Metformin Targets Cholesterol Biosynthesis Pathway, GM1 Lipid Raft Stabilization, EGFR Signaling and Proliferation in Triple Negative Breast Cancers
Reema S Wahdan-Alaswad*, Susan M Edgerton, Hiba S Salem and Ann D Thor
Department of Pathology, University of Colorado Anschutz Medical Campus, USA
Submission: February 01, 2018; Published: February 22, 2018
*Correspondence Address: Reema S Wahdan-Alaswad, Department of Pathology, School of Medicine, University of Colorado Anschutz Medical Campus, Mail Stop B216, 12631 E. 17th Ave, Rm 2215A, Aurora, CO 80045; USA, Tel: 303-724-3704; Email: ann.thor@ucdenver.edu
How to cite this article: Reema S W-A, Susan M E, Hiba S S, Ann D Thor. Metformin Targets Cholesterol Biosynthesis Pathway, GM1 Lipid Raft Stabilization, EGFR Signaling and Proliferation in Triple Negative Breast Cancers. Canc Therapy & Oncol Int J. 2018; 9(3): 555765. DOI: 10.19080/CTOIJ.2018.09.555765
Abstract
Triple negative breast cancer (TNBC) cells are particularly dependent on dysregulation of carbohydrate and lipid metabolism. We have demonstrated that the anti-diabetic agent metformin is particularly effective against TNBC, inhibiting proliferation, motility, pro-carcinogenic signaling, fatty acid synthesis and reducing the cancer stem cell subpopulation. In this study we evaluated the downstream effects of metformin on lipid metabolism and subcellular functions, using human derived TNBC cells MDA-MB-468, MDA-MB-231, BT-549 and HCC-70. Metformin inhibited transcription and translation of over 20 genes/enzymes in the cholesterol biosynthesis pathway, including HMG-CoA, a critical rate- limiting step in cholesterol synthesis and the target of statin drugs. It also down regulates cholesterol/GM1 incorporation into the cell membrane, destabilizes GM1 but not caveolin lipid rafts and down-regulates EGFR expression, activation and signaling (dependent in part on GM1 raft localization).
The antiproliferative activity of metformin was compared to growth inhibition induced by a cholesterol-depleting agent and statinmimic methyl-beta-cylodextrin (MβCD). When metformin and MβCD were used in combination, growth inhibition of TNBC was significantly increased. These studies provide novel mechanistic data on the action of metformin against TNBC. Given the potent effects of metformin and statin-mimetics on lipid biosynthesis (fatty acids and cholesterol), destabilization of GM1 rafts and EGFR signaling further studies to explore a combinatorial approach against dysregulation of carbohydrate and lipid metabolism in TNBC cancers are warranted. This will offer a new treatment paradigm for TNBC patients to inhibit tumor growth, enhance patient survival and modulate resistance to anti-EGFR tyrosine kinase inhibitors (TKIs).
Keywords: Metformin, Triple Negative Breast Cancer; Cholesterol; Lipid rafts; Statin
Abbreviations: Akt: Protein kinase B; ATP: Adenosine Triphosphate; DAPI: 4,6-diamidino-2-phenylindole; DHCR24: 24-Dehydrocholesterol Reductase; EGFR: Epidermal Growth Factor Receptor; FASN: Fatty Acid Synthase; GAPDH: Glyceraldehyde 3-phosphate Dehydrogenase; GM1: Monosialo Tetra Hexosyl Ganglioside; HMG-CoA: 3-hydroxy-3-methyl-glutaryl Coenzyme A Reductase; HMGCR: 3-Hydroxy-3-Methylglutaryl- CoA Reductase; HMGCS1: 3-Hydroxy-3-Methylglutaryl-CoA Synthase 1; HRP: Horseradish Peroxidase; IC50: Inhibitory Concentration 50; IGF1R: Insulin-like growth factor 1 Receptor; LBR: Lamin B Receptor; LSS: Lanosterol Synthase; MAPK: Mitogen-activated Protein kinase; M p CD: Methyl-beta-cyclodextrin; Met: Metformin, 1,1-dimethylbiguanide hydrochloride; MTS: 3-(4,5- dimethylthiazol-2-yl)-2,5-diphenltetrazolium Bromide; MVD: Mevalonate Diphosphate Decarboxylase; PI3K: Phosphoinositide 3-kinase; PTEN: Phosphatase and Tensin Homolog; RT-PCR: Reverse Transcriptase Polymerase Chain Reaction; SC5DL: Sterol-C5-Desaturase; SQLE: Squalene Epoxidase; Src: Proto-Oncogene Sarcoma, Tyrosine-Protein kinase; TGF- p : Transforming growth factor-beta; TKI: Tyrosine kinase Inhibitor; TNBC: Triple Negative Breast Cancer; UCCC: University of Colorado Cancer Center.
Introduction
Triple negative breast cancers (TNBC) are phenotypically aggressive and molecularly diverse, with a high rate of chemoresistance and a few potent targeted therapeutic options. The incidence of TNBC is increased in women with dysregulation of carbohydrate and lipid metabolism. Furthermore, these patients have a shortened disease free and overall survival. Cancer cells undergo shifts in carbohydrate and lipid metabolism, known as the ʽWarburg effectʼ and the ʽlipid switchʼ, respectively. These metabolic changes provide more intracellular energy (ATP) derived from glucose, glucosamine and other precursors, as well as fatty acids and cholesterol to facilitate cancer growth and survival. The abundance of lipids, including cholesterol, in TNBC facilitates specific cell substructures, such as lipid rafts, whose integrity is required for membrane receptor activity, cell adhesion, proliferation, migration, cell signaling and survival [1,2]. The majority of TNBC have significant expression of the epidermal growth factor receptor (EGFR), whose activity is closely associated with cancer progression, Akt and PI3K/MAPK signaling. It localizes to specific lipid rafts known as GM1 [3], comprised of cholesterol, sphingolipids, gangliosides, Src and EGFR [4,5]. This subcellular location has been associated with resistance to anti-tyrosine kinase inhibitors (TKI), whereas disruption of lipid rafts may alter EGFR signaling, induce TKI sensitivity and result in cancer cell death [6].
Chronic energy excess and physical inactivity are associated with alterations of carbohydrate and fatty acid metabolism, often resulting in obesity, metabolic syndrome and type 2 diabetes. Women with metabolic dysregulation often associated with increased insulin and insulin-like growth factor, serum cholesterol and glucose, experience higher rates of cancer, including breast cancer and TNBC. The anti-diabetic agent metformin (1,1-dimethylbiguanide hydrochloride) typically stabilizes metabolic dysregulation, at least early on in both metabolic syndrome and type 2 diabetes. Epidemiological data has shown that metformin reduces both the incidence and mortality of several types of cancer [7-11]. Many, but not all epidemiologic studies show a reduction in breast cancer incidence and prolonged survival in breast cancer patients treated with metformin. For example, a retrospective study of breast cancer patients treated with neoadjuvant chemotherapy, with and without metformin, showed a significantly higher chemotherapy response in patients who also received metformin [12].
Numerous studies of metformin in breast cancer patients, including those treated with a wide variety of chemotherapeutic agents are ongoing [13]. The majority of preclinical animal model studies have also demonstrated a reduction in mammary tumor formation and growth with metformin administration [14-18]. Mechanisms of metformin action against breast cancer have been subdivided into insulin-dependent and insulin-independent [19,20]. We have shown that metformin induces unique biological and molecular responses in different molecular subtypes of breast cancer [17,21-26]. TNBCs are exquisitely more sensitive to metformin than other subtypes of breast cancer [17].
We have shown that in TNBC cells, metformin abrogates growth, reduces oncogenicity and induces cell death via both internal and external pathways at a lower IC50 than other breast cancer subtypes [17,24]. We have also shown that glucose control within the normal physiologic range; as compared to conditions of hyperglycemia enhance metformin activity [24]. We have demonstrated that metformin targets STAT3, reduces cell growth, and induces apoptosis of TNBCs [22]. In mesenchymal stem like/claudin-low TNBC, we have shown that metformin abrogates transforming growth factor-β (TGF-β) oncogenic signaling by directly blocking translational and transcriptional activation of Smad2/3 [25]. Several groups have demonstrated that metformin selectively targets breast cancer stem cells, although the mechanisms underlying this interaction are not well understood [13,26-30].
As noted above, obesity and dysregulation of lipid metabolism are associated with a significant risk of breast cancer and a worse outcome for women who develop the disease [31]. While some of the risks may be attributed to hormonal shifts with the peripheral conversion of testosterone to estrogen by adipose tissue in the post-menopausal setting, other factors including up regulation of cytokines, growth factors, fatty acids and cholesterol, as well as disruption of immune function with obesity/metabolic shifts promote breast carcinogenesis. In contrast to metformin, epidemiologic data does not support a reduction in the incidence of all types of breast cancer with statin use. Cholesterol lowering drugs (particularly simvastatin) have been associated with longer disease specific survival in patients with breast cancer [32]. For example, in a population registry study from Finland, statin use was associated with lower rates of breast and colorectal cancer mortality [33]. Some have suggested that statins are more effective against TNBC than other breast cancer subtypes. A Kaiser Permanente Northern California Cancer Registry showed that women who took statins had proportionately fewer hormone receptor negative breast cancers (including TNBC), and cancers that did occur were significantly lower in grade and stage [34]. Shaitelman and colleagues at the MD Anderson Cancer Center recently performed a literature review of the impact of statin use on breast cancer. They also concluded that statins may benefit a subset of TNBC patients [35].
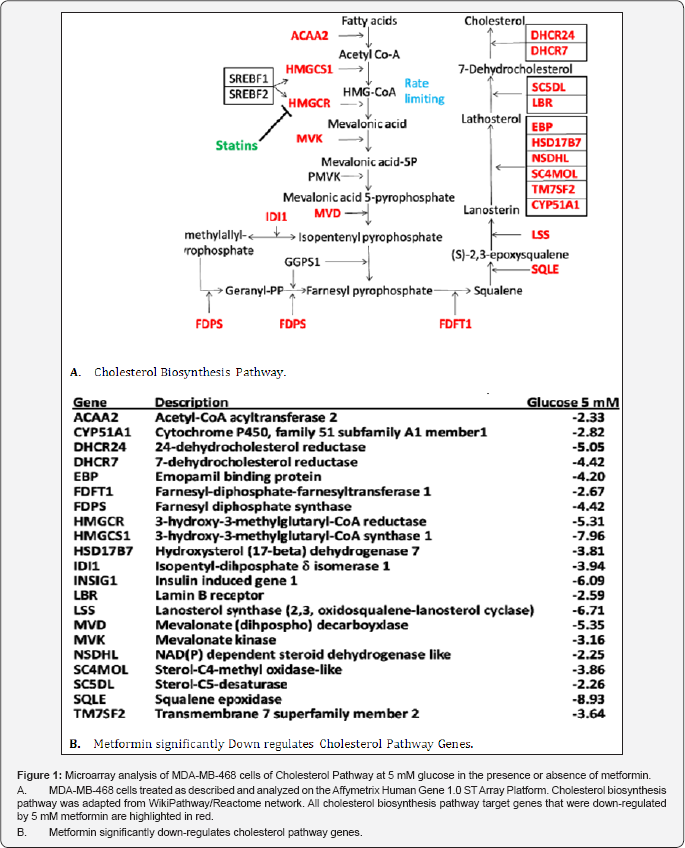
The current study is a derivative of our interest in carbohydrate and fat dysregulation associated with obesity and type 2 diabetes, and the use of metformin to decrease the risk or improve outcomes in women with breast cancer. We have previously reported that metformin directly targets fatty acid synthase (FASN), through activation of miR-193b [26]. Since FASN plays a critical role in de novo fatty acid synthesis and is important for survival of TNBC, we hypothesized that metformin might also target critical genes in the cholesterol synthesis and downstream biological functions of cholesterol, such as specific subsets of lipid rafts. In this report, we demonstrate that metformin inhibits the majority of genes included in the cholesterol biosynthesis pathway in TNBC (Figure 1), including the target of lipophilic statins HMG-CoA reductase. This is the rate limiting enzyme for cholesterol synthesis and is regulated via complex feedback mechanisms in normal mammalian cells. We also show for the first time that metformin has a major impact on cholesterol localization and integration into GM1 lipid rafts, as well as the co-localization and signaling induced by EGFR (a constitutive component of those rafts in TNBC).
Methods
Cell Culture Treatments
Human TNBC derived cell lines: MDA-MB-468, MDA-MB-231, BT-549, and HCC70 were obtained from American Type Culture Collection or University of Colorado Cancer Center (UCCC) Tissue Culture shared resource. The identity of all cell lines was authenticated UCCC Tissue Culture core. All cells were confirmed to be free of mycoplasma. Prior tissue culture conditions were previously reported [24,26]. For this study, cells were plated in 10% FBS, then starved with glucose free media (Invitrogen) DMEM/F12 media with 0-5% FBS for 24 h, then replenished with pure glucose supplemented at 5 mM or 17 mM glucose (Sigma Aldrich Co.). Metformin (MP Biomedicals) was dissolved in sterile water and used at the indicated concentration for each specific study. Methyl-beta-cyclodextrin (M β CD; Sigma Aldrich Co.) was used at 2.5 mM concentration in these studies.
Microarray
Methods used for Affymetrix Gene Chip© Whole Transcript (WT) sense target labeling assay protocol for MDA-MB-468 cells were previously published [24,26]. For these studies, Wiki Pathways/Reactome was used to outline cholesterol biosynthesis pathway. Analysis was performed using Partek and/ or Genespring software (Agilent). Gene expression was filtered on statistical significance (Student's t test, p<=0.05, Benjamini Hochberg correction) and genes with fold changes over control ≥ 2.0 or ≤ -2.0 were accepted for further analysis.
Real Time quantitative Reverse Transcriptase-PCR (qRT-PCR) for Cholesterol Biosynthesis Pathway
MDA-MB-468 and BT-549 cells were seeded in glucose 5 mM for 24 hrs then treated with 5 mM metformin for 24 hrs, RNA was harvested, and purified for cDNA synthesis as previously described [24,26]. Predesigned gene specific primer and probe sets were obtained from SA Biosciences. qRT-PCR synthesis of cholesterol pathway genes (HMGCS1, HMGCR, MVD, SQLE, LSS, SC5DL, LBR, and DHCR24) was performed using DyNAmo Flash SYBR Green qPCR kit (#F-415L, ABI) according to the manufacturer's protocol on an ABI 7500 Fast Real Time PCR System. The relative mRNA levels were calculated using the comparative Ct method (ΔΔCt) as previously published [26]. Reported values are the means and standard errors of three biological replicates.
Western Blot
Whole cell protein extracts preparation was previously described [24,26]. Blots were probed with antibodies to HMGCS1 (#87246), HMGCR (#174830), LSS (#124785), Seladinl (DHCR24) (#137845) obtained from Abcam, and EGFR (F4; #E3138), p-actin (AC-75) (Sigma Aldrich Co.). Secondary antibody goat anti-mouse HRP and goat anti-rabbit HRP were obtained from Jackson Immuno Research.
Cell Proliferation Assay
MTS assay was used as described [24] to determine cell viability. For reported experiments, cells were grown in glucose at either 5 mM or 17 mM levels with 0.5% FBS, or the same medium containing various doses of metformin (5 mM or 10 mM). After 6 days, wells were read at 490 nm with a microplate reader. All conditions were tested in triplicate samples, and the percentage of surviving cells from each group was calculated relative to controls. Control conditions were defined as 100% survival.
Lipid Raft Labeling Kit
Vybrant® lipid raft labeling kit (V-34405) was purchased from Life Technology and performed per manufacturer guidelines. After permeabilization, cells were washed, stained for EGFR (green) or GM1 (red) overnight, counterstained, mounted with DAPI and visualized using an Olympus microscope at 100x.
Statistics
Statistical calculations of metformin-mediated inhibition of cholesterol pathway genes were performed using Graph Pad Prism 7 software. Statistical analysis of the experimental data was performed using a 2-sided Student t-test. Significance was set at P<0.05. All bar graphs are representative of at least three biological replicates and/or three independent experiments. All western blots and immunofluorescent experiments are representative of three independent experiments.
Results
Metformin Targets Cholesterol Biosynthesis
We have previously reported that metformin has unique biological and molecular anti-cancer effects against TNBC and that the IC50 of metformin needed to induce those changes is significantly lower (~10 fold) at normal levels of glucose (5 mM, as compared to supra-physiologic levels as may be encountered in untreated diabetes [17,26]). We have recently shown that metformin inhibits FASN (upstream of the cholesterol synthesis pathway), through a novel mechanism involving miR-193b [26]. As an extension of our prior work [24,26], we used the Affymetrix Human Gene 1.0 ST Array data from two TNBC cell lines, MDA- MB-468 and MDA-MB-231, treated with or without metformin. Signal intensity estimates and P values were performed to determine the relative fold change ratios; those over ≥ 2.0 or ≤ 2.0 were considered significant. Metformin significantly inhibited over 20 key enzymes/genes in the cholesterol biosynthesis pathway (highlighted in red, Figure 1A). These genes and fold differences are shown in Figure 1B.
Metformin Inhibits transcriptional and translational expression of key components of cholesterol pathway
We next validated the microarray data, using quantitative change to RT-PCR methods comparing metformin treated to untreated TNBC cell lines MDA-MB-468 and BT-549, for levels of specific gene expression. Cells treated with 5 mM metformin showed significant attenuation of HMGCS1, HMGCR, MVD, SQLE, LSS, SC5DL, LBR, and DHCR24, all critical genes involved in cholesterol biosynthesis (Figure 2A and 2B). Next, we examined metformin-mediated inhibition of gene encoded proteins. As shown in Figure 2C, significant reductions in the expression of several proteins by metformin treated, as compared to control untreated cells was observed including HMGCS1, HMGCR, Seladinl (identified by gene DHCR24), and to a lesser degree LSS.


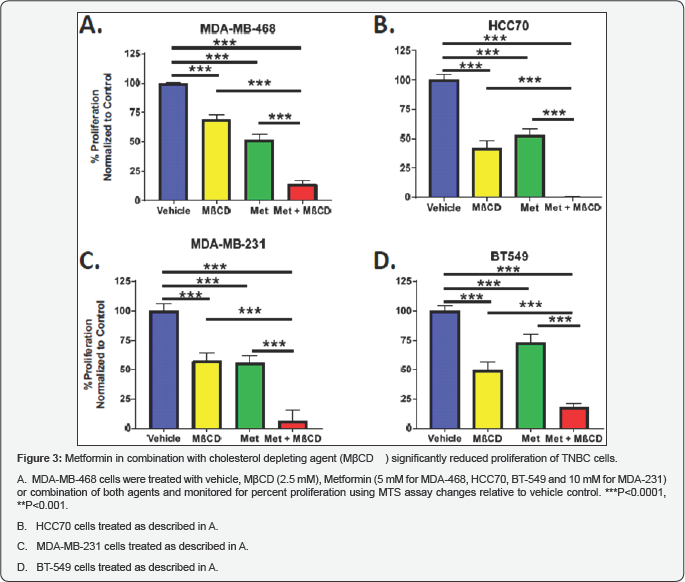
Metformin alone, or in combination with a cholesterol blocking agent M p CD, blocks proliferation of TNBC cells
We next compared the activity of metformin to methyl-beta- cyclodextrin (M β CD), as well as these agents in combination, to inhibit TNBC cell growth. M β CD is a cholesterol depleting agent that is used widely in cell culture experiments as a statin- mimetic for membrane studies. It neither binds to nor inserts into the plasma membrane, although it reduces the availability of cholesterol for GM1 lipid rafts. Four TNBC cell lines were treated with increasing concentrations of M p CD, relative to increasing concentrations of metformin to identify the IC50 for each agent and cell line tested (data not shown). These concentrations were then used for studies shown in Figure 3 (A: MDA-MB-468, B: HCC70, C: MDA-MB-231, D: BT-549). Both M p CD and metformin inhibited TNBC cell proliferation. In combination, these agents were superior to either drug alone as shown in gure 3A-3D.
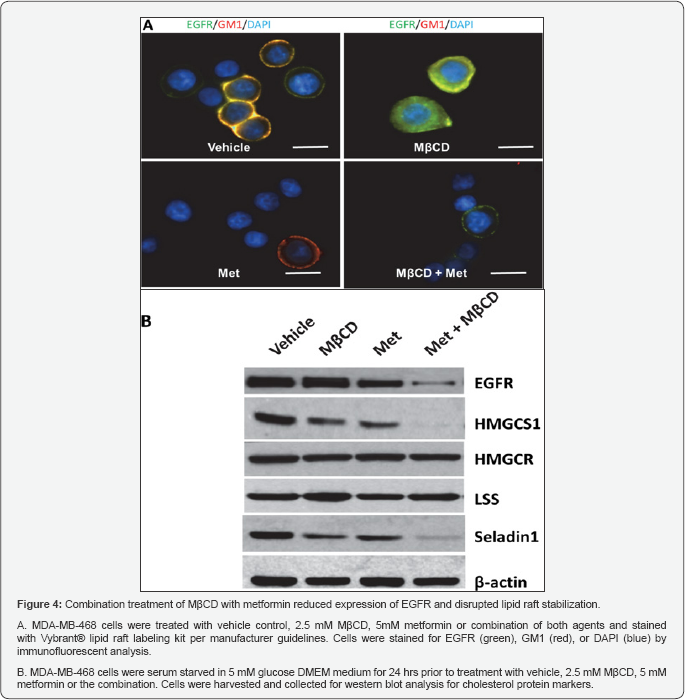
Metformin Reduces EGFR Expression and Cholesterol- Enriched GM1 Lipid Rafts
Given the role lipid rafts play as a membrane platform for signaling molecules and receptors, we next interrogated whether these agents alone or in combination would disrupt GM1 rafts and/or EGFR expression or activity. As compared to the vehicle control (Figure 4A, left upper panel), M β CD significantly reduced GM1 (red) as well as co-localization (yellow) of EGFR and GM1. M β CD treatment resulted in higher cytoplasmic expression of EGFR, not localized to cell membrane relative to the vehicle control. (Figure 4A, upper right panel). Metformin alone abolished EGFR expression (green signal) and co-localization with GM1 (yellow, lower left panel). The combination of M β CD and metformin is shown in Figure 4, lower right panel. With both agents there was a marked reduction in EGFR, no GM1 or co-localization signal. Our results were also reproducible in other TNBC cells in culture, including: BT-549, HCC70 and MDA-MB-231 cells (data not shown). Using similar methods M p CD had no effect on another subtype of lipid rafts in TNBC, identified by caveolin expression (data not shown but similar to previous reports [6]). Finally, we tested whether metformin alone, or in combination with M β CD, could reduce the expression of components of the cholesterol pathway in MDA-MB-468 cells. Metformin reduced HMGCS1, HMGCR, Seladin1 and EGFR, whereas M β CD lowered the first three but had no effect on EGFR expression. In combination, there was significant reduction of EGFR, HMGCS1, HMGCR, and Seladinl proteins.
Discussion
TNBC are associated with particularly poor patient outcomes, due in part to a lack of effective therapeutic targets and a propensity to develop resistance to standard chemotherapy. We have previously demonstrated that metformin induces cell death and blocks stem cell expansion in TNBC, via an increase in miR-193b that down-regulates FASN. The latter is critical for de novo fatty acid synthesis, cholesterol biosynthesis, TNBC survival and maintenance of the stem cell pool. TNBC have also been shown to up-regulate a number of genes that play a role in cholesterol biosynthesis. Cholesterol induces potent pro- carcinogenic effects on the tumor cells, promoting signaling through membrane bound receptors, facilitating intracellular signaling pathways, serving as an anchor for intracytoplasmic filaments that facilitate motility, invasion, providing precursors for energy and replication.
Lipophilic statins that interfere with cholesterol synthesis have been shown to reduce the incidence of TNBC, disproportionately in comparison to hormone receptor positive breast cancer. Breast cancer patient treatment with statins has also been shown to confer a survival benefit for affected patients. In vitro studies have further shown that cholesterol depletion in TNBC reduces cell growth and induces cell death. Others have previously reported that metformin inhibits HMGCR (the rate limiting enzyme and key regulator of the mevalonate pathway, the primary target of statins) in other cancers [36,37]. We are the first to perform a comprehensive survey of metformin's action on the cholesterol biosynthesis pathway. We also sought to determine if metformin and other agents like M β CD that deplete cholesterol or block its synthesis may induce greater anti-cancer activity than metformin alone. Our data shows that metformin inhibits over 20 key enzymes and genes that play a critical role in cholesterol biosynthesis. It was not only able to directly attenuate transcriptional activation of these key genes involved in cholesterol biosynthesis, it also reduced translational activation of down-stream signaling pathways. Although some of these metformin targets overlap with those inhibited by statins, the majority are unaffected by statin alone. Thus, metformin associated anti-TNBC activity shows far greater redundancy in mechanisms involving lipid metabolism, abrogating FASN and fatty acid synthesis, reducing cholesterol synthesis and causing disassociation of GM1 lipid rafts with internalization of EGFR.
EGFR expression typically correlates with a worse poor prognosis in breast cancers. It is usually over expressed in TNBC, most notably in the molecular subsets known as basal-like-1 and basal-like-2 [38,39]. Despite expression of the receptor, anti- EGFR TKI therapy has typically not been associated with patient benefit or prolonged survival in women with TNBC [40]. In addition to frequent TKI resistance, TNBC may also demonstrate activation of IGF1R and mutation/loss of PTEN, providing additional molecular mechanisms for TKI resistance.
Conclusion
On the basis of novel data presented above, non-insulin dependent inhibition of the cholesterol biosynthesis pathway by metformin action may provide significant benefits for patients with TNBC. Although preliminary, our data also suggests that a combination of metformin and statin may provide even greater inhibition of cell proliferation, especially in patients with TNBC. Although not specifically addressed by our studies, destabilization of GM1 rafts with internalization of EGFR by metformin, or metformin and statin, may induce sensitivity to anti-EGFR TKIs, in addition to reducing cell proliferation and inducing cell death. Finally, we note that metformin and statins are FDA-approved, well tolerated, and widely available inexpensive oral agents.
Further studies of the role of cholesterol biosynthesis on cancer cell growth, survival and oncogenicity, as well as its blockade by metformin and statins may provide a new treatment paradigm of benefit to TNBC patients (particularly those with metabolic dysregulation, obesity and type II diabetes).
Acknowledgment
We thank professional research assistant Zeying Fan for her help and contribution. Support for the UCDenver Micro Array Core Facility and the UCDenver Cancer Canter DNA Sequence and Analysis Shared Resource Core for cell line authentication is provided in part by Colorado’s NIH/NCI Cancer Center Support Grant P30CA043934.
Disclosures of Potential Conflict of Interest.
No potential conflicts of interest were disclosed.
Grant Support
Grant support provided in part by Susan G Komen for the Cure K100575 to RWA, ZF, SME, and ADT and ACS-IRG 16-184¬56 RWA from the American Cancer Society.
References
- George KS, Wu S (2012) Lipid raft: A floating island of death or survival. Toxicol Appl Pharmacol 259(3): 311-319.
- Liu Y, Sun R, Wan W, Wang J, Oppenheim JJ, et al. (2007) The involvement of lipid rafts in epidermal growth factorinduced chemotaxis of breast cancer cells. Mol Membr Biol 24(2): 91-101.
- Zhang Z, Wang L, Du J, Li Y, Yang H, et. al. (2016) Lipid raft localization of epidermal growth factor receptor alters matrix metalloproteinase-1 expression in SiHa cells via the MAPK/ERK signaling pathway. Oncol Lett 12(6): 4991-4998.
- Caruso JA, Stemmer PM (2011) Proteomic profiling of lipid rafts in a human breast cancer model of tumorigenic progression. Clin Exp Metastasis 28(6): 529-540.
- Patra SK (2008) Dissecting lipid raft facilitated cell signaling pathways in cancer. Biochim Biophys Acta 1785(2): 182-206.
- Irwin ME, Mueller KL, Bohin N, Ge Y, Boerner JL (2011) Lipid raft localization of EGFR alters the response of cancer cells to the EGFR tyrosine kinase inhibitor gefitinib. J Cell Physiol 226(9): 2316-2328.
- Chlebowski RT, McTiernan A, Wactawski-Wende J, Manson JE, Aragaki AK, et al. (2012) Diabetes, metformin, and breast cancer in postmenopausal women. J Clin Oncol 30(23): 2844-2852.
- Col NF, Ochs L, Springmann V, Aragaki AK, Chlebowski RT (2012) Metformin and breast cancer risk: a meta-analysis and critical literature review. Breast Cancer Res Treat 135(3): 639-646.
- Decensi A, Puntoni M, Goodwin P, Cazzaniga M, Gennari A, et al. (2010) Metformin and cancer risk in diabetic patients: a systematic review and meta-analysis. Cancer Prev Res (Phila) 3(11): 1451-1461.
- Del Barco S, Vazquez-Martin A, Cufi S, Oliveras-Ferraros C, Bosch- Barrera J, et al. (2011) Metformin: multi-faceted protection against cancer. Oncotarget 2(12): 896-917.
- Evans JM, Donnelly LA, Emslie-Smith AM, Alessi DR, Morris AD (2005) Metformin and reduced risk of cancer in diabetic patients. BMJ 330(7503): 1304-1305.
- Jiralerspong S, Palla SL, Giordano SH, Meric-Bernstam F, Liedtke C, et al. (2009) Metformin and pathologic complete responses to neoadjuvant chemotherapy in diabetic patients with breast cancer. J Clin Oncol 27(20): 3297-3302.
- Niraula S, Dowling RJ, Ennis M, Chang MC, Done SJ, et al. (2012) Metformin in early breast cancer: a prospective window of opportunity neoadjuvant study. Breast Cancer Res Treat 135(3): 821-830.
- Checkley LA, Rudolph MC, Wellberg EA, Giles ED, Wahdan-Alaswad RS, et al. (2017) Metformin Accumulation Correlates with Organic Cation Transporter 2 Protein Expression and Predicts Mammary Tumor Regression In Vivo. Cancer Prev Res (Phila) 10(3): 198-207.
- Giles ED, Wellberg EA, Astling DP, Anderson SM, Thor AD, et al. (2012) Obesity and overfeeding affecting both tumor and systemic metabolism activates the progesterone receptor to contribute to postmenopausal breast cancer. Cancer Res 72(24): 6490-6501.
- Kasznicki J, Sliwinska A, Drzewoski J (2014) Metformin in cancer prevention and therapy. Ann Transl Med 2(6): 57.
- Liu B, Fan Z, Edgerton SM, Deng XS, Alimova IN, et al. (2009) Metformin induces unique biological and molecular responses in triple negative breast cancer cells. Cell Cycle 8(13): 2031-2040.
- Rice S, Pellat L, Ahmetaga A, Bano G, Mason HD, et al. (2015) Dual effect of metformin on growth inhibition and oestradiol production in breast cancer cells. Int J Mol Med 35(4): 1088-1094.
- Camacho L, Dasgupta A, Jiralerspong S (2015) Metformin in breast cancer - an evolving mystery. Breast Cancer Res 17: 88.
- Morales DR, Morris AD (2015) Metformin in cancer treatment and prevention. Annu Rev Med 66: 17-29.
- Alimova IN, Liu B, Fan Z, Edgerton SM, Dillon T, et al. (2009) Metformin inhibits breast cancer cell growth, colony formation and induces cell cycle arrest in vitro. Cell Cycle 8(6): 909-915.
- Deng XS, Wang S, Deng A, Liu B, Edgerton SM, et al. (2012) Metformin targets Stat3 to inhibit cell growth and induce apoptosis in triple¬negative breast cancers. Cell Cycle 11(2): 367-376.
- Liu B, Fan Z, Edgerton SM, Yang X, Lind SE, et al. (2011) Potent anti¬proliferative effects of metformin on trastuzumab-resistant breast cancer cells via inhibition of erbB2/IGF-1 receptor interactions. Cell Cycle 10(17): 2959-2966.
- Wahdan-Alaswad R, Fan Z, Edgerton SM, Liu B, Deng XS, et al. (2013) Glucose promotes breast cancer aggression and reduces metformin efficacy. Cell Cycle 12(24): 3759-3769.
- Wahdan-Alaswad R, Harrell JC, Fan Z, Edgerton SM, Liu B, et al. (2016) Metformin attenuates transforming growth factor beta (TGF-beta) mediated oncogenesis in mesenchymal stem-like/claudin-low triple negative breast cancer. Cell Cycle 15(8): 1046-1059.
- Wahdan-Alaswad RS, Cochrane DR, Spoelstra NS, Howe EN, Edgerton SM, et al. (2014) Metformin-induced killing of triple-negative breast cancer cells is mediated by reduction in fatty acid synthase via miRNA-193b. Horm Cancer 5(6): 374-389.
- Hirsch HA, Iliopoulos D, Struhl K (2013) Metformin inhibits the inflammatory response associated with cellular transformation and cancer stem cell growth. Proc Natl Acad Sci U S A 110(3): 972-977.
- Hirsch HA, Iliopoulos D, Tsichlis PN, Struhl K (2009) Metformin selectively targets cancer stem cells, and acts together with chemotherapy to block tumor growth and prolong remission. Cancer Res 69(19): 7507-7511.
- Iliopoulos D, Hirsch HA, Struhl K (2011) Metformin decreases the dose of chemotherapy for prolonging tumor remission in mouse xenografts involving multiple cancer cell types. Cancer Res 71(9): 3196-3201.
- Zhu P, Davis M, Blackwelder AJ, Bachman N, Liu B, et al. (2014) Metformin selectively targets tumor-initiating cells in ErbB2- overexpressing breast cancer models. Cancer Prev Res (Phila) 7(2): 199-210.
- Blucher C, Stadler SC (2017) Obesity and Breast Cancer: Current Insights on the Role of Fatty Acids and Lipid Metabolism in Promoting Breast Cancer Growth and Progression. Front Endocrinol (Lausanne) 8: 293.
- Ahern TP, Lash TL, Damkier P, Christiansen PM, Cronin-Fenton DP (2014) Statins and breast cancer prognosis: evidence and opportunities. Lancet Oncol 15(10): e461-468.
- Haukka J, Niskanen L, Auvinen A (2017) Risk of Cause-Specific Death in Individuals with Cancer-Modifying Role Diabetes, Statins and Metformin. Int J Cancer 141(12): 2437-2449.
- Kumar AS, Benz CC, Shim V, Minami CA, Moore DH, et al. (2008) Estrogen receptor-negative breast cancer is less likely to arise among lipophilic statin users. Cancer Epidemiol Biomarkers Prev 17(5): 1028-1033.
- Shaitelman SF, Stauder MC, Allen P, Reddy S, Lakoski S, et. al. (2017) Impact of Statin Use on Outcomes in Triple Negative Breast Cancer. J Cancer 8(11): 2026-2032.
- Karlic H, Thaler R, Gerner C, Grunt T, Proestling K, et. al. (2015) Inhibition of the mevalonate pathway affects epigenetic regulation in cancer cells. Cancer Genet 208(5): 241-252.
- Nilsson S, Huelsenbeck J, Fritz G (2011) Mevalonate pathway inhibitors affect anticancer drug-induced cell death and DNA damage response of human sarcoma cells. Cancer Lett 304(1): 60-69.
- Cancer Genome Atlas N (2012) Comprehensive molecular portraits of human breast tumours. Nature 490(7418): 61- 70.
- Lau YK, Du X, Rayannavar V, Hopkins B, Shaw J, et al. (2014) Metformin and erlotinib synergize to inhibit basal breast cancer. Oncotarget 5(21): 10503-10517.
- Viale G, Rotmensz N, Maisonneuve P, Bottiglieri L, Montagna E, et. al. (2009) Invasive ductal carcinoma of the breast with the "triple-negative” phenotype: prognostic implications of EGFR immunoreactivity. Breast Cancer Res Treat 116(2): 317-328.






























