamster buccal-pouch squamous-cell carcinomas. Arch. Oral Biol 50(6): 593–598.
- Hsue SS, Wang WC, Chen YK, Lin LM (2008) Expression of inhibitors of apoptosis family protein in 7,12-dimethylbenz[a]anthracene-induced hamster buccal-pouch squamous-cell carcinogenesis is associated with mutant p53 accumulation and epigenetic changes. Int J Exp Pathol 89: 309–320.
- Kulkarni V, Saranath D (2004) Concurrent hypermethylation of multiple regulatory genes in chewing tobacco associated oral squamous cell carcinomas and adjacent normal tissues. Oral Oncol 40(2): 145–153.
- Kato K, Hara A, Kuno T, Mori H, Yamashita T, et al. (2006) Aberrant promoter hypermethylation of p16 and MGMT genes in oral squamous cell carcinomas and the surrounding normal mucosa. J Cancer Res Clin Oncol 132(11): 735–743.
- Ha PK, Califano JA (2006) Promoter methylation and inactivation of tumour suppressor genes in oral squamous-cell carcinoma. Lancet Oncol 7(1): 77–82.
- Hasegawa M, Nelson HH, Peters E, Ringstrom E, Posner M, et al. (2002) Patterns of gene promoter methylation in squamous cell cancer of the head and neck. Oncogene 21(27): 4231–4236.
- Sanchez-Cespedes M, Esteller M, Wu L, Nawroz-Danish H, Yoo GH, et al. (2000) Gene promoter hypermethylation in tumors and serum of head and neck cancer patients. Cancer Res 60(4): 892–895.
- Rosas SL, Koch W, da Costa Carvalho MG, Wu L, Califano J, et al. (2001) Promoter hypermethylation patterns of p16, O6-methylguanine-DNAmethyltransferase, and death-associated protein kinase in tumors and saliva of head and neck cancer patients. Cancer Res 61(3): 939–942.
- Mittag F, Kuester D, Vieth M, Peters B, Stolte B, et al. (2006) DAPK promotor methylation is an early event in colorectal carcinogenesis. Cancer Lett 240(1): 69–75.
- Levine AJ, Momand J, Finlay CA (1991) The p53 tumour suppressor gene. Nature 351(6326): 453–456.
- Boyle JO, Hakim J, Koch W, van der Riet P, Hruban RH, et al. (1998) The incidence of p53 mutations increases with progression of head and neck cancer. Cancer Res 53(19): 4477–4480.
- Caamano J, Zhang SY, Rosvold EA, Bauer B, Klein-Szanto AJ (1993) p53 alterations in human squa mous cell carcinomas and carcinoma cell lines. Am J Pathol 142(4): 1131–1139.
- Baral R, Patnaik S, Das BR (1998) Co-overexpression of p53 and c-myc proteins linked with advanced stages of betel- and tobacco-related oral squamous cell carcinomas from eastern India. Eur. J. Oral Sci 106(5): 907–913.
- Huang KH, Huang SF, Chen IH, Liao CT, Wang HM, et al. (2009) Methylation of RASSF1A, RASSF2A, and HIN-1 is associated with poor outcome after radiotherapy, but not surgery, in oral squamous cell carcinoma. Clin Cancer Res 15(12): 4174–4180.
- Taioli E, Ragin C, Wang XH, Chen J, Langevin SM, et al. (2009) Recurrence in oral and pharyngeal cancer is associated with quantitative MGMT promoter methylation. BMC Cancer 9: 354.
- Huang KH, Huang SF, Chen IH, Liao CT, Wang HM, et al. (2009) Methylation of RASSF1A, RASSF2A, and HIN-1 is associated with poor outcome after radiotherapy, but not surgery, in oral squamous cell carcinoma. Clin Cancer Res 15(12): 4174–4180.
- Omar EA (2013) The Outline of Prognosis and New Advances in Diagnosis of Oral Squamous Cell Carcinoma (OSCC). Journal of Oral Oncology Volume 519312: 519313.
- Tanaka C, Uzawa K, Shibahara T, Yokoe H, Noma H (2003) Expression of an Inhibitor of Apoptosis, Survivin, in Oral Carcinogenesis. J Dent Res 82(8): 607-611.
- Li YF, Hsiao YH, Lai YH, Chen YC, Chen YJ, et al. (2015) DNA methylation profiles and biomarkers of oral squamous cell carcinoma. Epigenetics 10(3): 229--236.
- Asokan GS, Jeelani S, Gnanasundaram N (2014) Promoter Hypermethylation Profile of Tumour Suppressor Genes in Oral Leukoplakia and Oral Squamous Cell Carcinoma J Clin Diagn Res Vol8(10).
- Basu B, Chakraborty J, Chandra A, Katarkar A, Baldevbhai JRK, et al. (2017) Genome-wide DNA methylation profile identified a unique set of differentially methylated immune genes in oral squamous cell carcinoma patients in India. Clinical Epigenetics 9: 13.
- Camillo CMC, Lourenço SV, Puga RD, Damascena AS, Teshima THN, et al. (2017) Profile of apoptotic proteins in oral squamous cell carcinoma: A cluster analysis of 171 cases. Applied Cancer Research 37: 2.
- Bhatia V, Goel MM, Makker A, Tewari S, Yadu A, et al. (2014) Promoter Region Hypermethylation and mRNA Expression of MGMT and p16 Genes in Tissue and Blood Samples of Human Premalignant Oral Lesions and Oral Squamous Cell Carcinoma. BioMed Research International 248419: 10.
- Irimie AI, Braicu C, Zanoaga O, Pileczki V, Gherman C, et al. (2015) Campian, R.S. Epigallocatechin-3-gallate suppresses cell proliferation and promotes apoptosis and autophagy in oral cancer ssc-4 cells. Onco Targets Ther 8: 461–470.
- Kumar A, Sarode SC, Sarode GS, Majumdar B, Patil S, et al. (2017) Beyond gene dictation in oscc progression and its therapeutic implications. Translational Research in Oral Oncology 2: 1–14.
- A Varshitha (2015) Prevalence of Oral Cancer in India. Pharm Sci & Res 7(10): 845-848.
- Stewart BW, Greim H, Shuker D, Kauppinen T (2003) Defence of iarc monographs. Lancet 361(9365): 1300.
- Irimie AI, Braicu C, Zanoaga O, Pileczki V, Gherman C, et al. (2015) Epigallocatechin-3-gallate suppresses cell proliferation and promotes apoptosis and autophagy in oral cancer ssc-4 cells. Onco Targets Ther 8: 461–470.
- Guerrero-Preston R, Baez A, Blanco A, Berdasco M, Fraga M, et al. (2009) Global DNA methylation: A common early event in oral cancer cases with exposure to environmental carcinogens or viral agents. P R Health Sci J 28(1): 24–29.
- Kerr J, Wyllie, Currie A (1972) Apoptosis: a basic biological phenomenon with wide-ranging implications in tissue kinetics. Br J Cancer 26(4): 239-257.
- Fracalossi A, Silva M, Oshima C, Ribeiro DA (2010) Wnt/beta-catenin signalling pathway following rat tongue carcinogenesis induced by 4-nitroquinoline 1-oxide. Exp Mol Pathol 88(1): 176-183.
- Noguti J, Moura De CFG, Hossaka TA, Franco M, Oshima CTF, et al. (2012) The Role of Canonical WNT Signaling Pathway in Oral Carcinogenesis: A Comprehensive Review. Anticancer research 32(3): 873-878.
- Mascolo M, Siano M, Ilardi G, Russo D, Merolla F, et al. (2012) Epigenetic Disregulation in Oral Cancer. Int J Mol Sci 13(2): 2331-2353.
- Lingen MW, Pinto A, Mendes RA, Franchini R, Czerninski R, et al. (2011) Genetics ⁄ epigenetics of oral premalignancy: current status and future research. Oral Diseases 17(1): 7–22.






























