Promoter Hypermethylation of FHIT and P14 Genesin OSCC Patients among North Indian Population
Meenakshi Jha1, Sandesh Kumar Patel2, Abhimanyu Kumar Jha2 and Anju Shrivastava1*
1Department of Zoology, Delhi University, India
2Department of Biotechnology, IMS Engineering College, Ghaziabad, India
Submission: May 09, 2017; Published: June 08, 2017
*Corresponding author: Anju Shrivastava, Department of Zoology, Delhi University, Delhi-110021, India, Email: Anjus6768@gmail.com
How to cite this article: Meenakshi J, Sandesh K P, Abhimanyu K J, Anju S. Promoter Hypermethylation of FHIT and P14 Genes in OSCC Patients among North Indian Population.Canc Therapy & Oncol Int J. 2017; 5(2): 555660. DOI: 10.19080/CTOIJ.2017.05.555660
Abstract
Cancer is the leading cause for death worldwide causing millions of death. Oral squamous cell carcinoma (OSCC) is one the leading cancers causing mortality across the world. Major cause for oral cancer is the smoking, chewing of tobacco and alcohol consumption. It has been found that epigenetic changes are also responsible for the oral cancer. These are caused due to environmental and dietary factors. FHIT is a tumor suppressor gene which gets hypermethylated due to several factors. It was observed that the FHIT hypermethylation was found to be significant in the north Indian population (p-value= 0.0007). FHIT hypermethylation was observed in 55% (22 out of 40) of the samples.It was also found to be significant with respect to all the three risk factors. This statistical calculation was done through Epiinfo 7.2 software. However p14 hypermethylation was not found to be significant in OSCC patients among north Indian population. This is the first report on the promoter hypermethylation of FHIT gene in OSCC patients among north Indian population.
Keywords: OSCC; Epigenetic changes; FHIT gene; p14 gene; Promoter Hypermethylation
Abbreviations: OSCC: Oral Squamous Cell Carcinoma; DNA: Deoxyribonucleic acid; DNMTs: DNA methyltransferases; EDNRB: Endothelin receptor type B; OPLs: oral premalignant lesions; FHIT: Fragile Histidine Triad (gene)
Introduction
Cancer stems from the genetic corruption of a single cell. It is now recognized that cancer, in its simplest form, is a genetic disease, or more precisely, a disease of abnormal gene expression [1]. Cancer is the second most common cause of morbidity and mortality in the world today after cardiovascular problems. More than six million people die due to cancer every year. According to GLOBOCAN 2012, an estimated 14.1 million new cancer cases and 8.2 million cancer-related deaths occurred in 2012, compared with 12.7 million and 7.6 million, respectively, in 2008. Prevalence estimates for 2012 shows that there were 32.6 million people (over the age of 15 years) alive who had a cancer diagnosed in the previous five years.
Oral Cancer
a. Worldwide and Indian Perspective: Oral cancer or oral cavity cancer, a subtype of head and neck cancer, is any cancerous tissue growth located in the oral cavity [2]. There are several types of oral cancers out of which around 90% are squamous cell carcinomas. Most oral cancers are superficial and easily detected, but deeply located tumors may not be noted until they have grown large and reached an advanced stage. The second most common location is the floor of the mouth [3]. Due to its aggressiveness, it invades surrounding organs and causes regional or distant metastases [4]. The overall survival rate for oral cancer is considerably lower than that of other cancers due to metastasis and recurrence [5].
Many other different types of carcinomas of oral cavity can finally become malignant and result in a squamous cell carcinoma [6]. Oral cancer is a heterogeneous group of cancers arising from different parts of the oral cavity, with different predisposing factors, prevalence, and treatment outcomes. It is the sixth most common cancer reported globally with an annual incidence of over 300,000 cases, of which 62% arise in developing countries. In India it ranks in the top three after cervical and breast cancer, which accounts for over thirty per cent of all cancers, reported in the country and is quickly becoming a global health priority.
Risk factors
The highest incidence and prevalence of OSCC (Oral Squamous Cell Carcinoma) is found in the Indian subcontinent where the risk of developing OSCC is increased by the very prevalent habits of chewing tobacco, betel quid and areca-nut. Tobacco use, smoking and alcohol are known risk factors for cancers of the oral cavity. Additionally, these risk factors have showed a synergist effect when they have been combined [7].
The probability of developing OSCC increases with the period of exposure to risk factors, and increasing age adds a further dimension of age-related mutagenic and epigenetic changes. In India, the morbidity and mortality in males are 6.6 and 3.1 while in females 2.9 and 1.4. Petti revised investigations about prevalence and/or incidence of OSCC and concluded that % of cases are attributable to smoking, between 7 to 19% to alcohol consumption and 10 to 15% to micronutrient deficiency [8]. A case-control study was conducted in India, with patients diagnosed with oral cancer (n=388) and an equal number of age and sex- matched controls to assess the effect of lifestyle factors (tobacco chewing, smoking, alcohol drinking, diet and dental care) on the risk of oral cancer. They emphasized that these risk factors are modifiable and it is important to increase awareness among the general public and policy makers as a first step in the prevention and control of OSCC [9].
The oral habits of tobacco and betel quid chewing, bidi (tobacco flakes wrapped in a tendu leaf) smoking, Cannabis and alcohol consumption have been documented as risk factors for OSCC in Indian population and the most prevalent tumor sites are mouth and oropharynx [10]. All the above forms of tobacco are known to contain hydrocarbons and several potent nitrosamines which are carcinogenic and act via initiation and promotion of oral cancer [11].
Although recent research has given deeper insight into the etiology of the disease, its occurrence as a potentially fatal disease continues unabated. It is therefore essential to identify and develop newer risk markers for diagnosis and therapy of OSCC [12]. The development of oral squamous cell carcinoma (OSCC) is a multistep process requiring the accumulation of multiple genetic alterations, influenced by a patient's genetic predisposition as well as by environmental influences [13]. Genetic alterations that lead to loss or changes in tumour- suppressor genes are known to contribute to oral carcinogenesis [12]. The knowledge of molecular biology of distinctive tumors and research to find predictive biomarkers for targeted therapy plays a pivotal role in determining the fate of OSCC patients. Besides genetic changes, epigenetic alterations may lead to changes in gene expression as well; these include modifications of the genome without changes in the underlying DNA sequence.
Epigenetics
The word "epigenetics" was coined by the developmental biologist C.H. Waddington in 1942. Robin Holliday defined epigenetics as "the Study of the mechanism of temporal and spatial control of the gene activity during the development of complex organism". Epigenetics refers to the change in gene expression without the change in the sequence of the gene [14]. Alterations in epigenetic modifications regulating essential cellular processes required for maintaining cellular identity have been found to be associated with cancer. Epigenetic changes include a wide array of chromatin remodeling complex co regulators, effectors, and transcription factors, synthesis of noncoding microRNAs (miRNAs), DNA methylation, as well as covalent modification of histones such as acetylation, deacetylation, phosphorylation, ubiquitination, and sumoylation are considered as important determinants for regulation of gene expression.
a. DNA Methylation: It is an important regulator of gene transcription, and its role in carcinogenesis has been a topic of considerable interest in the last few years. Aberrant promoter hypermethylation of cancer associated genes occur frequently during carcinogenesis and may serve as a cancer biomarker [15,16].
b. Histone modifications: Within the nucleus, DNA is wrapped around a histone octamer formed by four histone partners - an H3H4 tetramer and two H2AH2B dimers. Post-translational modifications at the N-terminal of histones including acetylation, methylation, ubiquitination, phosphorylation, biotinylation, sumoylation, ADP ribosylation, and proline isomerization contribute to genomic stability, DNA damage response, and cell cycle checkpoints integrity affecting gene transcription and DNA repair [17,18].
c. miRNA: MicroRNAs (miRNAs) are small noncoding RNAs of 20-22 nucleotides that function in RNA silencing and post transcriptional regulation of gene expression. MicroRNAs are involved in the regulation of biological processes such as cell cycle control, apoptosis, and several developmental and physiological processes including differentiation and development and are altered in cancer development [19]. The involvement of miRNA in suppressing gene expression can arise through numerous mechanisms, including genomic abnormalities, transcriptional regulation, and processing of miRNAs [20].
d. Unlike Genetic alterations, which are almost impossible to reverse, epigenetic aberrations are potentially reversible, allowing the malignant cell population to revert to a more normal state [21].
e. DNA Methylation- the black box of Epigenetics: DNA is methylated by DNA methyltransferases (DNMTs) at the 5-position (C5) of the cytosine ring, almost exclusively in the context of CpG dinucleotides, which are poorly represented in the genome overall due to spontaneous deamination of 5-methyl-cytosine into thymine. CpG islands are the sites present in the promoter of most of the tumor suppressor genes, and hypermethylation at CpG island leads to silencing of the expression of these genes. It is mediated by different DNA methyltransferase (DNMT) enzymes among which DNMT1, an ubiquitous enzyme, is considered to be the major methyltransferase that is involved in the maintenance of DNA methylation after replication while the other two DNA- methylating enzymes (DNMT3A and DNMT3B) are found to be involved in de novo methylation that set up DNA methylation patterns early in development [22].
f. Growth regulatory gene promoter region-associated CpG islands are found to be hypermethylated in cancer cells which leads to the transcriptional silencing of tumor suppressor genes [23], thus contributing to cancer progression, while derepression ofproto-oncogene transcription by demethylation leads to increased mutation rates and chromosome instability, which is an early characteristic of tumor cells [24].
g. DNA methylation in OSCC: DNA methylation is found to be closely related to OSCC tumor genesis [25]. From the analysis of DNA extracted from OSCC tissues and oral premalignant lesions (OPLs), it has been found that they exhibit more frequent and higher levels of DNA methylation, in comparison with healthy or corresponding normal tissue from neoplastic tissues [26]. The use of tobacco has been found to be associated with nonspecific global hypomethylation [27]. In contrast to smokers, patients who drink heavily have been found with an increased risk for CpG hypermethylation of multiple OSCC-related genes. Chronic inflammation of the oral mucosa is another risk factor that can potentially modify the methylation status of various genes in OSCC tumors [28]. The occurrence of multiple CpG methylation sites in a panel of tumor-related genes in OSCC was found to be highly associated with cancer stage and may also correlate with lymph node metastasis [29]. To date, CpG methylation of different genes that are involved in cell cycle control, apoptosis, signal transduction, and DNA damage repair has been reported [30].
h. Estimates of the global prevalence of oral potentially malignant disorders range from 1 to 5% [29]. Both genetic and epigenetic changes are known to contribute to tumor genesis in humans. While genetic alterations refer to irreversible changes in DNA sequence leading to oncogene activation or tumor suppressor gene inactivation [30], epigenetic changes denote reversible and heritable modifications in gene expression without any alterations in the DNA sequence [27].
i. A plethora of genes showing hypermethylation have been discovered in OSCC. Recent research suggest that p16 hypermethylation was associated with early incidence of OSCC, increased lymph node invasion in young patients, and poor prognosis in older patients. Further, p16 hypermethylation may also be implicated in age-related tumor invasion in carcinogenesis [31]. Aberrant methylation of EDNRB, KIF1A, DCC and p161NK4a genes is a common event in OSCC, suggesting that epigenetic alterations of these genes warrant validation in larger studies for their potential use as biomarkers [32]. Some of the main genes that frequently showed promoter methylation in OSCC are those that participate in diverse processes such as regulation of the cell cycle, DNA repair, proliferation, and apoptosis.
FHIT Gene
FHIT gene is a member of the fragile histidine triad gene family It encodes a diadenosine 5', 5”'-P1, P3-triphosphate hydrolase involved in purine metabolism. It was shown that the FRA3B/FHIT locus frequently exhibits deletions in preneoplasias and cancers, FHIT protein expression is lost or reduced in the majority of human cancers, and the orthologous mouse FHIT locus is fragile and exquisitely sensitive to carcinogen damage [33].
Evidence of aberrant transcripts was found in 53% of OSCCs and in two of seven premalignant lesions, one of which was from a patient developed OSCC during follow-up. In a study, investigators [34] compared the frequency of detection of FHIT aberrant expression using the RT- PCR assay for mRNA aberration and analysis of level of FHIT protein expression by immunohistochemical analysis in tissue sections using several polyclonal anti-FHIT sera. The findings were that 71% of the oral squamous cell carcinomas (OSCCs) showed reduced or absent FHIT protein and half of the cases with reduced FHIT showed aberrant RT -PCR products. In a study of OSCCs associated with betel and/or tobacco use [35], 28% of samples exhibited FHIT promoter methylation, and 36% of cancers and 50% of premalignant lesions showed aberrant RT-PCR products, suggesting that FHIT aberration could be an early event in oral carcinogenesis. But these changes vary from population to population due to difference in dietary and environmental factors.
p14 Gene
p14ARF (also called ARF tumor suppressor, ARF, p14ARF) is an alternate reading frame protein product of the CDKN2A locus (i.e. 1NK4a/ARF locus). ARF is induced in response to elevated mitogenic stimulation, such as aberrant growth signaling from MYC and Ras (protein). It accumulates mainly in the nucleolus where it forms stable complexes with NPM or Mdm2. These interactions allow p14ARF to act as a tumor suppressor by inhibiting ribosome biogenesis or initiating p53-dependent cell cycle arrest and apoptosis, respectively. p14ARF is an atypical protein, in terms of its transcription, its amino acid composition, and its degradation: it is transcribed in an alternate reading frame of a different protein, it is highly basic, and it is polyubiquinated at the N-terminus [36].
Both p16INK4a and p14ARF are involved in cell cycle regulation. p14ARF inhibits mdm2, thus promoting p53, which promotes p21 activation, which then binds and inactivates certain cyclin-CDK complexes, which would otherwise promote transcription of genes that would carry the cell through the G1/S checkpoint of the cell cycle. Loss of p14ARF by a homozygous mutation in the CDKN2A (INK4A) gene will lead to elevated levels in mdm2 and, therefore, loss of p53 function and cell cycle control. If the oral cancer is detectable at early stages of development, it has an 80 to 90% survival rate. Unfortunately, most of these cancers are hard to discover in the early stages because of a lack of public awareness and screening method, which would generally result in a poor prognosis and a low survival rate.. Currently, the most definitive method for oral cancer diagnosis and screening is a scalpel biopsy. Extraction of biopsy is a very painful process for the patients.
Epigenetic changes, like promoter hypermethylation of tumor suppressor genes can be detected in blood samples also and these changes can be used as diagnostic biomarker in OSCC. These epigenetic changes are dependent on environmental and dietary factors, which vary from one population to another. Hence, the present study was designed and focused on checking the promoter hypermethylation status of FHIT and p14 genes in OSCC patients among north Indian population and statistically correlating the promoter hypermethylation of these genes with risk of OSCC.
Materials and Methods
Sample Collection
Blood samples (40) were collected with informed consent of patients diagnosed with OSCC after obtaining the necessary ethical clearance from Dharamshila Cancer Hospital & Research Centre, New Delhi. The blood samples (20) from healthy individuals (as controls) were also obtained. The samples were further used for DNA extraction.
DNA extraction
Cells obtained from tissue biopsies and blood samples were lysed in digestion buffer (10 mM Tris-HCl, pH 8.0, 10 mM EDTA, 150 mM NaCl and 2% SDS) containing proteinase K (0.2 mg/ml). DNA was then purified using the standard phenol-chloroform extraction and ethanol precipitation.
Methylation-specific PCR (MS-PCR)
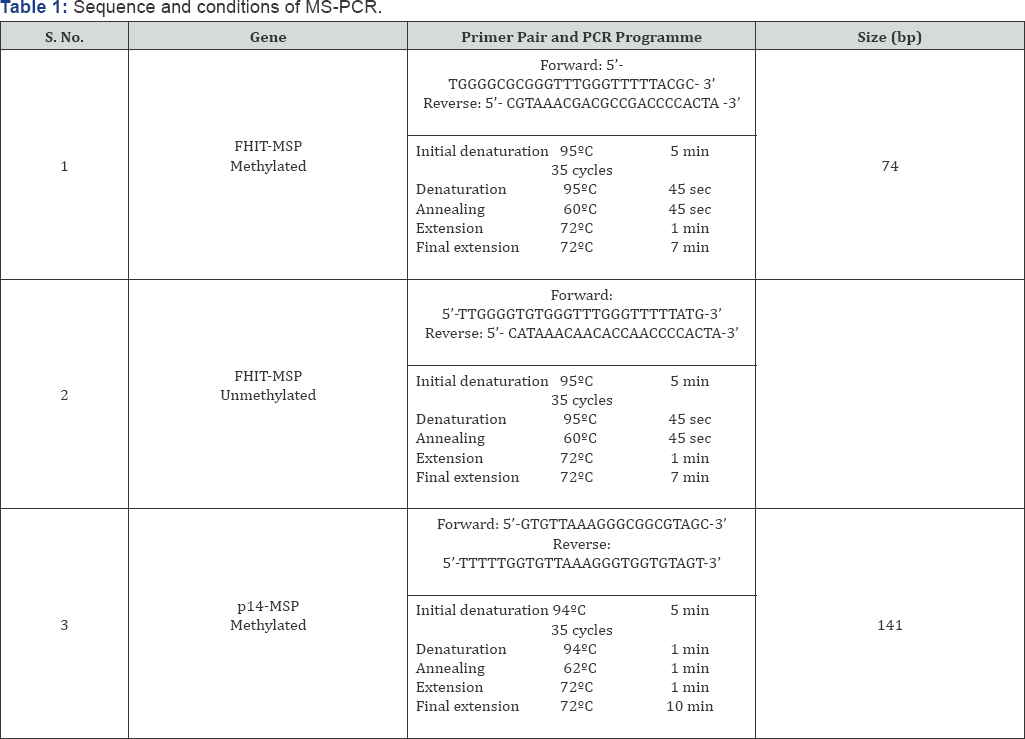
DNA isolated from the biopsy and blood samples was modified with sodium bisulphite and MS-PCR was carried out using specific primers for methylation and unmethylation for the p14ARF and FHIT genes (Table 1). The amplified products were run on a 2% agarose gel.
Statistical analysis
The association between hypermethylation of the genes and risk of OSCC was estimated by computing odds ratios (ORs) and 95% confidence intervals (CI) using the Chi-square test, Fisher's exact test and multivariate logistic regression analysis, which included several potential confounding variables. The reported OR may be interpreted as age-adjusted estimates of the relative risk of developing OSCC with the methylation of studied genes. Statistical analysis was performed using SPSS version 11.5 and Epi Info version 7.0. Px003C;0.05 was considered to indicate a statistically significant difference.
Statistical analysis
The association between hypermethylation in both genes and correlation with the risk of OSCC was estimated by computing ORs (Odds Ratios) and 95% CI (confidence interval) using Chi- square, Fisher’s exact test and multivariate logistic regression analysis which included several potential confounding variables. p value<0.05 was considered as significant. The Relative risk was calculated using Epiinfo Software version 7.0.
Results
The cases and the controls were well matched with respect to age, sex and place of living. The mean age ±SD of all cases and controls was 45.33±8.55 and 45.99±11.85 years respectively. According to place of living, patients and control were divided in two groups (a) urban (b) rural. 47.5% of patients were from urban and 52.5% were from rural background. The distribution for controls was quite similar to the cases (45% urban and 55% rural). Three risk factors were chosen for the present study: tobacco chewing, smoking and alcohol consumption.
DNA Extraction from blood sample
The DNA isolated from the blood was run on the 1% agarose gel and visualized under gel documentation unit. This was carried out to check the quality of DNA (Figure 1). The isolated DNA was also quantified using nanodrop (ND-1000).
Sodium Bisulphite modification
The Sodium bisulphite modification of the genomic DNA was performed using agarose bead method. It was carried out to distinguish methylated cytosine from unmethylated cytosine.
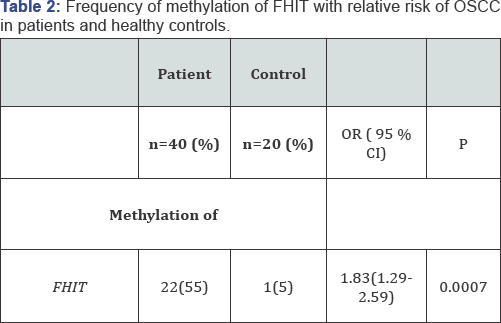
FHIT Hypermethylation
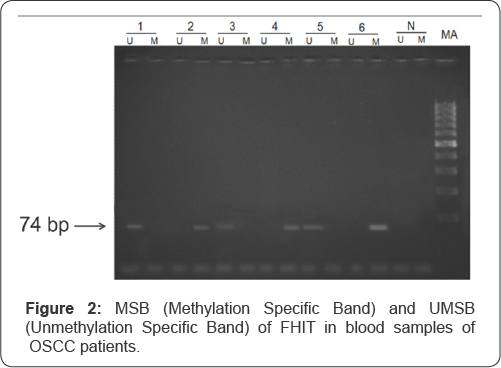
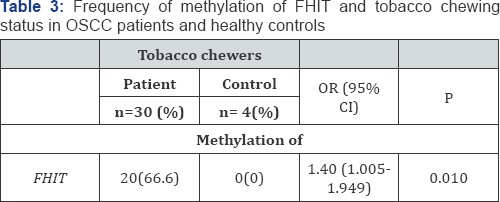
MSP was performed to check the promoter hypermethylation of FHIT gene in OSCC patients among north Indian population (Figure 2). Photomicrograph representing 2.5% agarose gel electrophoresis of PCR amplified FHIT gene that was carried out in 1X TAE buffer at a voltage supply of 80 V. L - 100bp ladder, U - unmethylation specific band, M - methylation specific band, N-Negative Control. Hypermethylation of FHIT was observed in 55% of OSCC patients. The methylation of FHIT showed significant risk of developing OSCC in patients (p-value<0.05) (Table 2). FHIT was observed to be methylated in 66.6% oftobacco chewers among patients and 0% among controls. Statistically significant relation in association with FHIT methylation was observed in the tobacco chewers (p-value<0.05) (Table 3).
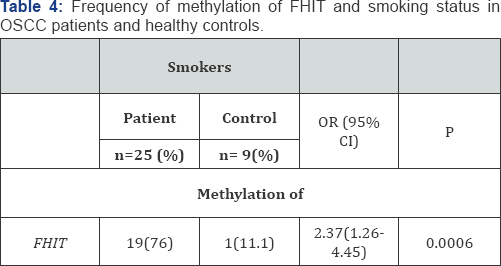
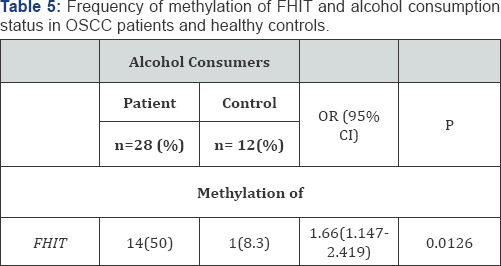
FHIT was observed to be methylated in 76% of smokers among patients and 11.1% among controls. A statistically significant increase in the risk of OSCC was also observed in case of smokers (p-value<0.05) (Table 4). FHIT was observed to be methylated in 50% of alcohol consumers among patients and 8.3% among controls. A statistically significant increase in the risk of OSCC was also observed in case of alcohol consumers (Table 5) (p-value<0.05).

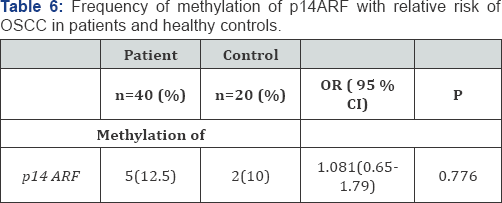
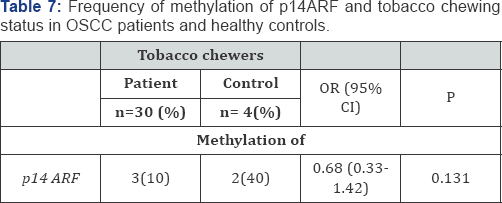
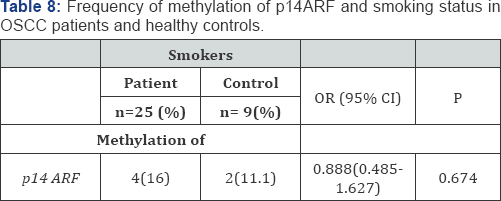
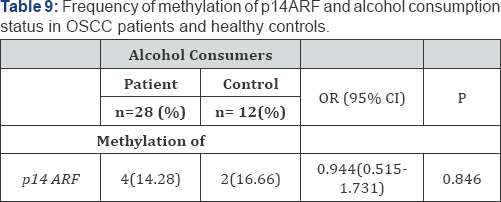
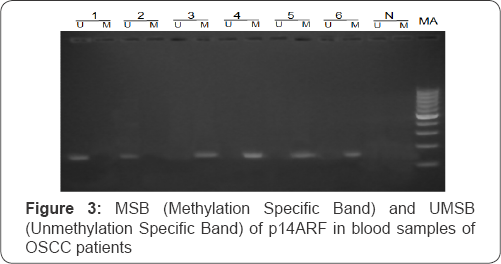
Hypermethylation of p14ARF was observed in 12.5% of OSCC patients (Figure 3). Any significant difference in methylation of p14ARF (p-value>0.05) between patients and controls was not observed. The methylation of p14ARF showed no risk of developing OSCC in patients (Table 6). P14ARF was observed to be methylated in 10% of tobacco chewers among patients and 40% among controls. No significant increase in the risk of OSCC was observed in case of tobacco chewers (Table 7). p14ARF was observed to be methylated in 16% of smokers among patients and 11.1% among controls. No significant increase in the risk of OSCC was observed in case of smokers (Table 8). p14ARF was observed to be methylated in 14.28% of alcohol consumers among patients and 8.3% among controls. No significant increase in the risk of OSCC was also observed in case of alcohol consumers (Table 9).
Discussion
Cancer is the second most common disease worldwide. According to Globocan 2012, 14.1 million new cancer cases, 8.2 million cancer deaths occurred, and 32.6 million people were living with cancer (within 5 years of diagnosis) worldwide. The majority of the cancer cases occurred in the less developed regions with an estimate of 57% (8 million) new cancer cases and 65% (5.3 million) cancer deaths. Cancer starts and progresses by synchronized genetic and epigenetic alterations, which cause changes in expression of multiple genes or leading to either activation of oncogenes or silencing of tumor suppressor genes.
Oral squamous cell carcinoma (OSCC) is one of the most common types of oral neoplasm, accounting for over 90 % of all mouth malignancies and 38 % of head and neck tumors. OSCC constitutes the eighth most common malignant tumors worldwide, being diagnosed with more than 500,000 new cancer cases and more than 100,000 cancer deaths every year. In fact, despite of recent advances in the field of oral cancer detection, prevention, and multimodality treatments, the overall 5-year survival for OSCC continues to be modest at best. The most important factor affecting OSCC survival after treatment is the stage of tumor at diagnosis. Although early-stage tumors (without lymph node involvement) have an ∼80 % 5-year survival rate, the 5-year survival rates for patients with regional lymph node spread or metastases are only about 56 and 34 %, respectively. Therefore, to improve long-term outcomes, an early detection in conjunction with primary and secondary prevention strategies is critical Oral carcinogenesis is a multistep process modulated by endogenous and environmental factors among which regular intake of chronic tobacco and alcohol as well as smoking constitutes three major risk factors, while chronic inflammation, human papillomavirus (HPV) persistent infection, betel quid chewing, and genetic predisposition are known to be supplementary factors that contribute toward its pathogenesis [36,37]. These predisposing factors may lead to a wide range of genetic and epigenetic alterations consisting of aberrant DNA methylation, histone modifications, and altered expression of miRNAs that promote genomic instability and tumor development and progression.
The genetic alterations are caused by irreversible changes in DNA sequence including gene deletions, amplifications, and mutations which lead to oncogene activation or tumor suppressor gene inactivation [38-40]. The epigenetic changes refer to any mitotically heritable alteration in gene expression without alterations of the DNA sequence. These changes occur more frequently than gene mutation. Epigenetic changes include DNA methylation, histone modifications, and RNA-mediated silencing. Disruption of any of these three distinct mutually reinforcing epigenetic mechanisms leads to dysregulation of gene expression, resulting in cancer development and other epigenetic diseases [41]. Epigenetics is defined as heritable changes in gene expression caused by chemical modification of genes that do not involve changes in primary DNA sequence. These epigenetic modifications are considered as potential initiating events that occur during early carcinogenesis. Epigenetic mechanisms including chromatin remodeling in an appropriate manner via recruitment of a wide array of chromatin remodeling complex co regulators, effectors, and transcription factors, synthesis of noncoding microRNAs (miRNAs), DNA methylation, as well as covalent modification of histones such as acetylation, deacetylation, phosphorylation, ubiquitination, and sumoylation are considered as important determinants for regulation of gene expression[21].
Epigenetic changes play a big role in cancer as it is the phenomena in which the functionality of the gene is lost without any genetic alteration or mutation [14]. DNA methylation is an important regulator of gene transcription, and its role in carcinogenesis has been a topic of considerable interest in the last few years.DNA methylation is a biochemical process involving the addition of methyl group to the cytosine or adenine DNA nucleotides. A methyl group is covalently attached to cytosine C5'. When DNA is treated with bisulphite, unmethylated cytosines are converted to uracil, but methylated cytosines are protected. In this phenomena the promoter region of the gene is hypermethylated which caused global hypomethylation to the gene. When these methyl groups are detached from the cytosine nucleotide then the expression of the sequence is changed. This is called reversal of methylation. This cause the expression to expressed which is hindered due to methylation. Epigenetic changes varies from population to population place to place as it mainly depend upon the environmental and dietary factors which varies greatly.
Bis (5'-adenosyl)-triphosphatase also known as fragile histidine triad protein (FHIT) is an enzyme that in humans is encoded by the FHIT gene. This gene, a member of the histidine triad gene family, encodes a diadenosine 5',5”’-P1,P3- triphosphate hydrolase involved in purine metabolism FHIT gene has been shown to be hypermethylated in oesophageal, lung, breast, prostate, bladder, cervical, and oral cancers (Dhillon et al., ). p14 gene is one of the splice variants of the CDKN2A gene, located on chromosome 9p21. The p14 protein activates the tumor suppressor gene p53 by inhibiting MDM2, an ubiquitin ligase that marks p53 for degradation which in turn leads to cell cycle arrest or apoptosis in cells [42]. Till now, no report exists on the hypermethylation of promoter region of FHIT and p14 genes in OSCC patients among north Indian population. Epigenetic changes vary from one population to another because of the different dietary and environmental factors. To the best of our knowledge, no published work is there from India on the promoter hypermethylation of the genes selected for the present study. Hence, the above mentioned genes were selected for the present study.
Regular intake of chronic tobacco and alcohol as well as smoking constitutes three major risk factors in the progression of OSCC. Hence, the present study also focused on correlating the methylation status of these genes with these risk factors in OSCC patients among north Indian population. In the present study, the DNA was isolated from the collected blood samples of cancer patient (OSCC) and control samples (patients not suffering from cancer). Further isolation of DNA was followed by sodium bisulfite modification by agarose bead method and E Z DNA Gold Methylation Kit (Zymo Research, US), after which MSP (methylation specific PCR) was carried out using methylation specific primers. Hypermethylation of FHIT was observed in 55% of OSCC patients. Statistically significant difference in methylation of FHIT (p-value<0.05) between patients and controls was observed. The methylation of FHIT showed significant risk of developing OSCC in patients. Statistically significant relation in association with FHIT methylation was observed in the tobacco chewers, smokers and alcohol consumers (p-value<0.05).A statistically significant increase in the risk of OSCC was observed in case of all of these categories.
This proves that FHIT hypermethylation is a significant step in the OSCC patients among north Indian Population. If these results are obtained on a large scale study the promoter hypermethylation of FHIT gene may be used as a diagnostic biomarker in OSCC. Hypermethylation of p14ARF was observed in 12.5% of OSCC patients. Any significant difference in methylation of p14ARF (p-value>0.05) between patients and controls was not observed. The methylation of p14ARF showed no risk of developing OSCC in patients. Statistically significant relation in association with p14ARF methylation was not observed in the tobacco chewers, smokers and alcohol consumers (p-value>0.05). No significant increase in the risk of OSCC was observed in all of these cases.
Hence, based upon the present study p14 could not be used as a diagnostic biomarker in OSCC patients among north Indian population. It was concluded that several factors lead to promoter hypermethylation of FHIT gene among north Indian population. But this study needs to be carried out on a large scale as the sample size in present study was small to draw any complete statistical conclusion. But the present study clearly shows the trend of FHIT hypermethylation among in OSCC patients among north Indian population.
More prospective studies are needed to affirm the clinical applicability of FHIT hypermethylation in larger groups of patients. Early assessment of FHIT hypermethylation might enable the identification of subgroups of patients with poor prognosis, who might require a different therapeutic approach. Therefore, we recommend future research to explore the position of this potential biomarker in the clinical management of OSCC and to evaluate whether it can contribute to personalized treatment strategies.
Conclusion and Future Perspectives
Promoter methylation of tumor suppressor genes is an important factor in carcinogenesis of OSCC. Analysis of DNA methylation is a useful approach to evaluation of the biological characteristics of oral cancers and may be a useful diagnostic biomarker. Hence, the present study was designed to study the methylation status of the genes like FHIT and p14 genes and to correlate the methylation status of these genes with the risk of OSCC statistically. The risk of OSCC was also calculated in correlation with the three risk factors- tobacco chewing, smoking and alcohol consumption and was found to be significant. It was concluded that several factors lead to promoter hypermethylation of FHIT gene among north Indian population. This proves that FHIT hypermethylation is a significant step in the OSCC patients among north Indian population and it can possibly be used as a diagnostic biomarker in OSCC patients. Based upon the present study, p14 could not be used as a diagnostic biomarker in OSCC patients among north Indian population. This study needs to be carried out on a large scale as the sample size in present study was small to draw any complete statistical conclusion. This is the first study on methylation status of these genes in OSCC among north Indian population. The research on methylation of these genes which leads to their inactivation in the OSCC will contribute to explore anti-apoptotic pathway of tumor cells and provide guidance for future treatment.
References
- Coleman WB, Tsongalis G J (2002) The role of genomic instability in the development of human cancer. The Molecular Basis of Human Cancer Humana Press Totowa pp: 115-142.
- Werning JW (2007) Oral cancer: diagnosis, management, and rehabilitation 1.
- Tanaka T, Ishigamori R (2011) Understanding carcinogenesis for fighting oral cancer. J Oncol pp: 6037-6040.
- Mishra R (2012) Biomarkers of oral premalignant epithelial lesions for clinical application. Oncol 48(7): 78-584.
- Kademani D (2007) Oral cancer. Mayo Clin Proc 82: 878-887.
- Sankaranarayanan R, Masuyer E, Swaminathan R, Ferlay J, Whelan S (1998) Head and neck cancer: a global perspective on Epidemiology and prognosis. Anticancer Res 18(6B): 4779-4786.
- Bundgaard T, Bentzen SM, Wildt J (1994) The prognostic effect of tobacco and alcohol consumption in intra-oral squamous cell carcinoma. Eur J Cancer B Oral Oncol 30B: 323-328.
- Petti S (2009) Lifestyle risk factors for oral cancer. Oral Oncol 45: 340350.
- Subapriya R, Thangavelu A, Mathavan B, Ramachandran CR, Nagini S, et al. (2007) Assessment of risk factors for oral squamous cell carcinoma in Chidambaram, Southern India: a case-control study. Eur J Cancer 7: 251-256.
- Nair U, Bartsch H, Nair J (2004) Alert for an epidemic of oral cancer due to use of the betel quid substitutes gutkha and pan masala: a review of agents and causative mechanisms. Mutagenesis 19 (4): 251-262.
- Datta K, Saha RK, Chakrabarti RN (1997) A simple risk estimates study for oral cavity cancer, practical approach in Indian context. Journal of the Indian Medical Association 95(3): 70-71.
- Viswanathan M, Tsuchida N, Shanmugam G (2003) Promoter hypermethylation profile of tumor associated genes p16,p15, Hmlh1,MGMT and E-Cadherin in oral squamous cell carcinoma. International Journal of Cancer 105: 41-46.
- Choi S, Myers JN (2008) Molecular Pathogenesis of Oral Squamous Cell Carcinoma implications for Therapy. Journal of Dental Research 87 (1): 14-32.
- Jha AK, Nikbakht M, N Kaur J (2011) Cervical cancer: Promoter hypermethylation and its reversal. Journal of Pharmacy Research 4(10): 3378-382.
- Jha AK, Nikbakht M, Jain V, N Capalash, J Kaur (2012) p16INK4a and p15INK4b gene promoter methylation in cervical cancer patients. Oncol Lett 3: 1331-1335.
- Jha AK, Nikbakht M, Jain V, Capalash N, Kaur J (2012) Promoter hypermethylation of p73 and p53 genes in cervical cancer patients among North Indian population. Mol Biol Rep 39(9): 9145-9157.
- Jha AK, Nikbakht M, Capalash N, Kaur J (2001) Demethylation of RARß2 gene promoter by Withania somnifera in HeLa cell line. Euro J Med Plants 4(5): 503-510.
- Fullgrabe J, Kavanag, E, Joseph B (2011) Histone oncomodifications. Oncogene 30: 3391-3403.
- Calin G A, Croce C M (2006) MicroRNA signatures in human cancers. Nat Rev Cancer 6(11): 857-866.
- Winter J, Jung S, Keller S, Gregory R I, et al. (2011) Many roads to matu- rity:microRNA biogenesis pathways and their regulation. Nat Cell Biol 11: 228-234.
- Yoo CB, Jones PA (2006) Epigenetic therapy of cancer: past, present and future. Nat Rev Drug Discov 5: 37-50
- Board RE, Blackhall FH, Hughes A, Andrew Hughes, Caroline Dive, et al. (2008) DNA methylation n circulating tumour DNA as a biomarker for cancer. Biomarker Insight 2: 307-319.
- Eden A, Gaudet F, Waghmare A, Jaenisch R (2003) Chromosomal instability and tumors promoted by DNA hypomethylation. Science 300: 300-455.
- Feinberg AP, Cui H, Ohlsson R (2002) DNA methylation and genomic imprinting Insights from cancer into epigenetic mechanisms. Seminars in Cancer Biology 12(5): 389-398.
- Piyathilake CJ, Bell WC, Jones J, Olga L Henao, William E Grizzle, et al. (2005) Pattern of nonspecific (or global) DNA methylation in oral carcinogenesis. Head & Neck 27(12): 1061-1067.
- Baba S, Yamada Y, Hatano Y, Shibata T, Hara A, et al. (2009) Global DNA hypomethylation suppresses squamous carcinogenesis in the tongue and esophagus. Cancer Science 100(7): 1186-1191.
- Gasche JA, Hoffmann J, Boland CR, Goel A (2011) Interleukin-6 promotes tumorigenesisby altering DNA methylation in oral cancer cells. International Journal of Cancer 129(5): 1053-1063.
- Supic G, Kozomara R, Jovic N, Zeljic K, Magic Z (2011) Prognostic significance of tumorrelated genes hypermethylation detected in cancer-free surgical margins of oral squamous cell carcinomas. Oral Oncology 47(8): 702-708.
- Johnson W, Jayasekara P, Hemantha K (2011) Squamous cell carcinoma and precursor lesions of the oral cavity: epidemiology and aetiology. Periodontology 57(1): 19-37.
- Saintigny P, Zhang L, Fan YH, El-Naggar AK, Papadimitrakopoulou VA, et al. (2011) Gene expression profiling predicts the development of oral cancer. Cancer Prevention Research 4: 218-229.
- Su PF, Huang WL, Wu HT, Liu TY, Kao SY, et al. (2010) p16(INK4A) promoter hypermethylation is associated with invasiveness and prognosis of oral squamous cell carcinoma in an age-dependent manner. Oral Oncology 46(10): 734-739.
- Kaur J, Demokan S, Tripathi SC, Macha, Shahnaz Begum, et al. (2010) Promoter hypermethylation in Indian primary oral squamous cell carcinoma. International Journal of Cancer 127(10): 2367-2373.
- Huebner K, Croce C M (2002) FRA3B and other common fragile sites, The weakest Links. Nat Rev Cancer 1: 214 -221.
- Heerden WFP, Swart Smith TL, Huebner K (2001) FHIT RNA and protein expression in oral squamous cell carcinomas. Anticancer Res 21: 2425-2428.
- Chang KW, Kao RJ, Lin SC (2002) Multiple molecular alterations of FHIT in betel-associated oral carcinoma. J Pathol 196: 300-306.
- Kuo ML, Den Besten Sherr CJ, Bertwistle D, Roussel MF, Sherr CJ (2004) "N-terminal polyubiquitination and degradation of the Arf tumor suppressor”. Genes Dev 18(15): 1862-1874.
- Gillison M L, D'Souza GS, Viscidi R, Begum S, Viscidi R, et al. (2008) Distinct risk factor profiles for human papillomavirus type 16-positive and human papillomavirus type 16- negative head and neck cancers. J Natl Cancer Inst 100: 407-420.
- Chaturvedi AK, Anderson WF, Gillison ML (2008) Incidence trends for human papillomavirus-related and unrelated oral squamous cell carcinomas in the United States. Journal of Clinical Oncology 26(4): 612619.
- Lingen MW, Franchini R, Tilakaatne W M, Peterson DE, Woo SB, et al. (2011) Genetics/epigenetics of oral premalignancy: current status and future research. Oral Diseases 17: 7-22.
- Kyrgidis A, Tzellos TG, Triaridis S (2010) Melanoma: stem cells, sun exposure and hallmarks for carcinogenesis, molecular concepts and future clinical implications. Journal of Carcinogenesis 9: 1477-3163.
- Egger G, Liang G, Jones PA (2004) Epigenetics in human disease and prospects for epigenetic therapy. Nature 429: 457-463.
- Leemans CR, Braakhuis BJ M, Brakenhoff RH (2011) The molecular biology of head and neck cancer. Nature Reviews Cancer 11(1): 9-22.






























