The Third Domain Ligand Binding Fragment of Alpha-fetoprotein: Detection of Metastasis-associated Molecular Targets
*Gerald J Mizejewski
New York State Department of Health, Wadsworth Center, USA
Submission: August 31, 2017; Published: September 11, 2017
*Correspondence address: Gerald J Mizejewski, Division of Translational Medicine, Molecular Diagnostics Laboratory, Wadsworth Center, New York State Department of Health, PO Box 509, Empire State Plaza, Albany, NY 12201-0509, USA, Tel: 518-486-5900; Email: gerald.mizejewski@health.ny.gov
How to cite this article: Mizejewski GJ. The Third Domain Ligand Binding Fragment of Alpha-fetoprotein: Detection of Metastasis-associated Molecular Targets. Canc Therapy & Oncol Int J. 2017; 6(5): 555700. DOI: 10.19080/CTOIJ.2017.06.555700
Abstract
The vast majority of cancer deaths are due to metastasis to distal sites rather than from the primary tumor mass. Previous reports have established that the third domain of the alpha-fetoprotein polypeptide (AFP3D) consists of amino acid sequence stretches that participate in protein-to-protein docking interactions. These were detected by means of a computer software program configured to recognize and localize peptide sequences on specifically targeted proteins. The present report identified sequence fragments on AFP3D that could potentially interact with multiple metastasis-related proteins. The metastatic proteins were derived from four classes of such candidates, namely:
a. Cell adhesion and cell-to-cell contact proteins;
b. Extracellular matrix proteins;
c. Growth factor receptors; and
d. Growth factors and kinase enzymes.
Following detection and identification of the AFP3D-to-metastatic protein interaction sites, the "in silico” localized amino acid sequences were compared and aligned with prior published AFP interaction sites comprising hydrophobic ligands, various receptors, cell cycle proteins, and cation channels. Attempts were then made to assess the potential of multiple ligands/proteins competing for interaction sites on AFP3D. Previous publications and the present report have confirmed and verified the "in silico” identified interaction sites by experimental cell-based assays, mRNA microarrays, and kinase profile screenings. The experimentally-confirmed procedures and assays were performed in both in vitro MCF-7 cell cultures assays and in rodent animal models. Finally, the experimental findings on the AFP3D interactions with metastatic proteins were consistent with the computer-derived data.
Keywords: Alpha-fetoprotein; Growth factors; Cell adhesion; Receptors; Extracellular matrix; kinases; Cell contact; Metastasis
Abbreviations: AA: Amino Acid; AFP: Alpha-fetoprotein; 3D: Third Domain; PI3K: Phosphoinositol Kinase; mTOR: Mechanistic Target of Rapamycin; GAAD153: Growth Arrest and DNA Damage-Inducible-protein-153; PTEN: Phosphatase and Tensin Homolog; CHK: Checkpoint; GIP: Growth Inhibitory Peptide; CdK: Cyclin Dependent Kinase; PKC: Protein Kinase-C; MAP: Metastasis-associated Proteins; ECM: Extracellular Matric; MMP: Matrix Metalloproteinase's; ADAM: A Disintegrin and Metalloproteinase; KISSR: Kisspeptin Receptor; TSHR: Thyroid- Stimulating Hormone Receptor; LAMR: Laminin Receptor; TGF: Transforming Growth Factor
Introduction
Historical Background
Human alpha-fetoprotein (AFP) was one of the first biomarkers developed as a cancer biomarker and is still today considered the "gold standard" biomarker for liver cancer. AFP is a glycopolypeptide exhibiting a molecular mass of 69 kD with a 3-5% carbohydrate content [1]. Structurally, the full-length AFP molecule consists of three domains forming nine layers of peptide loops configured by 15 cysteine disulfide bridges [2]. The two-dimensional (secondary) image of AFP has been determined and been confirmed to be a V- or U-shaped molecular structure. Human AFP is classified as a member of the albuminoid gene family which consists of albumin, alpha-albumin, AFP, vitamin-D binding protein, and the AFP-related gene (ARG) protein [3]. Physiologically, AFP is known to be a serum carrier/transport molecule for fatty acids, bilirubin, retinoids, steroids, flavonoids, phytoestrogens, organic dyes, and various drugs [4]. This fetal glycoprotein can further serve as a regulator of both growth enhancement and inhibition. AFP has been denoted as an "oncofetal protein" being present during fetal development as well as adult tumorigenesis [5]. Hence, AFP has served both as a tumor and fetal defect biomarker in the clinical setting.
The existence of AFP receptors, binding, and interacting proteins have been reported for more than 25 years. It is now widely acknowledged that AFP binds to multiple cell surface receptors and cytoplasmic binding proteins [6-8]. However, AFP has never been reported to enter the intact nucleus, but is known to reside in the perinuclear space compartment following synthesis and/or cell uptake. Recent studies, reviews, and updates have established that the third domain of AFP is a multi-ligand binding fragment capable of binding and interacting with a vast array of steroids, retinoids, fatty acids, drugs, growth factors, and various protein and peptide receptors [9,10]. At least three major groups of cell surface receptors are known to bind AFP, namely:
i. The scavenger;
ii. Mucin; and
iii. Chemokine receptors.
The intracytoplasmic binding proteins interacting with AFP include the retinoic acid receptor, caspase-3, nuclear steroid receptors, phosphatase and tensin homolog (PTEN), and the growth arrest and DNA damage inducible protein 153(GADD153) [11]. It is further conceivable that AFP could interact with nuclear proteins during mitosis when the nuclear envelope temporarily dissolves. Other potential cell proteins interacting with AFP could include cell cycle proteins, lysolipid receptors, and cation channel proteins [12].
Data on mapping of protein-to-protein interaction sites on the third domain fragment of AFP (AFP3D) has been generated by means of computer" in silico" software. Many, if not most, of the computer-modeling protein interaction sites on AFP3D being reported have already been experimentally confirmed and verified in cell-based assays [12,13]. The present report describes a further type of in silico protein interaction with AFP3D, namely, metastasis-associated proteins such as:
a. Cell adhesion,
b. Extracellular matrix (ECM) proteins,
c. Growth factor receptors, and
d. Growth factor and kinase proteins.
Their cell functions encompass activities such as homophilic cell-to-cell adhesions, calcium-dependent stabilizing and maintaining cell connections, cell gap junction establishment, cell migration, anchorage-dependent cell death (anoikis), tumor invasion, degrading of extracellular matrix (ECM), integrin interaction, basement membrane adhesion and degradation, collagen proteolysis, and others (Table 1). The association of AFP with metastasis-associated proteins has not been previously reported, pursued, or addressed in the biomedical literature even though AFP has long been known to enhance metastasis in liver and other AFP-secreting tumors.
The continued search of new receptor and protein interacting sites on AFP3D can be scientifically justified in order to identify and provide insight into new and novel activation sites, receptor docking and occupation sequences, ligand targeting, and signal transduction networks. Such novel findings could pave the way for utilizing sub domains of AFP3D fragments as drug delivery platforms that target cells involved in cancer, autoimmune or inflammation disorders. Finally, new insights could be gained toward the identification of previously unknown binding/interaction partners for AFP that might enhance our understanding of AFP-to-target cells involved in the propagation of extra- and intracellular signaling.
Objectives and Aims
The objectives in the present report were to first pursue, search, and identify potential metastasis-associated protein interaction sites on AFP3D. Computer-modeling and protein- to-protein interaction and docking software were employed to identify specific amino acid (AA) sequence segments on AFP3D that could plausibly interact with known metastasis-associated proteins (MAPs) reported in the literature. Secondly, the different subtypes of MAPs are discussed in lieu of their specificity for engaging metastatic activation and/or deactivation processes and their possible interaction with AFP3D to alter or influence tumor cell detachment, spreading, and dissemination. Third, the MAP localization sites are compared to previously reported ligand and receptor binding sites already confirmed on AFP3D. Fourth, each member of the MAPs is discussed regarding its cellular activities and reported associations with ligand and receptor sites previously reported on AFP. Finally, experimental reports of AFP-derived peptide interaction with MAPs will be addressed in lieu of the present in silico findings.
Computer Modeling and Molecular Docking Software
The computer modeling of the 3D structure of human AFP was previously described [12-14]. Molecular docking (interaction) sites on AFP were identified and localized by means of proprietary computer software program (Peptimer Discovery Platform) developed and generously provided by Serometrix LLC, Syracuse, NY, as described in detail in prior reports [13,15,16]. The proprietary software modeling and simulation of protein-to-protein interaction sites are highly reproducible as described in previous research reports and have been repeatedly validated using in vitro whole cell-based assays and receptor binding measurements [17,18].
Metastasis-Associated Proteins
Cell Adherence and Cell-to-Cell Contact Proteins
The total cadherin super family of proteins includes protocadherins, desmogleins, and desmocollins all of which function in cell adhesion and the formation of adherin junctions to link cells together [19]. The cadherins are dependent on Table 1a: The properties of the metastasis-associated proteins (MAPs) calcium (Ca++) ions to function in cell-to-cell adhesion and these proteins contain extracellular Ca++ binding repeat domains and intracellular domains for adaptor and cell signaling functions. Different subtypes of cadherins can join cells together in both homotypic and heterotypic binding fashions and are classified depending on the cadherin prefix indicated (N = neural, E = epithelial, etc.) in their name [20].
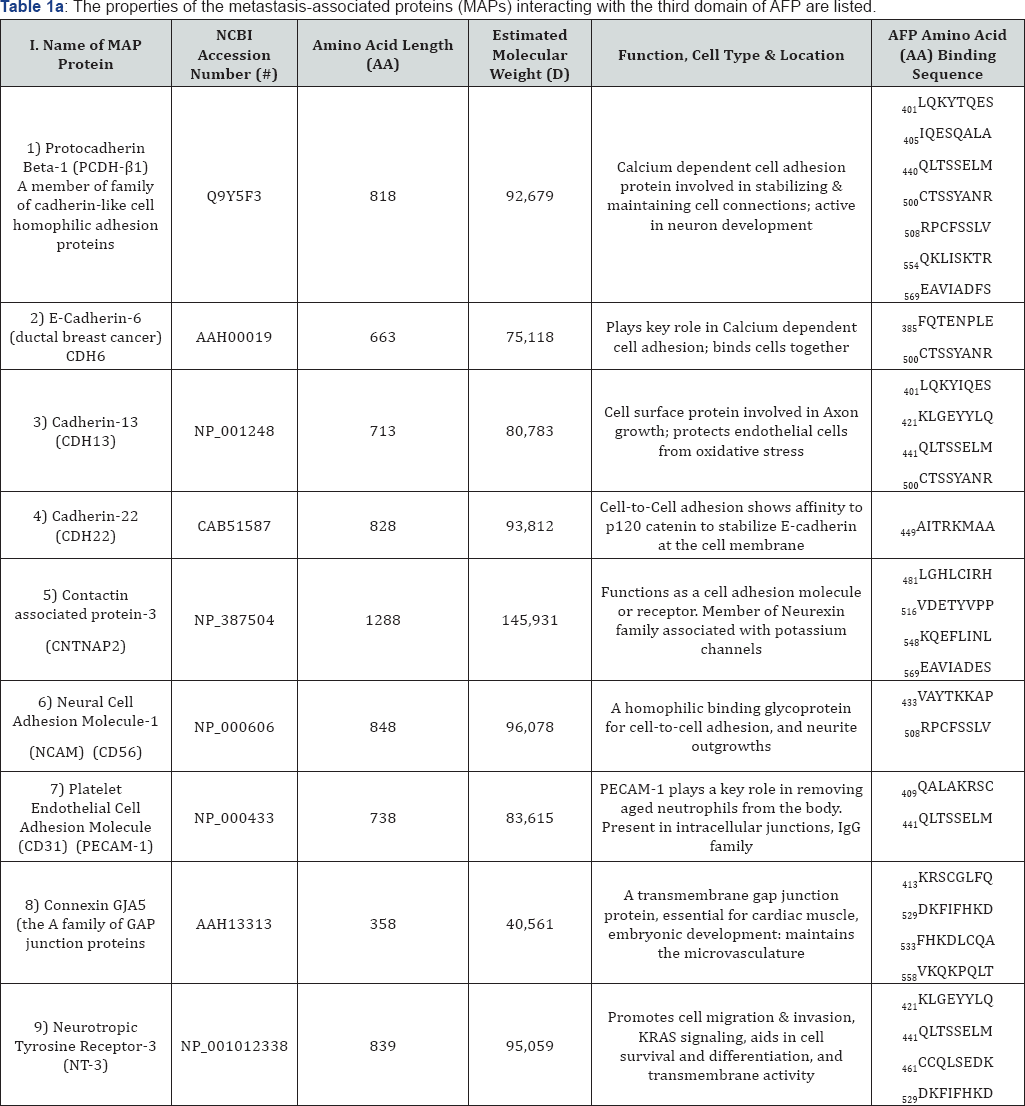
Group # I: Name: Cell Adherence and Cell-to-Cell Contact.
AA: Amino acid; KRAS: Cell Division Regulator Enzyme.
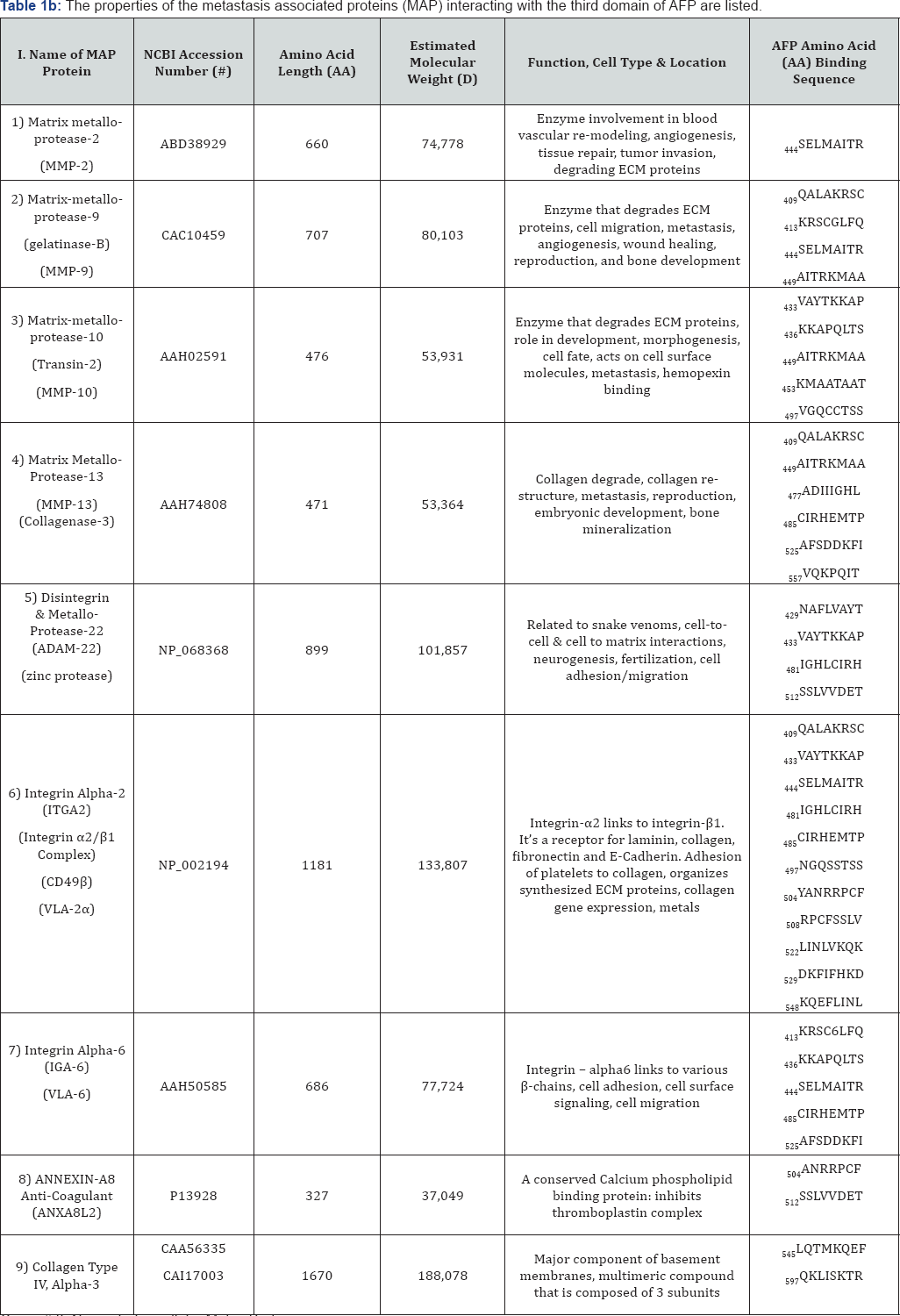
Group # II: Name: Extra-cellular Matrix Proteins.
AA: Amino Acid; ECM: Extra-Cellular Matrix.
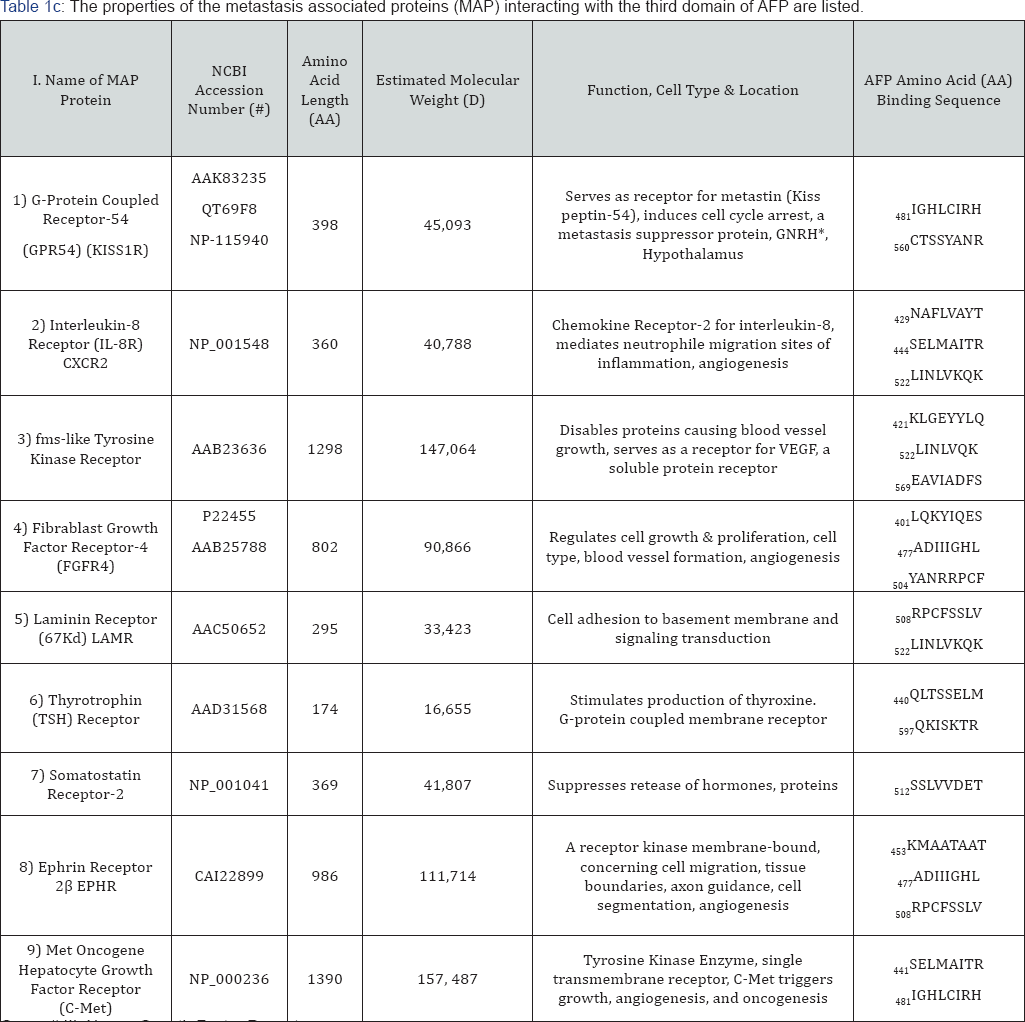
Group # III: Name: Growth Factor Receptors.
*GNRH: Gonadotrophia Releasing hormone; AA: Amino Acid.
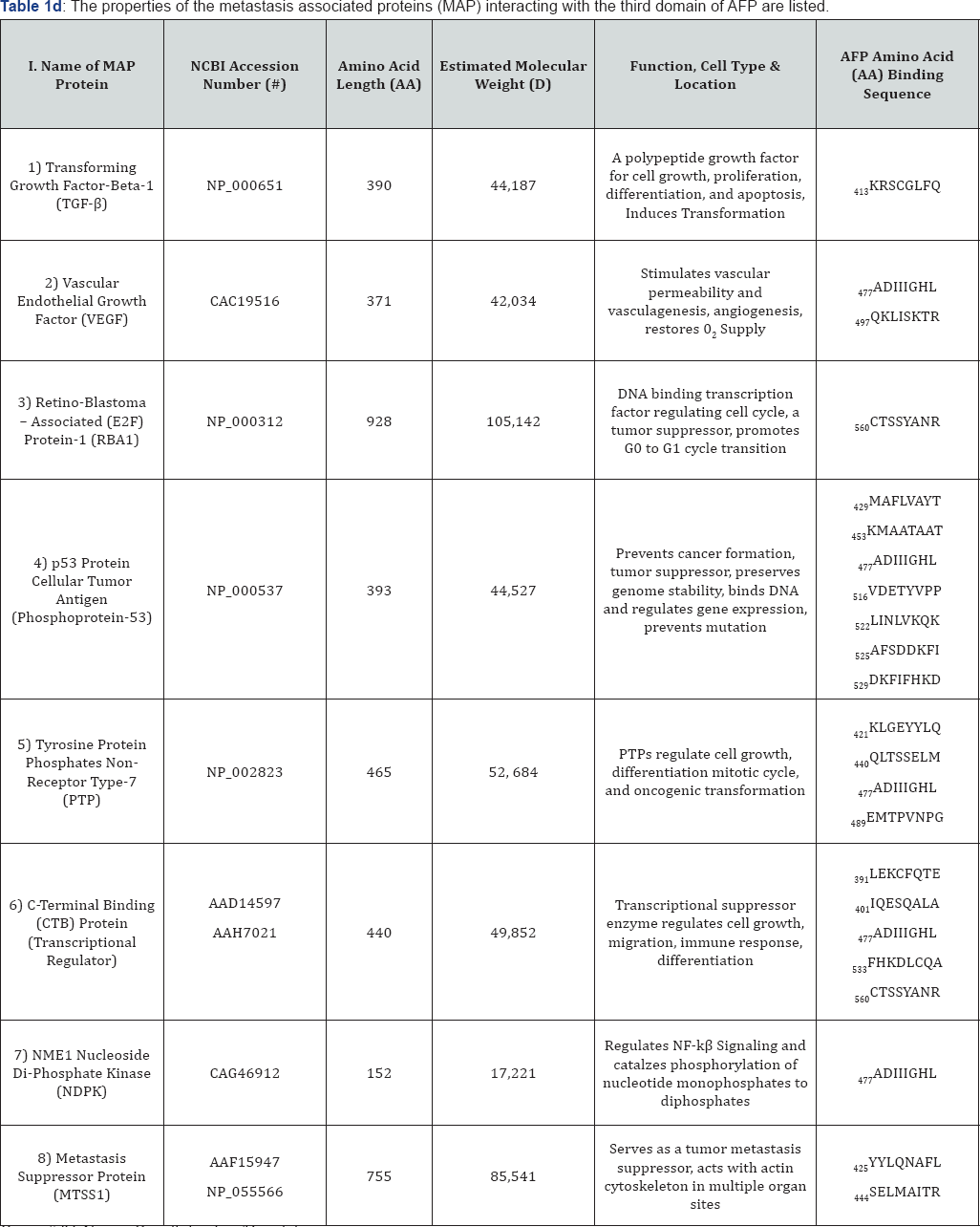
Group # IV: Name: Growth Factors/Regulators.
AA: Amino Acid; NF-kâ: Nuclear Factor B-cell activator.


The cadherins that interact in silico with AFP3D are listed in (Tables 1a-1d), Group 1 and Figures 1A & 1B and included protocadherin p 1 (neural development), E-cadherin-6 (ductal epithelium), cadherin-13 (endothelium) and cadherin-22 (epithelial). Contactin associated protein-3 was also found; such proteins function in concert with potassium channels and are sub-members of the Neurexin family [21]. The neural cell adhesion molecule-1 interaction with AFP also occurred and this protein functions in hemophilic cell-to-cell contact in neurite outgrowths [22]. The platelet endothelial cell adhesion molecule (CD31) also interacts with AFP3D and functions to remove aged white blood cells (i.e. neutrophils) from the bloodstream and intracellular tissue spaces [23]. Another AFP3D interacting adhesion protein found was connexin which plays a role in cell gap junction formation in vascular and heart tissue during embryo/fetal development [24]. Finally, a neurotrophic tyrosine receptor 3 sites was found which is known to function in cell migration following KRAS signaling and cell transmembrane activities (Table 1).
Extracellular Matrix (ECM) Proteins
which participates in collagen degradation. The MMPs are involved in the breakdown of the ECM in physiological events, development, disease, and metastasis. The ADAMS family members are membrane-anchored proteins structurally related to snake venom disintegrins being implicated in cell migration, metastasis, cell adhesion, and cell-to-cell contact during neurogenesis, fertilization, reproduction, muscle development, and integrin binding [26]. The collagenases and gelatinases are sub members of the MMP family involved in the breakdown of ECM proteins for skeletal bone mineralization, tissue remodeling, reproduction, and in diseases such as rheumatoid arthritis and osteoarthritis [27]. Integrins are cell surface transmembrane proteins, consisting of a / p chains, that participate in cell adhesion and cell membrane mediated signaling [28]. Finally, the annexins are Ca++ and phospholipids' binding proteins which function in the blood coagulation cascade [29].
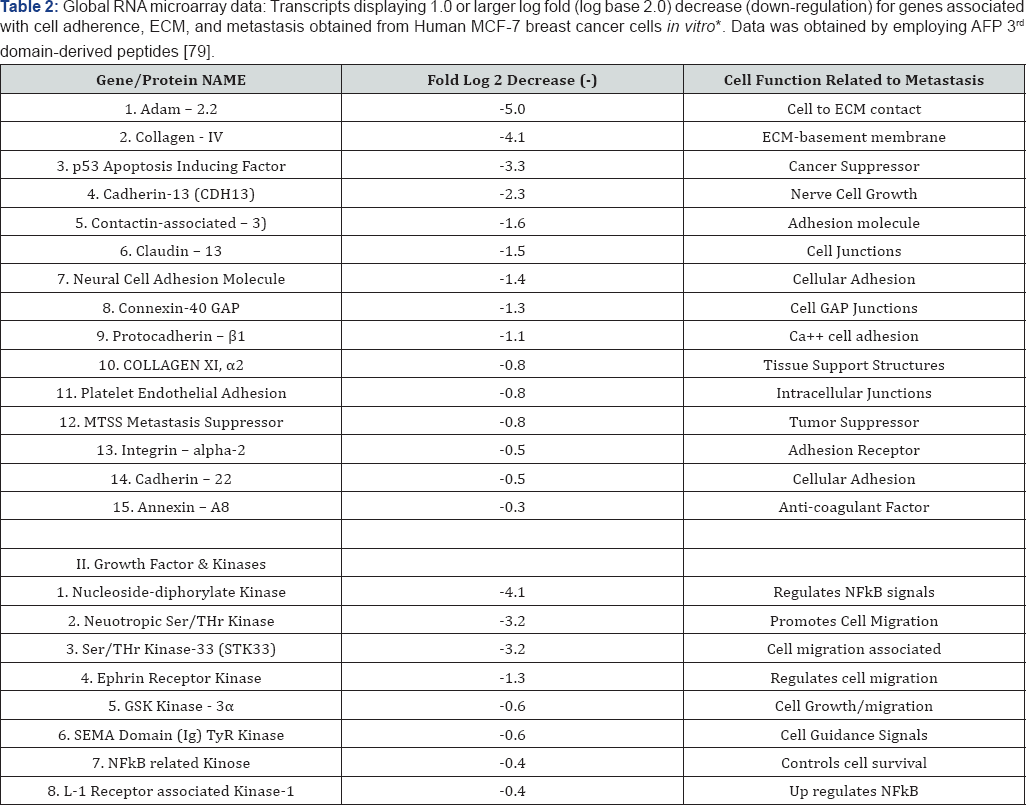
*Expression of 716 transcripts was significantly altered in MCF-7 cells after 8 days of treatment with AFP - derived peptides as compared to treatment with the scrambled peptide. Four hundred thirty RNAs were down regulated, while 286 RNAs were up regulated. Data provided in collaboration with Dr. Kathleen Arcaro, University of Massachusetts, Amherst, MA [32,33]. ECM: Extracellular Matrix.
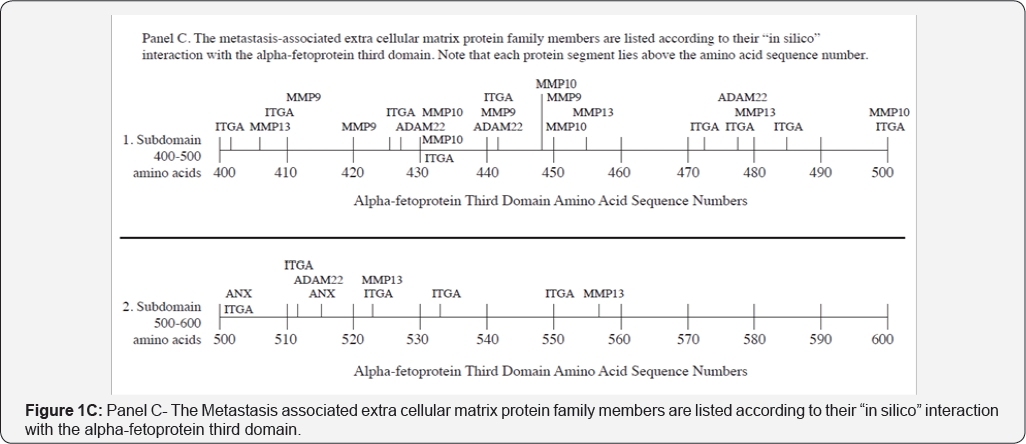
The specific metastasis-associated ECM proteins interacting angiogenesis, tissue repair, and tumor invasion, MMP-9, MMP- with AFP3D in silico are shown in Figure 1C and Table 2, Group 2. 10, and MMP-13 are largely involved with ECM degradation, The MMP family members which were detected included MMP- remodeling, and metastasis. AFP3D was also found to react with 2, MMP-9, MMP-10, and MMP-13 [25]. While MP-2 functions in ADAM-22 which has been implicated with cell migration, steroid responsiveness, myelin and brain development, and cell-to-cell interactions [26]. The integrins interacting with AFP3D included both integrin alpha-2 and integrin alpha-6. Integrin alpha-2 (ITGA2) serves as a receptor for laminin, collagen, collagen C-propeptides, fibronectin, and E-cadherin. ITGA2 is responsible for platelet and multiple cell adhesions utilized in for the tissue organization of collagen and other ECM proteins [29]. Integrin alpha-6 (ITGA2) also interacted with AFP3D in silico; ITGA2 (also known as VLA-6) combines with other integrin members to form heteropartner complexes. These partners participate in both cell adhesion and in cell surface-to-cytoplasmic signal transduction. The annexins are calcium and phospholipid binding proteins which inhibit the formation of the thromboplastin-specific complexes in clot formation [29]. Finally, the collagen type IV alpha-3 protein adds structure and support to tissues such as bone, cartilage, eye, and muscles [30].
The Growth Factor Receptors
The metastasis-associated growth factor receptors (GFRs) include a diverse array of receptor proteins encompassing biological processes such as angiogenesis, growth enhancement and inhibition, kinase activity, mitogenesis, differentiation, and cancer tumorigenesis [31]. Many, if not most, are G-coupled seven-transmembrane receptors with various kinase domains involving intracellular signaling and direct cell-to-cell contact interactions. Their cell surface signaling sets in motion a cascade of downstream signals which ultimately influence, regulate, or mitigate mitogenesis, cell invasion and migration, and metastasis. Some receptors containing tyrosine kinases can serve to activate and/or disable proteins (fMS-like) that cause growth in a variety of tissues including blood vessels [32]. Other such receptors (somatostatin receptor) can inhibit release of many types of hormones and secretory proteins [33]. For example, the ephrin receptor is involved in multiple regulation activities in tissue formation, cell migration, axon guidance, cell- to-cell interactions, stem cell differentiation, and cancer growth. Furthermore, the G-coupled receptor-54 serves as a receptor for the KISS gene product, a metastasis suppressor protein that inhibits metastasis, invasion, and chemotaxis of melanomas and breast cancer [34]. In summary, metastasis-associated growth factor receptors are intimately involved with cell-matrix adhesion, cell invasion and migration, angiogenesis, and cancer growth.
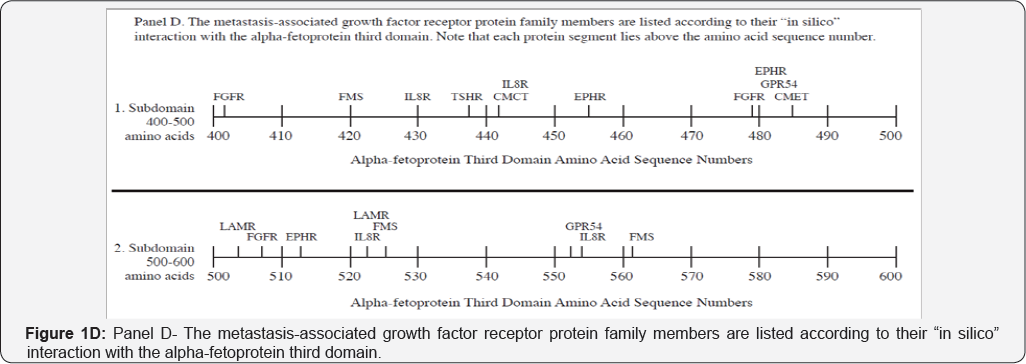
The metastasis-associated growth factor receptors interacting with AFP3D in silico are presented in Figure 1D and in Table 2, Group 3. The GFR group members detected were comprised of G-coupled receptors and other transmembrane proteins such as GPR54 (KISSR), interleukin-8 receptor (IL- 8R), fMS-like tyrosine receptor (fMS-KR), fibroblast growth factor receptor (GFGR4), laminin receptor (LAMR), thyrotrophin receptor (TSHR), somatostatin receptor-2 (SR2), ephrin receptor (EPHR), and c-Met receptor. The KISSR, which is known to interact with AFP3D serves to regulate the KISS tumor suppressor by inhibiting cell proliferation, cell migration, and tumor metastasis [34] . The second GFR to interact with AFP3D was found to be IL- 8R which binds IL-8 with high affinity and transduces a signal through the G-protein activated second messenger pathway [35] . IL-8R can inhibit embryonic oligodendrocyte precursor cell migration in the developing spinal cord and is involved with neutrophil chemotaxis activation. The fMS-KR is a soluble tyrosine kinase enzyme that serves as a receptor of vascular endothelial growth factor (VEGF), a potent angiogenic growth factor; fMS-KR serves to reduce free circulating levels of VEGF[36] .
The FGFR4 receptor, which interacts with AFP3D in silico, is known to bind to and interact with fibroblast growth factor to initiate cell growth and proliferation [37]. FGFR is over expressed in breast and ovarian cancer and other gynecological tumors. The LAMR that interacts with AFP3D is a 37 kD receptor precursor that forms complexes with p40 ribosome chromosome associated protein [38]. LAMR is a multi-functional metastasis- associated protein involved with cell adhesion, invasion and migration. The TSHR is located on thyroid cells and binds the thyroid-stimulating trophic hormone secreted by the pituitary. It stimulates production of thyroid hormone which regulates overall body metabolism [39]. The EPHR is the receptor for the Ephrin protein, serving as a receptor tyrosine kinase to enact cell migration and guidance functions especially in axon growth cones of the brain [40]. Ephrin signaling is also a factor in angiogenesis, stem cell development, and growth of tumors. Finally, the C-met oncogene product is a hepatocyte growth factor receptor tyrosine kinase. C-met receptor plays a role in cell survival, embryogenesis, and cell migration and invasion [41].
Growth Factors and Kinases
The metastasis-associated growth factors (GF) and kinases include multiple and diverse types of kinases involved in angiogenesis, vasculogenesis, oxygen supply to tissues, metastasis suppression, tumor formation, genome mutation, di- and tri-phosphate chemical exchange cell growth and proliferation, development, signal transduction, and cell cycle phase transition. The GFs and kinases localized on AFP3D sequences include transforming growth factor- p 1 (TGF p 1), vascular growth factor (VEGF), retinoblastoma-associated E2F protein-1 (RBA1), p53 phosphoprotein-1 (TB53), tyrosine protein phosphatase type 7 (PTP7), c-terminal binding protein (CTBP), nucleoside di-diphosphate kinase (NDPK), and metastasis suppressor protein (MTSS1). Most of these proteins serve as tyrosine kinase enzymes and are involved in blood vessel growth and cell cycle regulation.
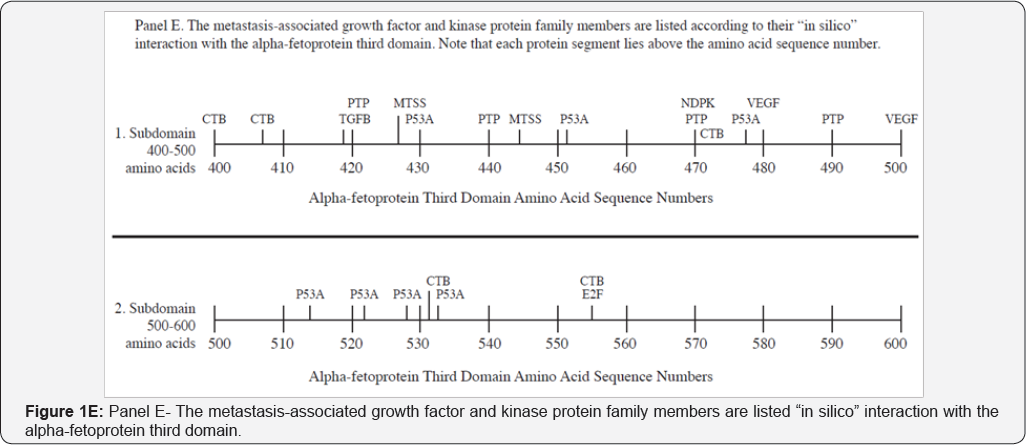
The metastasis-associated growth factors and kinases that interacted with AFP3D in silico are displayed in Figure 1E and on Table 2, Group IV. The GF and kinase protein (i.e. TGF p 1) interaction sites on AFP3D reflect proteins that promote cell growth and proliferation, tumor cell transformation, activation of transcription factors, and regulation of gene expression. It can further activate factors such as interferon- y, tumor necrosis factor-a and multiple cytokines. VEGF is a growth factor for blood vessel formation which regulates vascular permeability [42] . This GF can further serve to restore blood vessel injury and form new vessels to bypass blocked vasculature. The RBP1 is a retinoblastoma-related member of the E2F family of DNA-binding transcription factors, which act as key proteins regulating the cell cycle and tumor suppressor proteins [43]. The RBP-encoded E2Fs are also targets of transforming proteins of small DNA tumor viruses. The TBp53, a cellular tumor antigen, is a phosphoprotein which functions as a tumor suppressor protein providing protection to the genome [44]. Hence, it serves as a conserving stability agent of genes in preventing cancer cell formation. The tyrosine protein phosphatase non-receptor-7 (PTP7) is a signaling molecule that regulates multiple processes including cell growth, differentiation, the mitotic cycle, and cancer cell transformation [45].
PTP7 can interact with lymphokine secreting cells, hematopoietic cells, and exhibits MAP kinase activities. A further protein interacting with AFP3D is the C-terminal binding protein which functions as a regulatory tyrosine kinase [46]. This enzyme phosphorylates the C-terminal ends of Src-family kinases. The nucleotide-di-phosphate kinase (NDPK) that also interacted with AFP3D is an enzyme that catalyzes the exchange of terminal phosphates between different nucleoside di- and triphosphates. NDPK is involved with cell growth and proliferation, development, signal transduction, and G-protein coupled receptor activities. Finally, MTSS1 is a metastasis suppressor protein that interacted with the AFP3D fragment. This protein contains an acting binding segment and is present in various tissues and organs [47].
Reported Interactions of Metastasis-related with Hydrobolic Ligands and Various Receptors/Proteins
The co-localizations of hydrophobic ligands and various receptors/proteins are demonstrated in Figure 1, Panels A-E and have been reported in the literature as described below. The metastasis-related protein cluster site segments detected on AFP3D were identified by amino acid sequence numbers on the AFP polypeptide chain Figure 1A. In Figure 1A, the first metastasis-protein cluster is displayed for AFP3D amino acid (AA) #400-455; the second cluster was found on AA #480 to 500; and the third cluster ranged from AA #500-570 with outliers scattered throughout the third domain. Literature searches described herein revealed that these clustered domain segments may not be found there by mere chance. For example, the hydrophobic ligand binding sub domain in Figure 1A, extends and includes the entire AA lengths of the metastasis-related protein interaction segments on AFP3D. Interestingly, published reports have demonstrated that tumor invasion and metastases are clearly linked to activities of N-3 polyunsaturated fatty acids and estrogens [48-50]. These metastasis-to-hydrophobic linkages are known to involve cell adherence proteins such as cadherins, ADAM proteins, β -catenin, and others [51-53].
In other studies, reports of cation channel influence on metastasis have emerged. Such studies have implicated selective and non-selective cation channels such as transient receptor potential vallinoid (TRPV), voltage-gated sodium channels, ATP- release channels, and gap junctions involving connexin in breast, brain, pancreatic, and lymphoid tumors [54-57]. Additional reports have shown correlations between metastasis and cell cycle proteins such as β -catenin, ubiquitin ligases, F-box proteins, SKP2 proteins, and DNA-PK kinases [58-61]. Chemokine and scavenger receptors (SR) have also been associated with metastasis through proteins such as SR Class B type 1 Lysl oxidase (LOX), CXCR6, CXCL16, SR Class-A, and low-density lipoprotein receptor [62-65]. Lysophospholipid receptors are also associated with the process of metastasis through the sphingo-1-phophate receptor, VEGF receptor, lysophosphatidic acid receptors, Sp1 kinase, and G-coupled protein receptor-55 [66-68]. Finally, the mucins and their receptors have been linked to metastasis via N-cadherin, NCAM, mucin-1, mucin-2, mucin-6, mucin-16, and mucin-5AC [69-71]. In summary, it can be discerned that the ligand and receptor interaction sites on AFP3D clearly correlate in the literature with their respective sub domain localizations displayed in Figure 1A.
The Relationship of AFP3D-localized Amino Acid Segments with Metastasis
A literature survey in Pub Med and other search engines readily reveal that AFP and its derived peptides are directly involved with the metastatic process. AFP serum levels have long been associated as a biomarker for various metastatic tumors. Such cancers include hepatocellular carcinomas, germ cell tumors, and reproductive and gastrointestinal cancers. However, AFP is also a metastatic biomarker for tumors such as gastric cancer, breast tumors, mixed germ cell tumors such as seminomas, and neuroendocrine cancer [72-77]. Thus, full- length AFP has been shown to play a critical role in the metastatic spread of cancer cells.
Literature reports have also employed short peptides (8-34 amino acid) derived from the third domain of human AFP. These reports have experimentally demonstrated that AFP3D-derived peptides do in fact interact with cell adhesion, cell-to-cell contact, receptor, and kinase proteins that clearly influence and directly affect the metastatic process [76-80]. One such study with human follicular thyroid carcinoma (FTC) cells and the AFP- derived "Growth Inhibitory Peptide (GIP) directly demonstrated in vitro that GIP significantly inhibited cell migration and invasion in a dose-dependent fashion [78]. A second study reported that GIP suppressed the spread of tumor infiltrates and metastases in both human and mouse mammary cancers [76,77]. A third report showed that GIP interfered with cell- to-cell contact inhibition, blocked tumor cell adhesion to ECM proteins, and prevented platelet aggregation thus preventing tumor cell spreading/migration and attachment to anchorage surfaces [76].
Furthermore, the AFP-derived GIP peptide has been demonstrated to down-regulate proteins associated with the metastatic process using a mRNA global microarray [79]. As shown in Table 2 (Part I), GIP was found capable of down- regulating the mRNA of at least 15 different cell adherence and ECM proteins associated with metastasis; these included proteins such as ADAM-22, connexin, cadherins, contact in, collagen, and the MTSS metastasis suppressor. Moreover, the mRNA down-regulation (employing a log base 2) in the GIP study ranged downward from a -5.0 fold decrease to -0.3. In addition, AFP-derived GIP down-regulated the mRNA of metastatic growth factors and kinase from a -4.1 decrease to -0.4 (Table 2), Part II. Finally, an enzyme activity screen of the AFP-derived peptide Table 3: The percent of metastasis-associated kinase activity followir 100% and the inhibition or enhancement is listed as percent activity ol (GIP) incubated with metastasis-related kinases representing activities of protein kinase-C, MAP kinases, apoptosis signal kinases, and others, showed significant inhibitions ranging from 45 to 18% (Table 3) Part I. Moderately enhanced kinase activities ranging from 33 to 18% were further displayed in enzymes representative of ephrin, c-Met, fibroblast growth factor and NFkB-related kinases (Table 3) Part II.
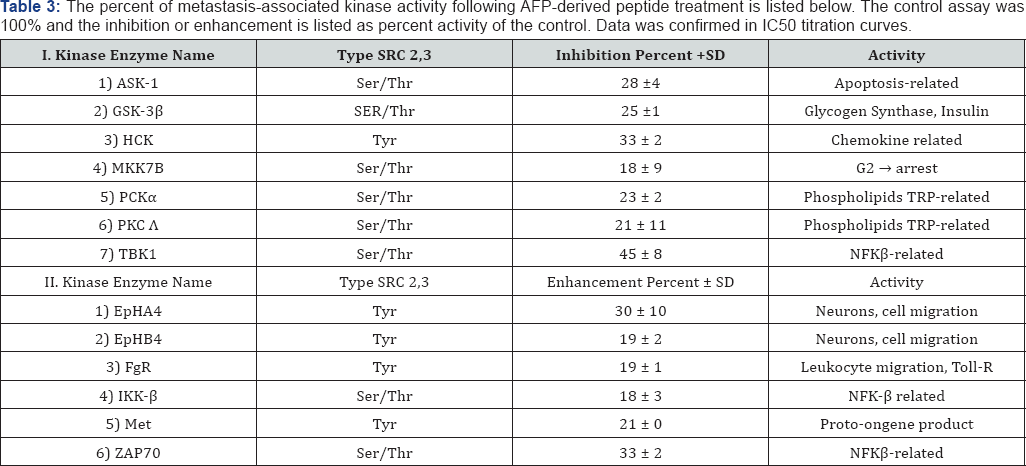
*Ser/Thr: Serine/thyronine kinase; Tyr: Tyrosine kinase
c-Src: A Non-Receptor kinase protein of the ser/Thr or tyrosine type that phosphorylates these residues in other proteins. The kinase activity screen for AFP-3D peptides was performed via the commercial “kinase profiler” by the Upstate Biosignaling Corp., Dundee Technology Park, Dundee, United Kingdom
Conclusion
Many previous reports have now documented in both computer-based and experimentally-verified studies that the AFP3D is comprised of short amino acid sequence that can associate with hydrophobic ligands and receptor/proteins [11-15]. It is now known that AFP3D amino acid sequences are capable of interacting with fatty acids, steroids, retinoids, ligands, and with various proteins including scavenger, mucin, chemokine, cation channel receptors, and cell cycle proteins. From Figure 1, it can be deduced that these ligands, receptors, and proteins appear to occupy and/or overlap various amino acid sequence sites. It is tempting to speculate that these compounds could act in combination with, or in competition with each other. AFP binding at overlapping sites in the case of fatty acids, estrogens, and retinoids have long been known and reported [1,2]. Thus, it would not be unreasonable to assume that competition of proteins to occupy binding/docking sites on AFP3D might occur. Presently, the metastasis interaction sites detected on AFP3D could also be subject to interference or conformational changes by ligands/proteins previously localized to those same sites. The literature reviews stated in this report confirmed that multiple metastatic proteins can and do interact with many of the ligands/proteins listed above. Therefore, it is plausible that co-localization of metastatic proteins with the multiple other ligands/proteins described herein, may be physiologically relevant.
It was further shown and addressed in the present report that metastasis-related proteins are indeed involved with cell adherence, cell-to-cell contact, and the extracellular matrix. These associations indicate that growth factors, and their receptors and kinases are involved in order to achieve the completed process of tumor cell detachment. Once tumor cells are separated and disseminated from the tumor mass, the segregated tumor cells must migrate and digest their way through various ECM environments and basement membranes to gain entrance and transpassage across the endothelial cells into the lumen of the blood vasculature and lymphatic ducts. The extravasations of tumor cells to distant sites occur only after the loss of cell adhesion and release of protolytic enzymes to digest a tunnel through a myriad of tissue membrane barriers. After the tumor cells have egressed through the blood vessels, they can attach, bind, and cluster with circulating platelets to ensure a shielded passage while traveling through the blood vessels [81,82]. By means of the process of chemotaxis, tumor cells are attracted to distal sites of filtration organs such as liver, lungs, kidneys, and bone marrow.
The blood vessel or lymphatic transpassage is again repeated and the tumor cells finally become "nested" within the stromal tissue of the host organ [82]. It becomes obvious that AFP3D interactions with the metastatic proteins described in this report can play a pivotal role in the enhancement or inhibition of the multi-stage process of guiding metastatic cells to distal sites. As demonstrated in Tables 2 & 3, certain AFP-derived peptide segments are capable of up- and down-regulation of the mRNA of various metastatic proteins with the subsequent inhibition or enhancement of the relevant metastatic kinases.
Conflict of Interest
No US federal grants were used there are no known conflicts of interest in the preparation of this manuscript.
References
- Mizejewski GJ (2001) Alpha-fetoprotein structure and function: relevance to isoforms, epitopes, and conformational variants. Exp Biol Med 226(5): 377-408.
- Mizejewski GJ (1997) Alpha-fetoprotein as a biologic response modifier: relevance to domain and subdomain structure. Proceedings of the Society for Experimental Biology and Medicine Society for Experimental Biology and Medicine 215(4): 333-362.
- Naidu S, Peterson ML and Spear BT (2010) Alpha-fetoprotein related gene (ARG): a new member of the albumin gene family that is no longer functional in primates. Gene 449(1-2): 95-102.
- Mizejewski GJ (2004) Biological roles of alpha-fetoprotein during pregnancy and perinatal development. Experimental biology and medicine 229(6): 439-463.
- Mizejewski GJ (2002) Biological role of alpha-fetoprotein in cancer: prospects for anticancer therapy. Expert Rev Anticancer Ther 2(6): 709-735.
- Posypanova GA, Gorokhovets NV, Makarov VA, Savvateeva LV, Kireeva NN, et al. (2008) Recombinant alpha-fetoprotein C-terminal fragment: the new recombinant vector for targeted delivery. J Drug Target 16(4): 321-328.
- Posypanova GA, Makarov VA, Savvateeva MV, Bereznikova AV, Severin ES. (2013) The receptor binding fragment of alpha-fetoprotein is a promising new vector for the selective delivery of antineoplastic agents. J Drug Target 21(5): 458-465.
- Atemezem A, Mbemba E, Marfaing R, Vaysse J, Pontet M, et al. (2002) Human alpha-fetoprotein binds to primary macrophages. Biochemical and biophysical research communications 296(3): 507-514.
- Aussel C, Masseyeff R (1984) Interaction of retinoids and bilirubin with the binding of arachidonic acid to human alpha-fetoprotein. Biochemical and biophysical research communications 119(3): 11221127.
- Benassayag C, Savu L, Vallette G, Delorme J, Nunez EA (1979) Relations between fatty acids and oestrogen binding properties of pure rat alpha 1-foetoprotein. Biochimica et biophysica acta 587(2): 227-237.
- Mizejewski GJ (2015) Nonsecreted cytoplasmic alpha-fetoprotein: a newly discovered role in intracellular signaling and regulation. An update and commentary. Tumour biology the journal of the International Society for Oncodevelopmental Biology and Medicine 36(12): 9857-9864.
- Mizejewski GJ (2016) The alpha-fetoprotein (AFP) third domain: a search for AFP interaction sites of cell cycle proteins. Tumour biology -the journal of the International Society for Oncodevelopmental Biology and Medicine 37(9): 12697-12711.
- Mizejewski GJ (2017) Review of the Third Domain Receptor Binding Fragment of Alpha-fetoprotein (AFP): Plausible Binding of AFP to Lysophospholipid Receptor Targets. Current Drug Targets 18(7): 874886.
- Mizejewski GJ (2015) The alpha-fetoprotein third domain receptor binding fragment: in search of scavenger and associated receptor targets. Journal of drug targeting 23(6): 538-551.
- Mizejewski GJ (2013) Review of the adenocarcinoma cell surface receptor for human alpha-fetoprotein; proposed identification of a widespread mucin as the tumor cell receptor. Tumour biology: the journal of the International Society for Oncodevelopmental Biology and Medicine 34(3): 1317-1336.
- Mizejewski GJ (2016) The third domain fragments of alpha-fetoprotein (AFP): Mapping AFP interaction sites with selective and non-selective cation channels. Current Topics in Peptide & Protein Research 16: 6382.
- Carter DC, He XM, Munson SH, Twigg PD, Gernert KM, et al. (1989) Three-dimensional structure of human serum albumin. Science 244(4909): 1195-1198.
- Osmond RI, Das S, Crouch MF (2010) Development of cell-based assays for cytokine receptor signaling, using an AlphaScreen SureFire assay format. Analytical biochemistry 403(1-2): 94-101.
- Hulpiau P, Van Roy F (2009) Molecular evolution of the cadherin superfamily. The international journal of biochemistry & cell biology 41(2): 349-369.
- Angst BD, Marcozzi C, Magee AI (2001) The cadherin superfamily: diversity in form and function. Journal of cell science 114: 629-641.
- Qiao YQ, Huang ML, Zheng Q, Wang TR, An Tao Xu, et al. (2015) CNTNAP3 associated ATG16L1 expression and Crohn's disease. Mediators of inflammation 2015: 404185.
- Suzuki M, Angata K, Nakayama J, Fukuda M (2003) Polysialic acid and mucin type o-glycans on the neural cell adhesion molecule differentially regulate myoblast fusion. J Biol chem 278(49): 49459-49468.
- Newman PJ, Newman DK (2003) Signal transduction pathways mediated by PECAM-1: new roles for an old molecule in platelet and vascular cell biology. Arterioscler Thromb Vasc Biol 23(6): 953-964.
- Maeda S, Nakagawa S, Suga M, Yamashita E, Oshima A, et al. (2009) Structure of the connexin 26 gap junction channel at 3.5 A resolution. Nature 458(7238): 597-602.
- Nagase H, Woessner JF, Jr (1999) Matrix metalloproteinase's. The Journal of biological chemistry 274(31): 21491-21494.
- Zhu P, Sun Y, Xu R, Sang Y, Zhao J, et al. (2003) The interaction between ADAM 22 and 14-3-3zeta: regulation of cell adhesion and spreading. Biochem Biophys Res commun 301(4): 991-999.
- Collier IE, Bruns GA, Goldberg GI, Gerhard DS (1991) On the structure and chromosome location of the 72- and 92-kDa human type IV collagenase genes. Genomics 9(3): 429-434.
- White DJ, Puranen S, Johnson MS, Heino J (2004) The collagen receptor subfamily of the integrins. The international journal of biochemistry & cell biology 36(8): 1405-1410.
- Hauptmann R, Maurer-Fogy I, Krystek E, Bodo G, Andree H, et al. (1989) Vascular anticoagulant beta: a novel human Ca2+/phospholipid binding protein that inhibits coagulation and phospholipase A2 activity. Its molecular cloning, expression and comparison with VAC- alpha. European journal of biochemistry/FEBS 185(1): 63-71.
- Mariyama M, Leinonen A, Mochizuki T, Tryggvason K, Reeders ST, et al. (1994) Complete primary structure of the human alpha 3(IV) collagen chain. Coexpression of the alpha 3(IV) and alpha 4(IV) collagen chains in human tissues. The Journal of biological chemistry 269(37): 2301323017.
- Ornitz DM, Xu J, Colvin JS, McEwen DG, MacArthur CA, et al. (1996) Receptor specificity of the fibroblast growth factor family. J Biol Chem 271(25): 15292-15297.
- Dokanehiifard S, Yasari A, Najafi H, Jafarzadeh M, Nikkhah M, et al. (2017) A novel microRNA located in the TrkC gene regulates the Wnt signaling pathway and is differentially expressed in colorectal cancer specimens. The Journal of biological chemistry 292(18): 7566-7577.
- Yamada Y, Post SR, Wang K, Tager HS, G I Bell, et al. (1992) Cloning and functional characterization of a family of human and mouse somatostatin receptors expressed in brain, gastrointestinal tract, and kidney. Proceedings of the National Academy of Sciences of the United States of America 89(1): 251-255.
- Ohtaki T, Shintani Y, Honda S, Matsumoto H, et al. (2001) Metastasis suppressor gene KiSS-1 encodes peptide ligand of a G-protein-coupled receptor. Nature 411(6837): 613-617.
- Lee J, Horuk R, Rice GC, Bennett GL, Camerato T, et al. (1992) Characterization of two high affinity human interleukin-8 receptors. The Journal of biological chemistry 267(23): 16283-16287.
- Maynard SE, Min JY, Merchan J, Lim KH, Jianyi Li, et al. (2003) Excess placental soluble fms-like tyrosine kinase 1 (sFlt1) may contribute to endothelial dysfunction, hypertension, and proteinuria in preeclampsia. The Journal of clinical investigation 111(5): 649-658.
- Ron D, Reich R, Chedid M, Lengel C, Cohen OE, et al. (1993) Fibroblast growth factor receptor 4 is a high affinity receptor for both acidic and basic fibroblast growth factor but not for keratinocyte growth factor. The Journal of biological chemistry 268(8): 5388-5394.
- Jackers P, Minoletti F, Belotti D, Clausse N, Sozzi G, et al. (1996) Isolation from a multigene family of the active human gene of the metastasis- associated multifunctional protein 37LRP/p40 at chromosome 3p21.3. Oncogene 13(3): 495-503.
- Szkudlinski MW, Fremont V, Ronin C, Weintraub BD (2002) Thyroid- stimulating hormone and thyroid-stimulating hormone receptor structure-function relationships. Physiological reviews 82(2): 473502.
- Kullander K, Klein R (2002) Mechanisms and functions of Eph and ephrin signalling. Nature reviews Molecular cell biology 3(7): 475-486.
- Bottaro DP, Rubin JS, Faletto DL, Chan AM, Kmiecik TE, et al. (1991) Identification of the hepatocyte growth factor receptor as the c-met proto-oncogene product. Science 251(4995): 802-804.
- Muller YA, Li B, Christinger HW, Wells JA, Cunningham BC, et al. (1997) Vascular endothelial growth factor: crystal structure and functional mapping of the kinase domain receptor binding site. Proceedings of the National Academy of Sciences of the United States of America 94(14): 7192-7197.
- Merten L, Agaimy A, Moskalev EA, Giedl J, Kayser C, et al. (2016) Inactivating Mutations of RB1 and TP53 Correlate With Sarcomatous Histomorphology and Metastasis/Recurrence in Gastrointestinal Stromal Tumors. American journal of clinical pathology 146(6): 718726.
- Isobe M, Emanuel BS, Givol D, Oren M, Croce CM, et al. (1986) Localization of gene for human p53 tumour antigen to band 17p13. Nature 320(6057): 84-85.
- Critton DA, Tortajada A, Stetson G, Peti W, Page R, et al. (2008) Structural basis of substrate recognition by hematopoietic tyrosine phosphatase. Biochemistry 47(50): 13336-13345.
- Bergman M, Mustelin T, Oetken C, Partanen J, Flint NA, et al. (1992) The human p50csk tyrosine kinase phosphorylates p56lck at Tyr-505 and down regulates its catalytic activity. The EMBO journal 11(8): 29192924.
- Kauffman EC, Robinson VL, Stadler WM, Sokoloff MH, Rinker- Schaeffer CW, et al. (2003) Metastasis suppression: the evolving role of metastasis suppressor genes for regulating cancer cell growth at the secondary site. The Journal of urology 169(3): 1122-1133.
- Belkaid A, Ouellette RJ, Surette ME (2017) 17beta-estradiol-induced ACSL4 protein expression promotes an invasive phenotype in estrogen receptor positive mammary carcinoma cells. Carcinogenesis 38(4): 402-410.
- Yin X, Yu XW, Zhu P, Zhang YM (2016) Endogenously synthesized n-3 fatty acids in fat-1 transgenic mice prevent melanoma progression by increasing E-cadherin expression and inhibiting beta-catenin signaling. Molecular medicine reports 14(4): 3476-3484.
- Jiang WG, Sanders AJ, Katoh M, Ungefroren H, Gieseler F, et al. (2015) Tissue invasion and metastasis: Molecular, biological and clinical perspectives. Seminars in cancer biology 35 Suppl: S244-S275.
- Kossmann CM, Annereau M, Thomas-Schoemann A, Nicco-Overney C, Christiane Chereau, et al. (2017) ADAM9 expression promotes an aggressive lung adenocarcinoma phenotype. Tumour biology: the journal of the International Society for Oncodevelopmental Biology and Medicine 39(7): 1010428317716077.
- Serini S, Zinzi A, Ottes Vasconcelos R, Fasano E, Riillo MG, et al. (2016) Role of beta-catenin signaling in the anti-invasive effect of the omega-3 fatty acid DHA in human melanoma cells. J Dermatol Sci 84(2): 149159.
- Hartmann F, Sparla D, Tute E, Tamm M, Schneider U, et al. (2017) Low acyl-CoA synthetase 5 expression in colorectal carcinomas is prognostic for early tumour recurrence. Pathol Res Pract 213(3): 261266.
- Nakanishi M, Morita Y, Hata K, Muragaki Y (2016) Acidic microenvironments induce lymphangiogenesis and IL-8 production via TRPV1 activation in human lymphatic endothelial cells. Experimental cell research 345(2): 180-189.
- Klumpp L, Sezgin EC, Eckert F, Huber SM (2016) Ion Channels in Brain Metastasis. Int J Mol Sci 17(9). pii: E1513.
- Yee NS (2016) TRPM8 Ion Channels as Potential Cancer Biomarker and Target in Pancreatic Cancer. Adv protein chem struct Biol 104: 127155.
- Martial S (2016) Involvement of ion channels and transporters in carcinoma angiogenesis and metastasis. American journal of physiology Cell physiology 310(9): C710-C27.
- Pascale RM, Joseph C, Latte G, Evert M, Francesco Feo, et al. (2016) DNA-PKcs: A promising therapeutic target in human hepatocellular carcinoma? DNA Repair (Amst) 47: 12-20.
- Milano DF, Ngai NA, Muthuswamy SK, Asthagiri AR (2016) Regulators of Metastasis Modulate the Migratory Response to Cell Contact under Spatial Confinement. Biophysical journal 110(8): 1886-1895.
- Chen J, Rajasekaran M, Xia H, Zhang X, Kong SN, et al. (2016) The microtubule-associated protein PRC1 promotes early recurrence of hepatocellular carcinoma in association with the Wnt/beta-catenin signalling pathway. Gut 65(9): 1522-1534.
- Zheng N, Zhou Q, Wang Z, Wei W (2016) Recent advances in SCF ubiquitin ligase complex: Clinical implications. Biochimica et biophysica acta 1866(1): 12-22.
- Yuan B, Wu C, Wang X, Wang D, Liu H, et al. (2016) High scavenger receptor class B type I expression is related to tumor aggressiveness and poor prognosis in breast cancer. Tumour biology: the journal of the International Society for Oncodevelopmental Biology and Medicine 37(3): 3581-3588.
- Singh R, Kapur N, Mir H, Singh N, James W Lillard J, et al. (2016) CXCR6- CXCL16 axis promotes prostate cancer by mediating cytoskeleton rearrangement via Ezrin activation and alphavbeta3 integrin clustering. Oncotarget 7(6): 7343-7353.
- Hua YJ, Wang HY, Tang LQ, Chen QY, et al. (2016) LOX expression in primary nasopharyngeal carcinoma: correlation with prognostic parameters and outcome. Oncotarget 7(7): 8200-8207.
- Yeung OW, Lo CM, Ling CC, Qi X, Geng W, et al. (2015) Alternatively activated (M2) macrophages promote tumour growth and invasiveness in hepatocellular carcinoma. Journal of hepatology 62(3): 607-616.
- Pal SK, Drabkin HA, Reeves JA, Hainsworth JD, Hazel SE, et al. (2017) A phase 2 study of the sphingosine-1-phosphate antibody sonepcizumab in patients with metastatic renal cell carcinoma. Cancer 123(4): 576582.
- Lee BH, Choi SH, Kim HJ, Jung SW, Kim HK, et al. (2016) Plant Lysophosphatidic Acids: A Rich Source for Bioactive Lysophosphatidic Acids and Their Pharmacological Applications. Biol Pharm Bull 39(2): 156-162.
- Kargl J, Andersen L, Hasenohrl C, Feuersinger D, Stancic A, et al. (2016) GPR55 promotes migration and adhesion of colon cancer cells indicating a role in metastasis. British journal of pharmacology 173(1): 142-154.
- Qu Y, Zhao D, Mu J, Che N, Zhang C, et al. (2016) Prognostic analysis of primary mucin-producing adenocarcinoma of the lung: a comprehensive retrospective study. Tumour biology : the journal of the International Society for Oncodevelopmental Biology and Medicine 37(1): 887-896.
- Nguyen Hoang S, Liu Y, Xu L, Chang Y, Lin Zhou, et al. (2016) High mucin-7 expression is an independent predictor of adverse clinical outcomes in patients with clear-cell renal cell carcinoma. Tumour biology: the journal of the International Society for Oncodevelopmental Biology and Medicine 37(11): 15193-15201.
- Wang H, Li WY, Kuo YJ, Yeh YC, (2016) Primary pulmonary hyalinising clear cell carcinoma with mucin production and delayed metastases after 16 years. Pathology 48(5): 518-521.
- Lu S, Ma Y, Sun T, Ren R, (2016) Expression of alpha-fetoprotein in gastric cancer AGS cells contributes to invasion and metastasis by influencing anoikis sensitivity. Oncology reports 35(5): 2984-2990.
- Lu Y, Zhu M, Li W, Lin B, Dong X, et al. (2016) Alpha fetoprotein plays a critical role in promoting metastasis of hepatocellular carcinoma cells. J Cell Mol Med 20(3): 549-558.
- Jin J, Niu X, Zou L, Li L, Li S, et al. (2016) AFP mRNA level in enriched circulating tumor cells from hepatocellular carcinoma patient blood samples is a pivotal predictive marker for metastasis. Cancer letters 378(1): 33-37.
- Iwatsuki S, Naiki T, Kawai N, Etani T, Keitaro Iida, et al. (2016) Nonpalpable testicular pure seminoma with elevated serum alpha- fetoprotein presenting with retroperitoneal metastasis: a case report. Journal of medical case reports 10(1): 114.
- Mizejewski GJ, Butterstein G (2006) Survey of functional activities of alpha-fetoprotein derived growth inhibitory peptides: review and prospects. Current protein & peptide science 7(1): 73-100.
- Muehlemann M, Miller KD, Dauphinee M, Mizejewski GJ (2005) Review of Growth Inhibitory Peptide as a biotherapeutic agent for tumor growth, adhesion, and metastasis. Cancer metastasis reviews 24(3): 441-467.
- Hua SC, Chen SY, Lu CH, Kao YT, Yu HI, et al. (2010) The effects of growth inhibitory peptide on follicular thyroid cancer cell growth, migration, and invasion. Tumori 96(3): 448-451.
- Mizejewski GJ (2011) Mechanism of Cancer Growth Suppression of Alpha-Fetoprotein Derived Growth Inhibitory Peptides (GIP): Comparison of GIP-34 versus GIP-8 (AFPep). Updates and Prospects. Cancers 3(2): 2709-2733.
- Mizejewski GJ, Muehlemann M, Dauphinee M (2006) Update of alpha fetoprotein growth-inhibitory peptides as biotherapeutic agents for tumor growth and metastasis. Chemotherapy 52(2): 83-90.
- Mizejewski GJ (1999) Role of integrins in cancer: survey of expression patterns. Proceedings of the Society for Experimental Biology and Medicine Society for Experimental Biology and Medicine 222(2): 124138.
- Tran-Thanh D, Done SJ (2010) The role of stromal factors in breast tumorigenicity. The American journal of pathology 176(3): 1072-1074.






























