Breast Cancer Detection: Current Methods and Roadmap to Personalized Medicine
*Snijesh VP and Manoj R Kumar
Innov4Sight Health and Biomedical Systems Private Limited, India
Submission: June 30, 2017; Published: July 03, 2017
*Corresponding author: Snijesh VP, Innov4Sight Health and Biomedical Systems Private Limited, Jayanagar, Bengaluru, Karnataka, India-560041, Email: snijesh@gmail.com
How to cite this article: Snijesh VP, Manoj R.Breast Cancer Detection: Current Methods and Roadmap to Personalized Medicine. Canc Therapy & Oncol Int J. 2017; 5(5): 555672. DOI: 10.19080/CTOIJ.2017.05.555672
Abstract
Breast cancer is characterized by uncontrolled growth of malignant cells in the mammary epithelial tissue. The etiology of breast cancer is poorly understood and in spite of substantial therapeutic advances, breast cancer treatment remains as an unsolved issue. Though few genetic markers for hereditary breast cancer have been identified, the etiology of sporadic breast cancer remains poorly understood. These demands for systematic and reliable approaches to investigate the molecular mechanisms underlying the disease and identify biomarkers for better prediction of prognosis, and response to therapies. In this mini review, we highlight conventional methods used for breast cancer diagnosis and significance of personalized medicine in the cancer treatment.
Keywords: Breast Cancer; Hormone Receptor Test; Cancer Genome; Personalized Medicine
Abbreviations: HRT: Hormone Receptor Test; IHC: Immuno Histo Chemistry; FISH: Fluorescence In Situ Hybridization; HER2: Human Epidermal Growth Factor Receptor 2; ER: Estrogen Receptor; PR: Progesterone Receptor; NGS: Next Generation Sequencing
Mini Review
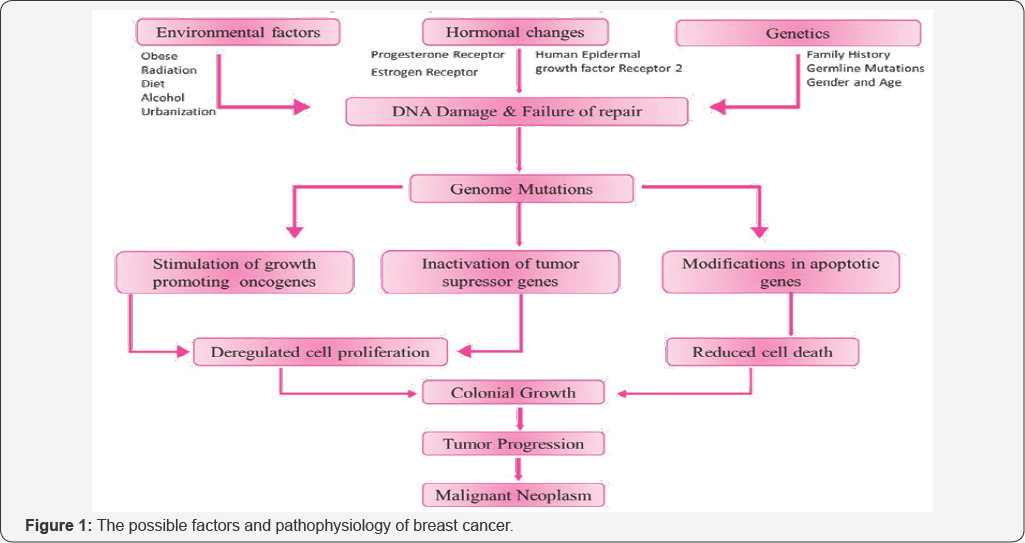
Breast cancer, the most commonly diagnosed cancer and leading cause of death in women, is a multifaceted disease characterized by abnormal and uncontrolled growth of malignant cells in the mammary epithelial tissue [1-3]. Approximately 1.67 million new cases and 522,000 deaths were reported worldwide, making breast cancer one among the leading source of cancer death. The exact roots of breast cancer are poorly known, but both genetic and environmental factors are involved. The possible factors and pathophysiology of breast cancer are depicted in the (Figure 1).
Generally, breast cancer progresses in cells from the lining of milk ducts and lobules that supply ducts with milk. Cancer progressing from duct and lobules are respectively known as ductal and lobular carcinomas. The present understanding of breast cancer pathogenesis is that invasive cancers ascend from a sequence of molecular modifications at the cell level. These modifications effect in breast epithelial cells with immortal features and uncontrolled growth. It encompasses of several biologically altered entities with distinct pathological features and clinical effects which makes breast cancer a complex disease that challenge to accurate diagnosis [4,5]. Recent evidences have proposed that breast cancers with different histopathological and biological features display distinct activities that lead to different treatment responses and should be given different therapeutic strategies [6,7]. In spite of substantial therapeutic developments, breast cancer treatment continues as an unsolved issue. Hence, there is a great need to understand the pathophysiology of breast cancer and to identify additional biomarkers for better prediction of prognosis and response to therapies. Therefore, systematic and reliable approaches has to be introduced to investigate the molecular mechanisms underlying the disease.
Current conventional methods employed for the detection of breast cancer is hormone receptor test (HRT), genetic test and gene panel tests. Most of these methods are used to detect recurrence of breast cancer. The only test which predicts the chance of hereditary breast cancer is BRCA1 and BRCA2 test.
Hormone Receptor Test
Understanding breast cancer cells have hormone receptors or not are a key piece of evidence for making treatment judgements. Largely used HRTs are ImmunoHistoChemistry (IHC) and Fluorescence In Situ Hybridization (FISH). IHC test is used to detect whether or not the cells have hormone receptors like Human Epidermal growth factor Receptor 2 (HER2), Estrogen Receptor (ER) and Progesterone Receptor (PR). FISH identifies specific region in the genome of person's cell. FISH in breast cancer is used to detect the over expression or extra copies of the HER2 genes. FISH is considered as more accurate than IHC for checking presence of HER2 [8].
Genetic Test
The well-known genes that can mutate and raise the breast cancer risk are BRCA1 and BRCA2. Women who carry mutations in the aforesaid genes have higher risk of developing breast cancer than average risk [9]. Genetic tests are carried out on these genes to detect whether the patient carry mutated genes from their parents. Genetic test of BRCA1 and BRCA2 is only established test which predicts chance of occurrence of breast cancer in a person. Rest of the tests are mainly focused on recurrence and TNM staging of breast cancer.
Gene Panels
Research evidences suggest that certain altered genes may increase the risk of developing different types of cancer. The genetic profile of a tumor supports in taking therapeutic decisions and establishing more specific prognosis. Gene panel test, simultaneous examination of many genes, is employed on the people who are genetically predisposed to cancer. The most widely used gene panel is MammaPrint [10], Oncotype DX [11] and Breast Next [12]. MammaPrint is a predictive diagnostic test early stage breast cancer based on 70 breast cancer gene signature which gives the results as high or low risk classification whereas Oncotype DX analyzes the expression of 21 genes and predicts likelihood of disease recurrence in people with ER+ breast cancer. The gene panel, BreastNext simultaneously analyzes 17 genes associated with increased risk for breast cancer, including BRCA1 and BRCA2.
Roadmap to Personalized Medicine
The aforementioned tests are mainly focused on recurrence of breast cancer. The only test that predicts the early breast cancer is BRCA1 and BRCA2 genetic test. But, most people who develop breast cancer have no family history of the disease and the genetic tests are done only for germline mutations which are carried from generation to generation. Till date, no methods are available to detect sporadic breast cancer which may arise due to somatic mutations or other environmental factors. Next Generation Sequencing (NGS), provides powerful platform to detect the alterations of genes in the cancer genome and is a key backbone of personalized medicine. The proposed workflow for personalized medicine is depicted in the (Figure 2).
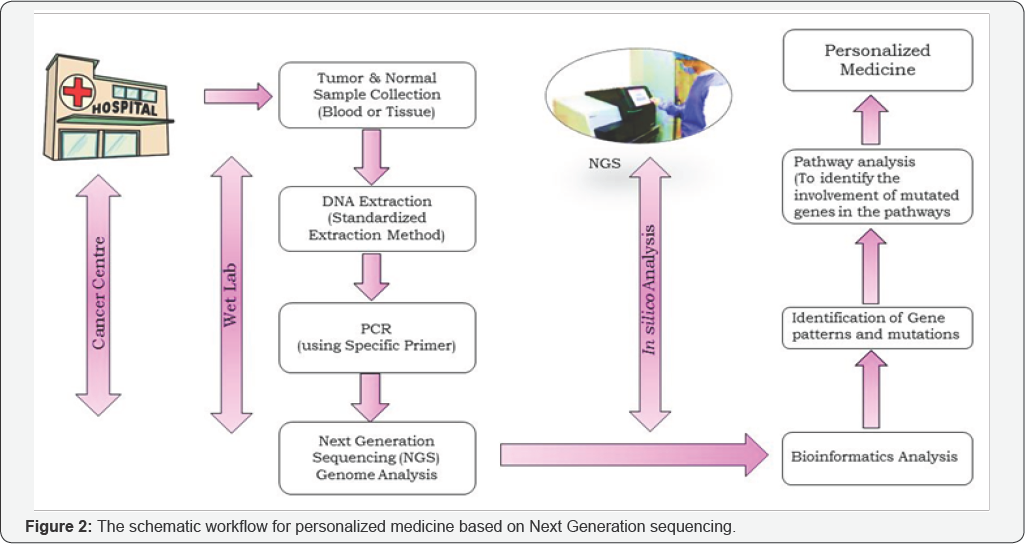
For clinical significance, initial step is to extract most frequently mutated genes in a population. Next, these mutations in a sample can be identified using NGS and Bioinformatics analytical methods. The role of altered genes or mutated genes has to be investigated comprehensively to predict the expected treatment for the patient. For example, if a gene is found to be mutated which increases the risk factor of breast cancer; the individual can change lifestyle which will decrease the chance of getting the breast cancer. In future, these predictions can be made much easier when we have enormous amount of data. In this case, machine learning with strong pattern recognition algorithms can be applied to predict the mutations in the genomic profile.
Conclusion
Personalized medicine is a medical technique that categorizes patients into different groups based on genomic profile and medical decisions. Developments in personalized medicine will help to create integrated treatment tactics specific to individual with respect to their genome and offers enhanced or improved diagnoses. The advent of next generation sequencing and machine learning may enable rapid detection of the disease based on alterations in individual's genome which will help the clinician to assign personalized therapy to the patient.
Acknowledgement
This work was supported by Innov4Sight Health and Biomedical Systems Private Limited, Jayanagar, Bangalore, INDIA. We thank Gopinath Varadharajan, Balachander Agoramurthy, Vijaygopal R and Geetha Sanjay for their support in the research.
References
- AU Mannan, J Singh, R Lakshmikeshava, N Thota, S Singh, et al. (2016) Detection of high frequency of mutations in a breast and/or ovarian cancer cohort: implications of embracing a multi-gene panel in molecular diagnosis in India, J Hum Genet 61(6): 515-522.
- X Dai, T Li, Z Bai, Y Yang, X Liu, et al. (2015) Breast cancer intrinsic subtype classification, clinical use and future trends. Am J Cancer Res 5(10): 2929-2943.
- CB Matsen, LA Neumayer (2013) Breast Cancer: a review for the general surgeon. JAMA Surg 148(10): 971-979.
- ML Gatza, JE Lucas, WT Barry, JW Kim, Q Wang, et al. (2010) A pathway- based classification of human breast cancer. Proc Natl Acad Sci USA 107(15): 6994-6999.
- FM Blows, KE Driver, MK Schmidt, A Broeks, FEV Leeuwen, et al. (2010) Subtyping of Breast Cancer by Immunohistochemistry to Investigate a Relationship between Subtype and Short and Long Term Survival: A Collaborative Analysis of Data for 10,159 Cases from 12 Studies. PLoS Med 7(5): e1000279.
- DI Rodenhiser, JD Andrews, TA Vandenberg, AF Chambers (2011) Gene signatures of breast cancer progression and metastasis. Breast Cancer Res 13(1): 201.
- J Ferlay, C Hery, P Autier, R Sankaranarayanan (2009) Global Burden of Breast Cancer. Breast Cancer Epidemiology, p. 1-19.
- Breastcancer.org - Breast Cancer Information and Awareness. (2017) Fish Test (Fluorescence In situ Hybridization). USA.
- X Yang, ME Lippman (1999) BRCA1 and BRCA2 in breast cancer. Breast cancer research and treatment 54(1): 1-10.
- EA Slodkowska, JS Ross (2009) MammaPrint™ 70-gene signature: another milestone in personalized medical care for breast cancer patients. Expert review of molecular diagnostics. Expert Rev Mol Diagn 9(5): 417-422.
- JJ Carlson, JA Roth (2013) The impact of the Oncotype Dx breast cancer assay in clinical practice: a systematic review and meta-analysis. Breast cancer research and treatment. Breast Cancer Res Treat 141(1): 13-22.
- CB Mauer, SM Pirzadeh-Miller, LD Robinson, DM Euhus (2013) The integration of next-generation sequencing panels in the clinical cancer genetics practice: an institutional experience. Genet Med 16(5): 407






























