Registration of Nasopharynoscope Videos with Radiation Treatment Planning CT Scans
Julian Rosenman1*, Qingyu Zhao2, True Price2, Rui Wang2, Junpyo Hong2, Marc Niethammer2, Ron Alterovitz2, Jan-Michael Frahm2, Bhishamjit Chera1 and Stephen Pizer1,2
*1Department of Radiation Oncology, The University of North Carolina at Chapel Hill, USA
*1Department of Radiation Oncology, The University of North Carolina at Chapel Hill, USA
Submission: April 23, 2017; Published: May 05, 2017
*Correspondence Address: : Julian Rosenman, PhD MD, Department of Radiation Oncology, The University of North Carolina at Chapel Hill, 101 Manning Drive, CB 7512, Chapel Hill NC 27599, USA, Tel: [919) 966-7681; Email: rosenmju@med.unc.edu
How to cite this article: Julian R,Qingyu Z, True P, Rui W. Registration of Nasopharynoscope Videos with Radiation Treatment Planning CT Scans. Canc Therapy & Oncol Int J. 2017; 5(1): 555652. DOI: 10.19080/CTOIJ.2017.05.555652
Purpose: We present a method that allows one to register endoscopic videos with CT scans to provide additional data for radiation treatment planning of the oropharyngeal region. We hypothesize that this additional data will allow for more accurate radiation treatment plans. Methods and Materials: Our approach is to reconstruct a 3D textured surface from the 2D video data as seen on multiple video frames. That reconstruction, which we call an endoscopogram, can then be deformably registered to the CT scan. To reconstruct an endoscopogram from raw video, we first compute a sparse 3D point cloud by tracking corresponding feature points from one video frame to nearby ones using standard structure from-motion (SfM) techniques. During this procedure we also calculate the most likely camera trajectory. We then employ a second technology, shape-from-shading (SfS), to produce high-quality 3D surfaces from each video frame and correct these SfS images using the global accurate point cloud as a constraint. Finally we co-register several of these frame-based surfaces to create an endoscopogram that can be deformably registered with a CT scan. Results: We have successfully computed a number of spatially accurate endoscopograms and have registered them with their planning CT scans with an average accuracy of about 2-3 mms. Conclusion: These technologies will allow for accurate data transfer from endoscopy images directly onto a radiation planning CT scan that may improve treatment of head and neck cancer In addition, the endoscopogram allows one to see the entire videoed mucosal surface at a glance. Keywords: Endoscopy; Deformable registration; Radiation treatment planning; Registering endoscopic videos with CT scans 2Abstract
Introduction
Modern radiation therapy treatment planning relies on imaging to determine tumor location and spread. Although for several reasons CT scans are preferred as the base image for radiation treatment planning, information from MRi and PhT scans can be added to the planning CT via registration using only rigid registration because CT and MRi are acquired with the patient in a similar position. in the case of PhT/CT these two scans are done in rapid sequence during the same imaging session. In the past few years a new and important kind of medical imaging has become readily available although we do not usually think of it that way. These are videos that can be taken during endoscopy.
For example, otolaryngologists and radiation oncologists regularly perform nasopharyngoscopy because they feel that direct visualization of the tumor provides information on tumor location (especially mucosal spread) not available on CT or even PhT/CT. The radiation oncologist, in particular, uses that information to improve the accuracy of gross tumor volume (GTV) delineation on CT. However, endoscopic data can be utilized only indirectly, through the physician's memory or notes, as there has been no way to directly transfer the endoscopic data to CT, that is, no way to register CT and video data.
This paper describes a new process whereby endoscopic video frames can be reconstructed into a 3D textured surface that we call an endoscopogram. The endoscopogram then can either viewed directly, as it shows the entire video graphed mucosal surface at a glance, or registered with the planning CT. After registration selected data from the endoscopy, such as gross tumor extent, can be added directly to the planning CT. The possible clinical benefits of this technology will be discussed and an ongoing clinical trial described. To our knowledge this approach to improve radiation treatment planning has not been previous attempted.
Methods And Materials
There are two major technical problems that must be solved before one can register an endoscopic video with the planning CT. The first is the reconstruction of an accurate 3D model from the 2D video data (the endoscopogram) and the second is the registration of the reconstructed endoscopogram with the CT.
Reconstruction of the endoscopogram
The problem of reconstructing a spatially accurate endoscopogram from the endoscopic video proved to require three separate processes, two of them new. The first process is that of determining the three-dimensional spatial position of as many "feature points" within the endoscopic video frames as possible, along with the camera position and orientation for each video frame. The method presupposes that, to first approximation, in nearby frames the scene remains still and the camera moves in a regular fashion, although the camera position and orientation are not known. Methods for computing this point cloud (set of points for which the spatial position has been determined) under these conditions have been developed over many years [1-7], but they have typically not been used to reconstruct human anatomy. More commonly, such techniques are used to reconstruct "urban scenes" composed of buildings, roads and Registering endoscopic videos with CT scans 4 other objects that contain many straight lines and right angles as well as a fixed light source. Although the output of structure from motion (SfM) algorithms is typically a only a relatively sparse point cloud, under the assumption that the buildings and other structures are mostly comprised of flat or near planar faces an accurate, high resolution 3D model can be reconstructed. That is not the case for endoscopy, as human anatomy is noticeably lacking in straight lines and planar surfaces.
Moreover, since the anatomy is continuously deforming, we need to limit the structure from motion reconstruction to short frame sequences where the rigidity and fixed lighting assumptions hold approximately. As a result of these problems, a high-resolution reconstruction of an endoscopogram from a sparse SfM-generated point cloud alone proved not to be possible, but we could obtain an accurate sparse point cloud and a spatial position and orientation of the camera for every video frame. The second process is the utilization of another historically successful approach to reconstructing 3D models from 2D data, but one that does not use parallax methods to generate a point cloud. Known as "shape-from-shading," SfS is a technique that attempts to recover the 3D shape from a 2D image using the gradual variation of shading within the image as a guide. This is the reverse of what artists do when they convey depth in a picture by using an appropriate shading method. The idea of obtaining shape from shading, stems from work begun in 1952 when it was used to calculate the slopes and heights of mountains and craters on the moon from their shadows [8]. In the 1980s a series of papers formalized the method for images with a known lighting model, i.e., known light sources and surfaces of known reflectivity. Presently there are at least six major approaches to doing shape-from-shading from a single image [9].
In our experience, rendering the pharynx with SfS typically results in high quality 3D surfaces with locally accurate curvature, but with globally inaccurate depth scaling, and significant connectivity problems, particularly in areas of occlusion, where part of the anatomy cannot be viewed because something is in the way. We now had two reconstruction methods, SfM that would produce a globally accurate but sparse point cloud that lacked information on local curvature, and an SfS technique that produced that local curvature correctly but lacked global fidelity. We felt that the two approaches should be combined to take advantage of the best qualities of both.
Unfortunately a straightforward attempt to compute a SfS surface under the SfM constraints proved to be unsatisfactory. Fortunately, we were able to develop a new iterative method to correct the global errors found in SfS-generated images. The approach begins with the assumption that there is some lighting model, varying across the surface that when used with current SfS algorithms would produce a surface with both locally and globally correct curvatures and depths. We start with a simple lighting model and use the SfM point cloud as a constraint to compute an improved lighting model. In an iterative fashion, the new lighting model is then used to compute an improved depth map. We call this method SfMS (Structure from Motion and Shading) [10]. SfMS is done for each video frame selected to be part of the endoscopogram. Each frame is thus reconstructed into a 3D surface that will make up part of the entire endoscopogram. So although we often refer to an SfMS surface as coming from a single video frame it is understood that the surface is corrected by an SfM point cloud derived from multiple temporally-local video frames.
Registering endoscopic videos with CT scans 5 The third process (which we call "fusion") is to combine these multiple SfS textured surfaces (usually 20-30) into a single textured surface (endoscopogram). Fusion is needed because a single 3D reconstructed frame cannot capture the entire mucosal surface seen at endoscopy. For example, one cannot see both the lingual and laryngeal sides of the epiglottis in a single 2D video frame. But because of inevitable patient motion between video frames that are temporally separated, a rigid registration between SfMS surfaces is not satisfactory. Rather, a deformable method is needed. It also proved to be the case that registering these surfaces all together, rather than one-by-one gave better and more consistent results. The registration method to fuse these surfaces needs to deal with non overlap of surfaces and holes due to occlusion and to use both shape and texture information on the surfaces being registered. The method we invented is called Thin-Shell Demons (TSD) [11,12].
Motivated by the demons framework developed by Thirion[13], we regard one surface as an elastic thin shell with additional structural energy that can be attracted via virtual forces produced between the surfaces. The result is a deformation process that tries to align structures similar in geometry and texture in a physically realistic way.
Deformable registration of the planning CT with the endoscopogram
Once the endoscopogram is produced, we must register it to the radiation planning CT scan. The deformable registration method that we have developed for registration of the endoscopogram with a mucosal surface generated from the CT considers mechanical aspects of the deformation and locational and curvature aspects of the surfaces. However, it differs of TSD described because it cannot use texture and because it recognizes that due to resolution differences and occlusions in the video, it estimates where there is disagreement of certain geometric structures. Experimentation has also shown that this variant of TSD gives satisfactory results.
Completing the system
To complete the system, we developed one more piece of software, one that would allow one to draw the gross tumor volume (GTV) onto the reconstructed endoscopogram, view the outline on the original video frames and then on the planning CT scan via registration.
Summary of the approach
A. We first reconstruct a spatially correct textured surface from multiple selected video frames using SfM and SfS.
B. We then register these frame-based textured surfaces with each other (fusion) so as to display the entire pharyngeal surface. We call this fused object an endoscopogram.
C. Once we have completed construction of this endoscopogram, we deformably register it to the treatment planning CT
D. Then we are able to identify any selected points (for example, the tumor interior) on the endoscopic video frames or on the endoscopogram, and compute their corresponding position on the treatment planning CT. This step allows us to complete our goal of bringing endoscopic video-generated data into the treatment planning process.
Testing the registration
Because the thin shell demon approach does not use point correspondences as a means of registration we can test the final registration result by manually selecting corresponding points on video frames and the CT scan and use them to measure the overall accuracy of the global registration under the assumption that there is low human error in making these correspondences. To reduce this error as much as possible, a team of physicians and students worked together to select the corresponding points, using software that displayed the points, both those selected on video and those selected on CT, in both 2D and 3D formats. In addition we extensively tested the accuracy of our system in deformably registering synthetic surfaces with known deformations.
Registering endoscopic videos with CT scans 6. We can also apply synthetic, realistic endoscopogram-to-CT deformations and apply them to the endoscopogram while also smoothing away geometric structures that are typically not seen in the CT. Measurements of the accuracy of the computed deformation between the endoscopogram and the synthetic CT surface can then be done. This eliminates errors due to manual registration. Finally, for clinical images we can view qualitatively how well the anatomic structures in the endoscopogram map onto the CT image slices.
Results
Displays

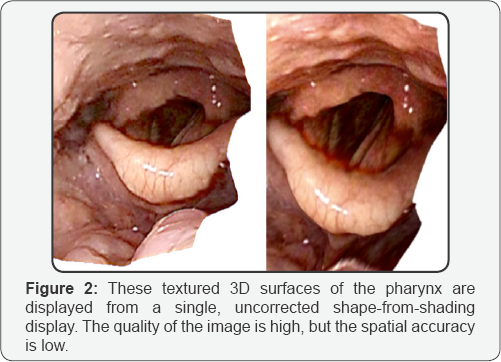
For our initial studies we collected endoscopic videos and CT planning scans, under IRB guidelines, from six anonymized patients. Our nasopharyngoscope is an Olympus model that records full color images at a resolution of 720 x 480 interlaced. CT scans, from a late model Philips CT, have a resolution of 1 x 1 x 3 mm. No special tracking hardware was used to determine endoscope camera position or orientation. (Figure 1) shows a sparse point cloud calculated using structure-from-motion methods. The calculated likely camera trajectories during the procedure are shown in blue. Originally we had hoped to use this kind of surface to register with the CT scan, but local curvature, needed for our deformable registration method, cannot be accurately calculated from such a surface (Figure 2).
These textured 3D surfaces of the pharynx are displayed from a single, uncorrected shape-from-shading display. The quality of the surface is high, but the spatial accuracy is low. Note that the anterior commissure of the vocal folds appears to be attached to the epiglottis in the surface on the right. This is an example of the error that is made due to the presence of occluding surfaces (Figure 3).
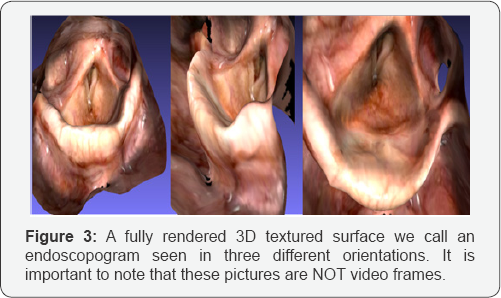
This is an endoscopogram, a fully 3D object, which is seen in three different orientations. In figure 3a one sees the vallecula and lingual surface of the epiglottis. In figure 3b one sees the top of the epiglottis, and the vocal folds and pyriform sinuses come into view. In figure 3c one sees the anterior commissure and laryngeal surface of the epiglottis. The 3D display is fully interactive and thus one can review the entire pharyngeal surface very quickly.
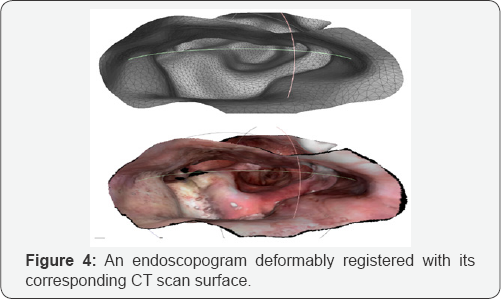
Accuracy of endoscopogram/CT registration
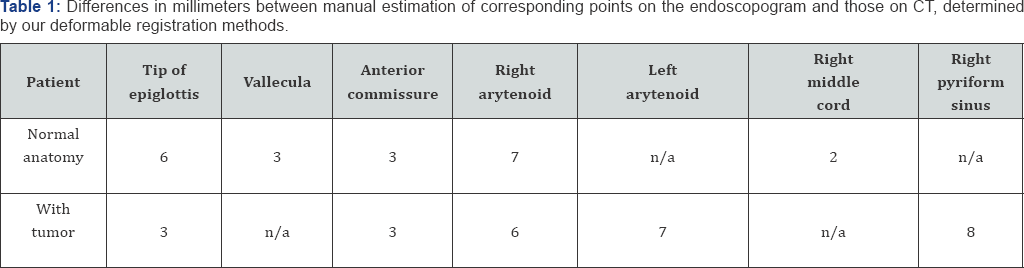
(Figure 4) shows an example of a co-registered endoscopogram and CT. The accuracy of the registration for two patients, one with tumor and one without tumor was tested, using an early version of our method. Table 1, below, shows the results between the team's determination of corresponding points and those determined by the deformable registration. For example, for the anterior commissure the team chose the most likely candidate point on both the CT and the clearest video frame. That particular point was relatively easy to choose on both modalities, but the team found it difficult to agree on the exact locations of many of the other points, particular the tip of the epiglottis because the modalities of endoscopic video and CT are so different and, in that organ at least, there appeared to be a good deal of deformation when the patient was lying down (CT) and sitting up (endoscopy). All measurements below are in millimeters. Registering endoscopic videos with CT scans 7 (Table 1).
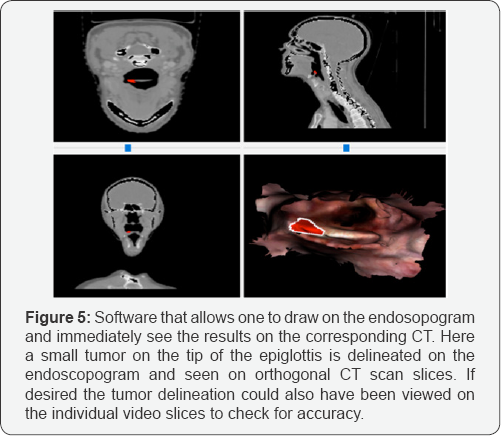
In addition to the foregoing measurements based on manually chosen corresponding points we also measured the closest point distances between the surfaces of the endoscopogram and CT surface from the epiglottis down to the larynx. The RMS distance was 3 millimeters. The maximum distance measurements based on the synthetic deformations on 5 patients and using more recent versions of our methods[14] were under 2 mm. As illustrated in (Figure 5), in the clinical cases the transfer of tumor appeared to be transferred with an accuracy of 1-2mm.
Discussion
Importance of this work
The immediate objective of our work is that of enabling a physician to outline the tumor extent on the endoscopy video and have region bounded by these contours accurately mapped onto the patient's planning CT scan. We do not envision that the importance of this process is that it will routinely result in small (millimeter) changes in GTV delineation so much as that on occasion it will cause substantial changes in GTV delineation. This situation will likely occur when CT and even PET/CT fail to show extensive mucosal spread of the tumor that is obvious on endoscopy. But this is unknown territory, and we have now begun clinical studies to test the above hypothesis.
This will be a 20-patient retrospective pilot study whose purpose is to determine the potential value of using information gathered from endoscopy directly in the treatment process. We will determine if the treatment plan on these 20 patients whose treatment has already been completed could have been improved, geometrically or dosimetrically, by the addition of information from endoscopy. By using this new technology after the fact, it cannot alter patient outcomes in any way and therefore poses no possible medical risk to the patient.
For each of the 20 previously anonymized patients we will locate a complete endoscopy video and the planning CT scan. Using the methods described above we will compute the position of the Registering endoscopic videos with CT scans 8 endoscopically derived GTV (eGTV) and compare it to the CT derived GTV (cGTV). Each plan it will be graded as to whether or not the tumor as seen on the eGTV would have been adequately treated by the plan based only on the cGTV.
In addition to its potential value as an aid to radiation treatment planning, high quality interactive doscopograms such as shown in Figures 3 can also serve a valuable function for medical review. Reviewing an entire endoscopic video is time-consuming and thus not routinely done. However viewing an endoscopogram should allow the clinician to find and study areas of concern quickly and easily. As such, the endoscopogram may become a standard review image for patient follow-up. In addition, the endoscopogram may be used for surgical planning or even to determine the best approach for a biopsy of a worrisome, persistent lesion. Finally, an endoscopogram can be used to detect (and document) small changes in the mucosal surface that occur over time, as seen on serial examinations. This could be registering previously endoscopograms from previous examinations with the present one.
Perhaps the most important consequence of this work will be the eventual routine incorporation of the data contained in all sorts of endoscopic images into any treatment planning or review process. Indeed, we have begun exploratory work on adapting these methods to the more difficult task of colonic reconstruction from colonoscopy in near-real time. Here we hope to provide feedback to the colonoscopist as to what areas of the colonic surface were not adequately visualized so that while the patient is still on the examining table the problem can be remedied. Other anatomic areas of interest that could be reconstructed include esophagoscopy with ultrasound, bronchoscopy, and cystoscopy. Registering endoscopic videos with CT scans 9.
Acknowledgement
We are grateful to NVidia, Inc. for a grant of a GPU unit that is being used in our research. This research was done under the partial support of NIH Grant #R01 CA158925 and a Lineberger Cancer Center tier 1 grant.
References
- Nguyen MH, Wunsche B, Delmas P (2013) Modeling of 3D objects using unconstrained and uncalibrated images taken with a handheld camera. CVIC 274: 86-101.
- Faugeras O, Robert L, Laveau S (1998) 3-D reconstruction of urban scenes from image sequences. CVIU 69(3): 292-309.
- Triggs B, McLauchlan PF, Hartley RI (2002) Bundle adjustment - a modern synthesis. Vision Algorithms: Theory and Practice. LNCS 1883(2000): 298-372.
- Lucas BD, Kanade T (1981) An iterative image registration technique with an application to stereo vision. International Joint Conference on Artificial Intelligence 674-679.
- Tomasi C, Kanade T (1991) Detection and tracking of point features. CMUTR CMU-CS 91-132.
- Lowe DG (1999) Object recognition from local scale-invariant features. Proceedings of the ICCV 2(2): 1150-1157.
- Bay H, Ess A, Tuytelaars T (2008) SURF: Speeded up robust features. CVIU 110(3): 346-359.
- Van Diggelen J (1952) A photometric investigation of the slopes and the heights of the ranges of hills in the Maria of the Moon. Bulletin of the Astronomical Institutes of the Netherlands 11: 283-289.?
- Zhang R, Tsai PS, Cryer JE (1999) Shape-from-shading: A Survey. IEEE Transactions on Pattern Analysis and Machine Intelligence. 21(8): 690-706.
- Price T, Zhao Q, Rosenman J. Shape from motion and shading in uncontrolled environments. Internal report.
- Zhao Q, Price T, Pizer S, Niethammer M, Alterovitz R, et al. (2015) Surface Registration in the Presence of Missing Patches and Topology Change. In Proceedings of Medical Image Understanding and Analysis 201: 8-13.
- Zhao Q, T Price, S Pizer, M Niethammer, R Alterovitz, et al. (2016) The Endoscopogram: a 3D Model Reconstructed from Endoscopic Video Frames. MICCAI 439-447.
- Thirion JP (1998) Image matching as a diffusion process: an analogy with Maxwell's demons. Med Image Anal 2(3): 243-260.
- Zhao Q, Pizer P, Niethammer M, Alterovitz, R, Rosenman J (2017) Orthotropic Thin Shell Elasticity Estimation for Surface Registration.






























