Drivers of Precision Medicine: Liquid Biopsy and Next-Generation Sequencing
James A Posey1 and Chen-Hsiung Yeh2*
1Sidney Kimmel Cancer Center, Thomas Jefferson University, USA
2Circulogene Theranostics, USA
Submission: December 11, 2016; Published: December 15, 2016
*Corresponding author: Chen-Hsiung Yeh, Circulogene Theranostics, 3125 Independence Drive, Suite 301, Birmingham, AL USA 35209; Tel: 1-205-234-0128
How to cite this article: James A. P, Chen-Hsiung Y. Drivers of Precision Medicine: Liquid Biopsy and Next-Generation Sequencing. Canc Therapy & Oncol Int J. 2016; 2(4): 555595. DOI: 10.19080/CTOIJ.2016.02.555595
cfDNA: Cell-Free DNA; NGS: Next-Generation Sequencing; WGS: Whole Genome Sequencing; WES: Whole Exome Sequencing; GH: Guardant Health; CT: Circulogene TheranosticsAbbreviations
Editorial
Targeted therapy specifically aims at tumor genetic alterations s the hallmark of precision medicine. Companion diagnostic testing utilized to determine the presence or absence of certain oncogenic mutations prior to targeted treatment under the current medical guidelines will enable improved clinical outcome, and thus serves as a vital component for precision medicine. Standard clinical practice to assess genetic mutations in cancer patients has historically been through direct sampling of tumor tissue with biopsy or surgical resection. Unfortunately, tissue biopsy is one-time single-site sampling, painful, costly, and can miss the dynamics of tumor clonally evolution during disease progression. The emerging liquid biopsy by circulating cell-free DNA (cfDNA), on the other hand, can fulfill the gap, not only providing actionable information for decision-making prior to treatment, but also longitudinal surveillance during and after the treatment regimen to assess for patient response, drug resistance[1-3] and cancer recurrence [4].
Genomic content of cfDNA that is collected from a befouled specimen must be tested with an appropriate analytical method in order to identify existing mutant alleles. Traditional methods for mutation detection include assays such as quantitative PCR and Sanger sequencing [5,6]. PCR-based assays are sensitive but provide only limited information on a handful of genes and hotspot mutations, whereas Sanger sequencing does not possess the sensitivity adequate for detection in the context of liquid biopsy [7]. Modern next-generation sequencing (NGS)-based technology involves sequencing DNA in a highly multiplex and parallel fashion with higher sensitivity [8].
Whole genome sequencing (WGS) or exome sequencing (WES) of tumor profiling using NGS is an unbiased approach that provides extensive and comprehensive genomic information about cancer. Although they can provide genetic resolution both at the single nucleotide level, as well as detect structural alterations such as large rearrangements, gross deletions and duplications [9-11],the cost of these sequencing are still too high and our current knowledge bases are too small for routine clinical application. Targeted gene panels are currently the best option for CLIA clinical utility in the field of liquid biopsy, as they allow multiple action able genes to be analyzed and can provide enough depth of coverage to detect minor allele frequencies in a clinically-relevant and cost-effective manner [12,13]. NGS gene panels can also eliminate the need for reflex testing and preserves precious specimens by limiting the number of tests required for full characterization. In order for NGS liquid biopsy diagnostics to be accurate and affordable, a balance must be reached between panel size and the level of multiplexing.
There are currently a handful of CLIA liquid biopsy companies who offer cfDNA NGS-based genetic testing. As more NGS gene panels begin to enter the market, it is important for users to understand the benefits and limitations of these methods and technologies being utilized. There are many nuances in designing NGS gene panels for liquid biopsy tumor profiling, and multiple options exist for each component of the workflow. Numerous factors can impact the accuracy, sensitivity and specificity of cfDNA sequencing including the sample preparation, target enrichment, library construction, sequencing platform and bioinformatics tool. Some degree of standardization of cfDNA NGS assays used for the detection of somatic mutations is essential before their deployment in clinical practice. Towards this direction, in addition to the concordance studies between tissue and liquid biopsies, comparison of cfDNA-based testing between different methodologies on different platforms in different laboratories are needed.
In the first side-by-side comparison study, the NGS results for a12-patient cohort from 2 commercially available gene panels (70 genes and 50 genes) are analyzed. These samples were processed and tested in two independent CLIA liquid biopsy laboratories, Guardant Health (GH) and Circulogene Theranostics (CT), with distinct sample volume input, sample preparation, library construction, NGS platform and different mutation calling bioinformatics. GH and CT tests resulted in a very similar 40- 50% rate of clinically relevant actionable information, i.e., either FDA drugs or active clinical trials. There are 2 cases reported “no mutation” by both tests. TP53 is the most frequent mutated gene identified in 41.7% (5/12) of cases. In 5 cases, there is at least one gene mutation matched. In another 5 cases, mutations did not match each other. It is noteworthy that there are 43 genes commonly covered by both panels, however, Guardant’s test surveyed all axons of these 43 genes, while Circulogene’s assay only scanned hotspot mutations within these genes. Nevertheless, the overall concordance between both test results was fairly high at the gene level, because the denominator in this calculation was 516 (43 genes for 12 patients), and the majority of genes were called wild-type alleles (no mutation detected).
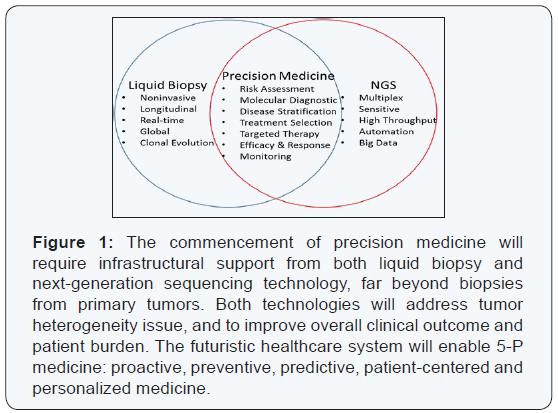
A key issue in clinical oncology practice is the ability to accurately interpret what is truly clinically meaningful and actionable on a report from a commercially available NGS test. The majority of cancer care is not delivered at tertiary comprehensive cancer centers, but rather in busy community oncology practices. Treating physicians will have time constraints to read and digest NGS reports, so there is a need for clear, concise reporting of clinically relevant targets. With the integration of additional levels of content enhancement and scientific/clinical evidence, liquid biopsy NGS reports can eventually deliver patient-centered context and substance that will lead to better healthcare outcomes and toward the goal of precision medicine (Figure 1).
References
- Yun CH, Mengwasser KE, Toms AV, Woo MS, Greulich H, et al. (2008) The T790M mutation in EGFR kinase causes drug resistance by increasing the affinity for ATP. Proc Natl Acad Sci U S A 105(6): 2070-2075.
- Gazdar AF (2009) Activating and resistance mutations of EGFR in nonsmall- cell lung cancer: role in clinical response to EGFR tyrosine kinase inhibitors. Oncogene 28(1): S24-S31.
- Massarelli E, Varella-Garcia M, Tang X, Xavier AC, Ozburn NC, et al. (2007) KRAS mutation is an important predictor of resistance to therapy with epidermal growth factor receptor tyrosine kinase inhibitors in non-small-cell lung cancer. Clin Cancer Res 13(10):2890- 2896.
- Crowley E, Di Nicolantonio F, Loupakis F, Bardelli A (2013) Liquid biopsy: monitoring cancer-genetics in the blood. Nat Rev Clin Oncol 10(8): 472-484.
- Bárbara Angulo, Elena García-García, Rebeca Martínez, Ana Suárez- Gauthier, Esther Conde, et al. (2010) A commercial real-time PCR kit provides greater sensitivity than direct sequencing to detect KRAS mutations: A morphology-based approach in colorectal carcinoma. J Mol Diagn 12(3): 292-299.
- Lopez-Rios F, Angulo B, Gomez B, Mair D, Martinez R, et al. (2013) Comparison of testing methods for the detection of BRAF V600E mutations in malignant melanoma: Pre-approval validation study of the companion diagnostic test for vemurafenib. PLoS One 8(1): e53733.
- Michaela Angelika Ihle, Jana Fassunke, Katharina König, Inga Grünewald, Max Schlaak, et al. (2014) Comparison of high resolution melting analysis, pyrosequencing, next generation sequencing and immunohistochemistry to conventional Sanger sequencing for the detection of p.V600E and non-p.V600E BRAFmutations. BMC Cancer 14:13.
- Masoudi-Nejad A, Narimani Z, Hosseinkhan N (2013) Next generation sequencing and sequence assembly. Springer New York, USA, (Volume 4), pp. 11-39.
- Erin D. Pleasance, R. Keira Cheetham, Philip J. Stephens, David J. McBride1, Sean J. Humphray, et al. (2010) A comprehensive catalogue of somatic mutations from a human cancer genome. Nature 463: 191- 196.
- Chen K, Wallis JW, McLellan MD, Larson DE, Kalicki JM, et al. (2009) Breakdancer: An algorithm for high-resolution mapping of genomic structural variation. Nat Methods 6(9): 677-681.
- Chiang DY, Getz G, Jaffe DB, O’Kelly MJ, Zhao X, et al. (2009) Highresolution mapping of copy-number alterations with massively parallel sequencing. Nat Methods 6(1): 99-103.
- Wagle N, Berger MF, Davis MJ, Blumenstiel B, Defelice M, et al. (2011) High-throughput detection of actionable genomic alterations in clinical tumor samples by targeted, massively parallel sequencing. Cancer Discov 2(1): 82-93.
- Pritchard CC, Salipante SJ, Koehler K, Smith C, Scroggins S, et al. (2014) Validation and implementation of targeted capture and sequencing for the detection of actionable mutation, copy number variation, and gene rearrangement in clinical cancer specimens. J Mol Diagn 16(1): 56-67.






























