Application Of (Bio)Chemical Engineering Concepts and Tools to Model Grcs, and Some Essential CCM Pathways in Living Cells
Gheorghe Maria1,2,*
1Department of Chemical and Biochemical Engineering, Politehnica University of Bucharest, Polizu Str. 1-7, Bucharest 011061, Romania
2Romanian Academy, Chemical Sciences section., Calea Victoriei 125, Bucharest 010071, Romania
Submission:November 06, 2023; Published:December 20, 2023
*Corresponding author:Gheorghe Maria, Dept of Chemical and Biochemical Engineering, Politehnica University of Bucharest, Polizu Str. 1-7, Bucharest 011061, Romania, email: gmaria99m@hotmail.com; https://en.wikipedia.org/wiki/Gheorghe_Maria
How to cite this article:Gheorghe M. Application Of (Bio)Chemical Engineering Concepts and Tools to Model Grcs, and Some Essential CCM Pathways in Living Cells. Curr Trends Biomedical Eng & Biosci. 2023; 22(1): 556080. DOI:10.19080/CTBEB.2023.22.556080
Summary
Biologically catalyzed reactions (with enzymes, or living cell cultures) can successfully replace complex chemical syntheses, being more selective, by using milder reaction conditions, and generating less waste. As proved by the recent literature, the developed in-silico (math-model-based) numerical analysis of such biochemical/biological systems turned out to be a beneficial tool to (i) off-line determine optimal operating policies of complex multi-enzymatic or biological reactors with a higher precision and predictability, or (ii) to design GMO (genetically modified micro-organisms) of desired characteristics for various uses. This work presents a holistic ‘closed loop’ approach that facilitate the control of the in vitro through the in silico development of dynamic models for living cell systems, by deriving deterministic modular structured cell kinetic models (MSDKM) (with continuous variables, and based on cellular metabolic reaction mechanisms). The ever-increasing availability of experimental (qualitative and quantitative) information about the tremendous complexity of cell metabolic processes, stored in large bio-omics databanks (including genomic, proteomic, metabolomic, fluxomic cell data for various micro-organisms), but also about the bioreactors’ operation necessitates the advancement of a systematic methodology to organise and utilise these data.
Keywords:Biochemical engineering concepts applied in bioinformatics; Deterministic modular structured cell kinetic model (MSDKM); Hybrid structured modular dynamic (kinetic) models (HSMDM);Whole cell variable cell volume (WCVV) modelling framework; Whole cell constant cell volume (WCCV) modelling framework; Individual gene expression regulatory module (GERM); Genetic regulatory circuits (GRC), or networks (GRN); Chemical and biochemical engineering principles (CBE), Rules of the control theory of nonlinear systems (NSCT).
Abbreviations:ADP: Adenosin-diphosphate; AMP: Adenosin-monophosphate; ATP: Adenosin-triphosphate; BCE- (Bio)chemical engineering; CBE: chemical and biochemical engineering; BR: Batch reactor; CABEQ Jl: Chemical and biochemical engineering journal; CCM- Central carbon metabolism; CIT: Citrate; CSTR: Continuous stirred tank reactor; FBA: Flux balance analysis; FBR: Fed-batch bioreactor; G: The active Gene (DNA); GRC, GRN: Genetic regulatory circuits; GERM: Individual Gene expression regulatory module; GMO: Genetically modified micro-organisms; GP: The inactive complex of G with the transcription factor P (its encoding protein in the reduced model here); GRC, GRN: Genetic regulatory circuits (GRC), or networks; GS- Genetic switch; HSMDM: Hybrid structured modular dynamic (kinetic) models; L: Species at which regulatory element acts; M: mRNA; MCA: Metabolic control analysis; Met (MetG, MetP): Metabolites (lumped DNA and protein precursor metabolites, respectively); MINLP- Mixed-integer nonlinear programming; MSDKM: Deterministic modular structured cell kinetic model; MFA: Metabolic flux analysis; NLP: (non)linear programming problem [48]; nM: Nano-moles/L, nano-molar (i.e. 10-9 mol/L concentration); NG: Negligible; NSCT - the control theory of nonlinear systems; Nut (NutG, NutP): Nutrient (external nutrients imported to produce metabolites involved in the G and P synthesis respectively); ODE: Ordinary differential equations set; P: Protein; PTS: Phosphotransferase GLC import system; PPP: Pentose-phosphate pathway; SCR: Semi-continuous bioreactor; SNP: Single nucleotide polymorphisms; SUCC: Succinate; Re(x): Real part of ”x” variable; TF: Transcription factor; TCA: Citric acid cycle (or tricarboxylic acid cycle); TPFB: Three-phase fluidized bioreactor; TRP: Tryptophan; QSS: Quasi steady-state; WC: Whole cell; WCCV: Whole cell of constant volume hypothesis; WCVV: Whole cell of variable volume hypothesis; [.]: Concentration.
Introduction
This work is aiming to prove the feasibility and the advantages of using the classical and novel concepts and numerical tools of the chemical and biochemical engineering (CBE) to develop MSDKM of the extended cell-scale CCM-based (central carbon metabolism), and of genetic regulatory circuits (GRC) / networks (GRN). These extended kinetic models will be further linked to those of the bioreactor dynamic models (including macro-scale state variables), thus resulting hybrid structured modular dynamic (kinetic) models (HSMDM) proved to successfully solve more accurately difficult bioengineering problems.
In such HSMDM, the cell-scale model part (including nano-level state variables) is linked to the biological reactor macro-scale state variables for improving the both model prediction quality and its validity range. By contrast, the current (classical/default) approach in biochemical engineering and bioengineering practice for solving design, optimization and control problems based on the math models of industrial biological reactors is to use unstructured Monod (for cell culture reactor) or Michaelis-Menten (if only enzymatic reactions are retained) global kinetic models by ignoring detailed representations of metabolic cellular processes. The applied engineering rules to develop MSDKM and HSMDM dynamic math models presented in the 1-st and 2-nd parts of this work are similar to those used in the CBE, and in the control theory of nonlinear systems (NSCT).
This 1-st part of the paper presents some general concepts of CBE, of the NSCT, and of Bioinformatics used to derive MSDKM and HSMDM models, with continuous variables and based on cellular metabolic reaction mechanisms. Such extended structured cell math (kinetic) models consider, with a degree of detail suitable to the each approached case study, the cellular key-metabolic reactions and the cell key-species dynamics. These structured models can satisfactorily represent the key steps of the CCM at a cell scale, by also can include GRC-s responsible for the CCM syntheses regulation, besides reaction modules responsible for the synthesis of cellular metabolites of interest for the industrial biosynthesis. Special attention is paid to the conceptual and numerical rules used to construct various individual GERM-s kinetic models, but also various GRC-s (e.g. toggle-switch, amplitude filters, modified operons, etc.) modular kinetic models from linking individual GERM-s.
The 2-nd part of the paper will briefly reviews the ’whole-cell of variable-volume’ (WCVV) modelling framework introduced and promoted by the author in previous works. Also, this 2-nd part points-out the features of the deterministic WCVV models, and its advantages when simulating GERM-s, and GRC-s dynamics in living cells, by contrast to the classical (default) WCCV (whole-cell constant-volume modelling framework); P.I.-s of GERM-s; rules to link GERM-s when modelling GRC-s, and other related theoretical aspects necessary to construct MSDKM and HSMDM models.
The 3&4 parts of the paper proves, by means of several demonstrative relevant examples the superiority of using MSDKM and HSMDM dynamic math models when solving various bioengineering problems (i) to in-silico off-line optimize the operating policy of various types of bioreactors, and (ii) to in-silico design/check some GMO-s of industrial use able to improve the performances of several bioprocess/bioreactors.
By contrast, by considering only the macroscopic key-variables of the process (biomass, substrate, and product concentrations), the unstructured (apparent, global) math models do not adequately reflect the metabolic changes of the bioreactor biomass, being inadequate to accurately predict the cellular response to the medium disturbances through the self-regulated cellular metabolism. These classical global/ unstructured dynamic models may be satisfactory for an approximate modeling of the biological process, but not for modeling of cellular metabolic processes, and they can not make any correlation between the bioreactor operation and the continuous adaptation of the biomass metabolism to the variable conditions of the bioreactor. Even worst, as proved by the author in previous papers, such global models may lead to biased and distorted conclusions about the GERM’s performances, thus making difficult the modular constructions of GRC-s by linking individual GERM-s.
Introduction
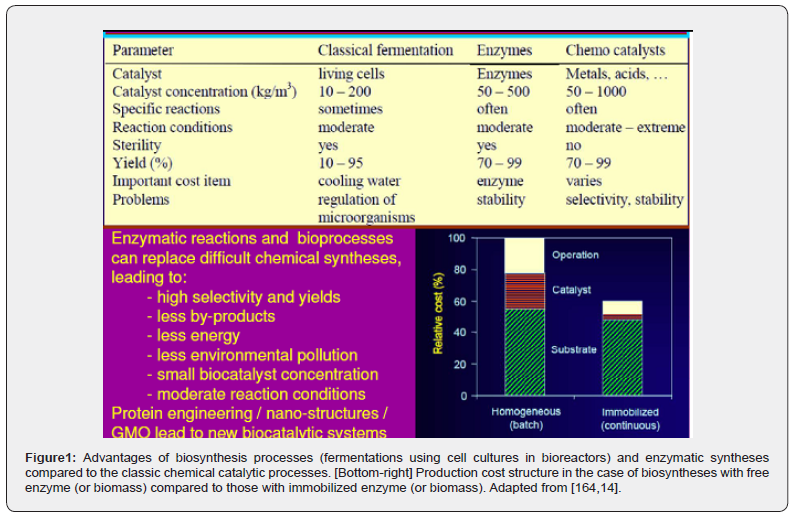
In the last decades, there has been a tendency to replace the complex processes of fine chemical synthesis, highly energy-consuming and generating large amounts of toxic waste, with biosynthesis processes (using isolated and purified enzymes, or cell cultures as bio-catalysts). The motivation is given by the multiple advantages offered by enzymatic processes (Figure 1): i) very high selectivity; ii) very high conversion; iii) does not generate toxic by-products; iv) very mild reaction conditions, easy to achieve without high costs (low temperatures of 20-60°C, normal pressure, pH within controllable limits). Thus, in recent years, a significant number of enzymatic or biological industrial processes have been reported [1-4] in order to obtain chemical products/derivatives in the fine organic synthesis industry, in the pharmaceutical industry, in the food industry or in the detergent industry, by using various bioreactors with cell or enzyme cultures [1,4]. Among these new processes are the production of derivatives of monosaccharides, organic acids, alcohols, amino acids, etc., using mono- or multi-enzymatic reactors, or bioreactors with cell cultures used in the production of yeast, food additives, recombinant proteins (enzymes, vaccines), biopolymers [1,2,5]. The development of a sustainable biological process must consider several aspects related to the characteristics of the biocatalyst, the integration of the process and the minimization of costs, satisfying economic, environmental / safety and social objectives [6-8].
The current approach in biochemical engineering and bioengineering practice for solving design, optimization and control problems based on the math models of industrial biological reactors is to use unstructured Monod (for cell culture reactor) or Michaelis-Menten (if only enzymatic reactions are retained) by ignoring detailed representations of metabolic cellular processes. The engineering rules applied are similar to those used in the chemical and biochemical engineering (CBE), and in the control theory of nonlinear systems (NSCT). However, by considering only the macroscopic key-variables of the process (biomass, substrate, and product concentrations), these unstructured (apparent) math models do not adequately reflect the metabolic changes of the bioreactor biomass, being inadequate to accurately predict the cellular response to the medium disturbances through the self-regulated cellular metabolism. These global dynamic models may be satisfactory for an approximate modeling of the biological process, but not for modeling of cellular metabolic processes, and they casn not make any correlation between bioreactor operation and the continuous adaptation of biomass metabolism to the variable conditions of the bioreactor.
The current trend to more accurately solve such engineering problems is to use deterministic modular structured cell kinetic models (MSDKM), with continuous variables, and based on cellular metabolic reaction mechanisms, that consider, with a degree of detail suitable to the each approached case study, the cellular metabolic reactions and the cell key-species dynamics. These structured models can satisfactorily represent the key steps of the central carbon metabolism (CCM) at a cell scale, by also including reaction modules responsible for the synthesis of cellular metabolites of interest for the industrial biosynthesis. As proved ny Maria [11-14,32], and Yang et al. [175], the modular structured kinetic models can reproduce the dynamics of complex metabolic syntheses inside living cells. This is why, the modular GRC and CCM dynamic models, of an adequate mathematical representation, seem to be the most comprehensive mean for a rational design of the regulatory GRC with desired behaviour [110]. The same MSDKM can satisfactorily simulate, on a deterministic basis, the self-regulation of cell metabolism for its rapid adaptation to the changing bioreactor reaction environment, by means of complex "genetic regulatory circuits" (GRC-s), which include chains of individual „gene expression regulatory modules" (GERM-s).
In this context, this work shortly review the essential CBE principles and rules used to elaborate MSDKM, but also the so-called ’’Hybrid structured modular dynamic (kinetic) models” (HSMDM) [9,10] that combine the characteristics of the cellular metabolic process involving species participating to the essential reaction modules of CCM at a nano-scopic level, with the macro-scopic processes involving the state variables of the industrial bioreactor. In this way, more accurate predictions are obtained both for the dynamics of the biological process at the cellular level, and for the dynamics of the operating parameters of the analyzed industrial bioreactor. The immediate applications of these MSDKM and HSMDM refer to (i) the more precise determination of the optimal operating policy of an industrial bioreactor, and (ii) facilitates, by means of an in-silico numerical analysis, determination of GMO-s with a cell metabolism of desired characteristics.
In this first part of this work, the general concepts to construct MSDKM models with continuous variables are presented by using CBE and NSCT principles/concepts and rules. Special attention is paid to the conceptual and numerical rules used to build-up modular CCM kinetic models, in direct connection to various individual GERM-s kinetic models, but also to various GRC-s (e.g. toggle-switch, amplitude filters, operons expression, etc.) modular kinetic models by linking a couple of GERM-s. To do such a complex modelling work in a consistent way, this part also briefly reviews the novel „Whole cell variable cell volume” (WCVV) modelling framework introduced and promoted by [11-18], as an essential modelling instrument to develop more realistic and precise MSDKM-s and HSMDM-s. Besides presenting the WCVV deterministic model hypotheses, this paper points-out its advantages when simulating GERM-s, and GRC-s dynamics in living cells, in a holistic approach, by contrast to the classical (default) WCCV (whole-cell constant-volume modelling framework). The WCVV discussion is extended by briefly reviewing the P.I.-s of GERM-s; and the rules to link GERM-s when modelling/build-up GRC-s.
However, industrial bioprocesses still have a limited spread due to the high costs of enzyme/biomass isolation and stabilization on a suitable support, as well as its high sensitivity in relation to the operating conditions, the rather low reproducibility of the biological process due to biomass changes from one cell cycle to another, and of the difficult controllability of the bioreactor. However, many of these drawbacks can be overcome by an efficient immobilization of the biomass on suitable supports, by using suitable GMO-s with superior catalytic activity, and/or by optimizing the working conditions and the operation mode of the selected biological reactor, by using an advanced off-line in-silico analysis of the engineering part of bioprocess development based on effective math models and numerical algorithms. This last alternative is briefly reviewed in the 2-nd part of the paper.
As proved in the literature, the in-silico (math/kinetic model-based) numerical analysis of biochemical or biological processes by using MSDKM or HSMDM models are proved to be not only an essential but also an extremely beneficial tool for engineering evaluations aiming (i) to determine with a higher accuracy the optimal operating policies of complex multi-enzymatic reactors, [5,19-23], or of bioreactors including the biomass adaptation to the variable bioreactor environment over hundreds of cell cycles [2,9,24-26], or even (ii) to easier and quickly simulate and analyse the performances/ characteristics of various GMO-s alternatives, by using the “metabolic flux analysis” (MFA), [26-30], together with the gene-knock-out technique) [9,10,14,30-32].
Biochemical Reactor Case
To solve engineering problems for this case, the trend is to use complex multi-enzymatic systems which successfully replace complex chemical syntheses, by using milder reaction conditions, and generating less waste. Progresses in enzymes immobilization, and genetic engineering lead to prolonging the biocatalyst life and efficiency [169-171].
Even if the multi-enzymatic systems are advantageous, the engineering part used to optimize such a complex process is not an easy task because it must account for the interacting enzymatic reactions, enzymes deactivation kinetics (if significant), multiple and often opposed optimization objectives, technological constraints, and uncertainties coming from multiple sources (model / constraints inaccuracies, disturbances in the control variables), and a highly nonlinear process dynamics [2,19,20,22-24,33-35]. All these parametric/model/data uncertainties require updating (with a certain frequency) the enzymatic process model, the optimal operating policies of the reactor being determined by using rather deterministic (model-based) optimization rules [36]. Multi-objective criteria, including economic benefits, operating and materials costs, product quality, etc., are used to off-line, or to on-line derive feasible optimal operating/control policies for various bioreactor types [34] by using specific numerical algorithms [20,22,23,25,32,35,37-39].
Biological reactor case
In thw last decades, the trend is to use biological processes conducted in complex biological reactors to successfully replace complex chemical syntheses, by using milder reaction conditions, and generating less waste. Progresses in biomass immobilization, and genetic engineering lead to prolonging the biocatalyst life and efficiency [172].
To solve engineering problems for this case, development of extended cell-scale CCM / GRC-s structured MSDKM or HSMDM kinetic models on a deterministic basis to adequately simulate in detail the cell metabolism self-regulation, cell growth, and its replication for such an astronomical cell metabolism complexity is practically impossible due to the lack of structured and comprehensive experimental information, and computational limitations. A review of some trials are reviewed by [40,11,12,14,32]. That is because the cell metabolism is highly sophisticated, involving O(103-4) components, O(103-4) transcription factors (TF-s), activators, inhibitors, and at least one order of magnitude higher number of (bio)chemical reactions, all ensuring a fast adaptation of the cell metabolism to the changing environment through complex genetic regulatory circuits (GRC-s), that includes individual or chains of „gene expression regulatory modules of reactions” (GERM-s), genetic switches (GS), operon expression, etc. [11,12,14,32]. The cell is highly responsive to the environmental stimuli and highly evolvable by self-changing its genome/proteome and metabolism, that is the stoichiometry and the reaction rates (fluxes) of the enzymatic reactions to get an optimized and balanced growth by using minimum resources (nutrients/substrates).
In spite of such tremendous modelling difficulties, the development of structured reduced deterministic (rather than stochastic) MSDKM or HSMDM models on a deterministic basis reported significant progresses over the last decades [10,11,12,14,32,31,41-43]. Such reduced cell models are able to adequately reproduce the dynamics of some CCM complex metabolic syntheses [10,31,32,40,44,45,46,47], but also the dynamics of some GRC-s [9,11,12,14,32,] tightly controlling the metabolic processes. Even if they are rather based on sparse information from various sources, unconventional statistical identification, and lumping algorithms [11,12,14,32,15,16,48,49], such structured reduced deterministic kinetic models have been proved to be extremely useful for in-silico (a) analyse and characterize the cell CCM, (b) for designing novel GRC-s conferring new properties/functions to the mutant cells, or (c) for engineering bioreactor evaluations [10,24,11,12,14,32, 50] (see the part 2 of this paper).
The current (default) approach to solve the model-based design, optimization and control problems of industrial biological reactors is the use of unstructured models of Monod type (for cell culture reactors)[3] or of Michaelis-Menten type (if only enzymatic reactions are retained)[21,22,23,35,51,52] that ignores detailed representations of cell processes. The applied engineering rules are similar to those used by the CBE and inspired from the NSCT [25,34,53,54-60].
However, by accounting for only key process variables (biomass, substrate and product concentrations), these global (unstructured) dynamic models do not properly reflect the metabolic changes, being unsuitable to accurately predict the cell response to environmental perturbations by means of (self-)regulated cell metabolism [9,61,11,12,14,32].
The alternative is to use structured kinetic models, by accounting for cell metabolic reactions and component dynamics. Such deterministic models lead to a considerable improvement in the predictive power, with the expense of incorporating a larger number of species mass balances including parameters (rate constants) difficult to be estimated from often incomplete data (by using complex NLP, MINLP, and statistical procedures [48,168,32,49,15,16]), and, consequently, difficult to be used for industrial scale purposes [9,10,15,16,32,62,].
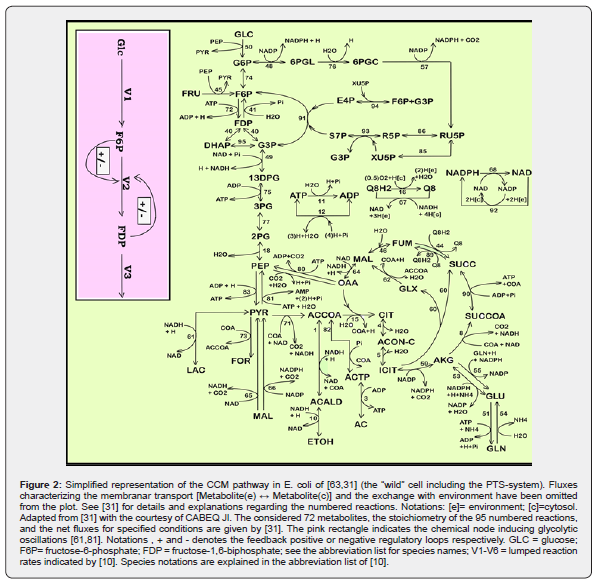
As a result, an impressive large number of valuable structured deterministic kinetic models (based on a mechanistic description of the metabolic enzymatic reactions tacking place among individual or lumped species of the cell) have been proposed in the literature to simulate the cell CCM dynamics, with including tenths-to-hundreds of key species. Here, it is worth mentioning the E. coli model of [63] used by [31,44,64,65,66,67,68] for various purposes, or the S. cerevisiae glycolysis model of [69], or the JWS platform of [70], or the MPS platform of [71] to simulate cell metabolism (dynamics and/or stationary fluxes), to mention only few of them. Simulation platforms, such as E-cell of [72,73], or V-cell of [74], accounting for thousands of species and reactions, display extended capabilities to predict the dynamics of the cell metabolism under various conditions, based on EcoCyc, KEGG, Prodoric, Brenda and other bio–omics databanks (review of [14]). A worthwhile CCM-based dynamic or stationary models were reported by Maria [10,31,44] based on a lumped reaction pathway schematically represented in (Figure 2).
Some of structured deterministic CCM kinetic models have been reviewed by [11,12,14,32]. Deterministic MSDKM kinetic models using continuous variables has been developed by Maria [44] for the glycolysis, and by [64,65,75,76,77,78] for the CCM in bacteria of industrial interest. Such MSDKM models can adequately reproduce the cell response to continuous perturbations. The MSDKM cell model structure and size is adapted based on the available bio–omics databanks, and experimental information. Even if such extended structured models are currently used only for research purposes, being difficult to be identified, it is a question of time until they will be adapted in the form of HSMDM–s for industrial / engineering purposes. As proved by this work, and by several examples given in the part 2 of this paper, already significant progresses have been reported in this respect.
The Parts 1 and 2 of this work presents a holistic ‘closed loop’ approach for the development of models of biological systems [79]. The ever-increasing availability of experimental (qualitative and quantitative) information, at the cell metabolism level, but also about the bioreactors’ operation necessitates the advancement of a systematic methodology to organise and utilise these data. The resulted HSMDM–s were proved to successfully solving more accurately difficult bioengineering problems. In such HSMDM-s, the cell-scale model part (that is the nano-level state variables) is linked to the biological reactor macro-scale state variables for improving the both model prediction quality and its validity range. The case studies presented and discussed in the parts 3&4 of this work prove this engineering aspect.
In fact, the use of HSMDM hybrid models realizes a valuable combination of the process modelling-scales, by linking unstructured with structured process characteristics to generate more precise predictions (see the examples given by [9,10,24]). Basically, the HSMDM hybrid models use a two-level hierarchy: the bioreactor macroscopic state variables linked with the nano-scale variables describing the cell key metabolic processes, and cell syntheses of practical interest (usually resulting in excretable valuable metabolites).
In fact, such a HSMDM (hybrid structured cell dynamic model) must include only the pathway responsible for the target metabolite synthesis, linked to the essential parts of the CCM, that is the lumped modules of the cell core. These CCM core modules include, among others, the glycolysis, the glucose (GLC) uptake system [i.e. the phosphotransferase (PTS), or an equivalent system], the “adenosine triphosphate” (ATP)-recovery system (i.e. the ATP-use-recovery cycle), TCA (tricarboxylic acid, or citric acid cycle), PPP (Pentose-phosphate pathway). Of course, additional CCM reaction pathways modules must be added to the structured cell model (in a detailed or reduced/lumped form) if their presence is absolutely necessary to derive consistent CCM simulations. See for instance the discussion of [10,24,31,32,44,46,61,80].
A special interest was given to the accurate modelling of the glycolysis dynamics and its self-regulation [44,61,81-83] as long as this core-module of the CCM includes intermediates which are starting nodes for the internal production of lot of cell metabolites (e.g. succinate (SUCC), citrate (CIT), amino-acids like cysteine, lysine, phenyl-alanine, tryptophan (TRP), etc., see Figure 3) [2,31,47,83]. This need to have good quality structured cell models to simulate the dynamics of the bacteria CCM (and its regulation via cell GRC-s/GRN-s) became a subject of very high interest over the last decades, allowing an in-silico design of GMO-s with desirable characteristics of various applications in the biosynthesis industry, civil engineering, medicine, and other fields [11,12,14,32].

MSDKM and HSMDM models advantages
Even if such complex / extended dynamic model requires more experimental and computational efforts to be built-up, as proved by the approached case studies in the parts 3&4 of this work, the resulted deterministic modular structured cell kinetic model (MSDKM) of the key-parts of cell CCM and GRC-s (of interest), as well as the hybrid (bi-level, that is the cell-species state variables, linked to the macro-level bioreactor state variables) dynamic models (HSMDM) present a large number of advantages compared to the classical (default) unstructured models of Monod type (for cell culture bioreactors), or of Michaelis-Menten type (if only enzymatic reactions are retained) that ignores detailed representations of cell metabolic processes. Thus, among the multiple advantages of HSMDM models are to be mentioned the following most relevant ones:
(i).- A higher prediction detailing degree. Thus,
(i-a). For the case study referring to the HSMDM used to design and simulate the GRC responsible for mercury (mer) operon expression in the wild or GMO design E. coli cell (see the part 4 of this work), a higher prediction detailing degree is reported, that is predicted dynamics of [26(cell species) + 3(bulk species)] vs. only [3 (bulk species)] by a classical macroscopic SCR- TPFB unstructured (global) dynamic model, while covering a wider range of input [Hg2+] loads, with also using cloned E. coli cells with various amounts of mer-plasmids [Gmer]. Here SCR denotes a semi-continuous bioreactor, that is a CSTR (continuous stirred tank reactor) of variable feeding, of TPFB (three-phase fluidized bioreactor) constructive type [9,173].
(i-b). For the case study referring to the HSMDM used to simulate the dynamics of the species at cell-level, and bulk-phase level in a fed-batch bioreactor (FBR), but also to maximize the tryptophan (TRP) production (see the part 3 of this work), a higher prediction detailing degree is reported, by characterizing the dynamics of [11(cell species) + 4(bulk species)] vs. only [3 (bulk species)] by a classical macroscopic FBR global model, while covering a wider range of control variables, and allowing the design/check of various GMO E. coli cells strains.
(ii).- Prediction of the inner cell key-species reaction rates (different from the apparent rates of the substrates and excreted products observed in the bioreactor). See the above case study (i-a) of [9,84,85], and the case study (i-b) of [10,24].
(iii).- The structured [cell-and-bioreactor] hybrid model predictions can cover a wider range of input/control variables of the bioreactor. Thus, for the above case study (i-a), the HSMDM of Maria and Luta [9] realizes a higher prediction detailing degree, that is simulated dynamics of [26(cell species) + 3(bulk species)] vs. the dynamics of only [3 (bulk) state variable] predicted by a classical macroscopic SCR- TPFB model. Such a an advanced HSMDM predictions can cover a wider range of input [Hg2+] loads (0–100 mg/L), and can be used to design cloned E. coli cells with various amounts of mer-plasmids [Gmer] in a wider feasible range of (3–140 nM).
(iv).- The HSMDM model can predict the bacteria metabolism adaptation to environmental changes over dozens of cell cycles, and the effect of cloning wild E. coli cells to obtain (GMO) with modified characteristics/properties/behaviour under stationary or perturbed bioreactor operating conditions. See the case study (i-a) for details (Part-4).
(v).- The extended HSMDM can offer predictions of a higher accuracy when they are used to in-silico (model-based) engineering developments (bioreactor design, or its off-line optimization) compared to unstructured/global models (see the parts 3&4 of this work, for several bioreactor cases). For instance, (i-b) the HSMDM could better predict the optimal time stepwise feeding policy of a fed-batch reactor (FBR) to increase the tryptophan (TRP) production (Part 3) [10,24]. In the above case study (i-a), the extended HSMDM allows optimizing the operating policy of a SCR-TPFB regarding the biomass concentration, the inlet feed flow-rate, the inlet [Hg2+], the immobilized biomass support size, and the [Gmer] concentration in the used cloned E. coli cells [Maria and Luta, 2013; Maria et al., 2013].
(vi).- A similar procedure of using hybrid dynamic models can be applied to optimize the multi-enzymatic systems, by linking several interacting complex enzymatic reactions to the reactor state-variables. For instance, in a bi-enzymatic reactor case, [21,22,23,35,86] used a HSMDM to derive optimal operating policies of a batch reactor (BR), a series of BR-s of equal size but of different (adjustable) initial load (SeqBR), or of a FBR by accounting for multiple competing optimization objectives.
(vii).- Complex HSMDM-s can also be used for bioinformatics studies, by evaluating the influence of the bioreactor operating conditions (that is the control macro-variables) on the dynamics of cell nano-scale key-intermediates and fluxes involved in the synthesis of the metabolite of interest. For instance, in the above case study (i-b) , the HSMDM including the reaction modules of glycolysis, ATP-recovery system, TRP-operon expression, and biomass [X] production allows a quick evaluation of the E.coli cell metabolic fluxes (related to the metabolic engineering topics of [27] ), thus directing the design of GMO cells with desirable properties (‘motifs’) [10,24].
(viii).- Extended HSMDM can successfully be used to obtain lumped dynamic models of the bioreactor useful for rapid engineering calculations, by employing specific model reduction rules and additional kinetic data valid in the local operating domains (in the control variable space). Specific kinetic model consistent reduction rules are given by [48,49,87] for nonlinear models, or [88,89] for linear models cases. As a result of such an approach, the bioprocess complexity may be described by a succession of local reduced models enfolded on the real process. The locally reduced models include only the key metabolic pathways to obtain the relevant process state predictions (of interest).
(ix).- As a corollary to the issue (vii), The structured HSMDM [cell-and-bioreactor] models are also useful for understanding the cellular bioprocess in direct connection to the bioreactor operating mode. For instance, in the above case study (i-b), such an extended cell model can in-silico determine the conditions of occurrence of oscillations for the cell glycolysis [44,47,80,81,83], or oscillations in the TRP-operon expression [47,61,], or those leading to a balanced cell growth (quasi-steady-state QSS, conditions, i.e. the cell homeostasis) [80]. As another example, in the above case study (i-a), the extended HSMDM can predict the mer-enzymes expression levels and the cytosolic mercury reduction rate depending on the mer-plasmids level [Gmer] in the cloned E. coli cells.
(x).- Some other case studies supporting the use of complex HSMDM are mentioned in the parts 3&4 of this work for various purposes. For instance,(a) to in-silico design GMO E. coli strains for maximizing the production of both succinate (SUCC) and of biomass in a batch reactor (BR) by using the gene-knockout and the Pareto front techniques [31]; (b) to off-line optimize the operating policy of a FBR to maximize the monoclonal antibodies (mAbs) production [2,90], etc.
Application of CBE and NSCT concepts/principles and rules to construct and HSMDM dynamic models
Basically, by reviewing the rules to construct MSDKM and HSMDM models, and their applications, this work is aiming to prove the feasibility and the advantages of using the classical and novel concepts and numerical tools of the chemical and biochemical engineering (CBE), and of the control theory of nonlinear systems (NSCT) to develop extended cell dynamic math models with including the key-reaction modules of the cell CCM, and of GRC-s involved in the regulation of the metabolites’ syntheses of practical interest.
While the 1-st part of this work is aiming at reviewing the general CBE concepts used for such an approach, the 2-nd part of the work will briefly reviews the ’whole-cell of variable-volume’ (WCVV) modelling framework introduced and promoted by the author in previous works, inspired by the CBE applied rules in the case of nonlinear reacting systems of variable volume. Also, this 2-nd part reviews and points-out the features of the deterministic WCVV models, and its advantages when simulating GERM-s, and GRC-s dynamics in living cells, by contrast to the classical (default) WCCV (whole-cell constant-volume modelling framework). The same review includes a short description of the regulatory performance indices (P.I.-s) of GERM-s (inspired from the NSCT), and also rules to link GERM-s when modelling GRC-s, as well as other related theoretical aspects necessary to construct MSDKM and HSMDM models. Special attention is paid to the conceptual and numerical rules used to construct various individual GERM-s kinetic models, but also various GRC-s (e.g. toggle-switch, amplitude filters, operon expression, etc.).
To apply the CBE and NSCT principles and rules, as documented by [11,12,14,32], to overcome the cell process dynamics complexity, the metabolic pathway representation with continuous and/or stochastic individual or lumped reactions/variables remains the most adequate and preferred representation of cell processes, the adaptable-size and structure of the lumped model (species and/or reactions) depending on available information and the utilisation scope.
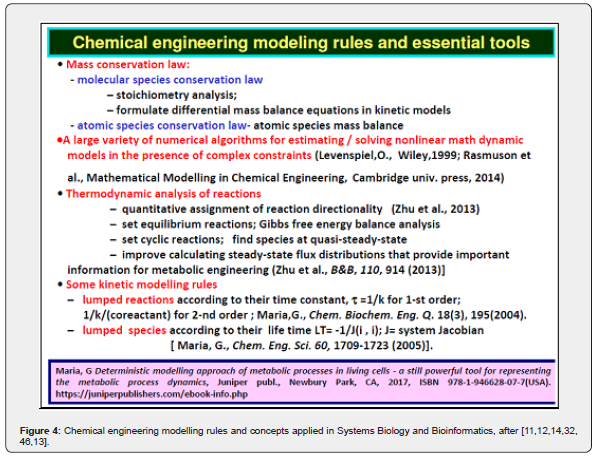

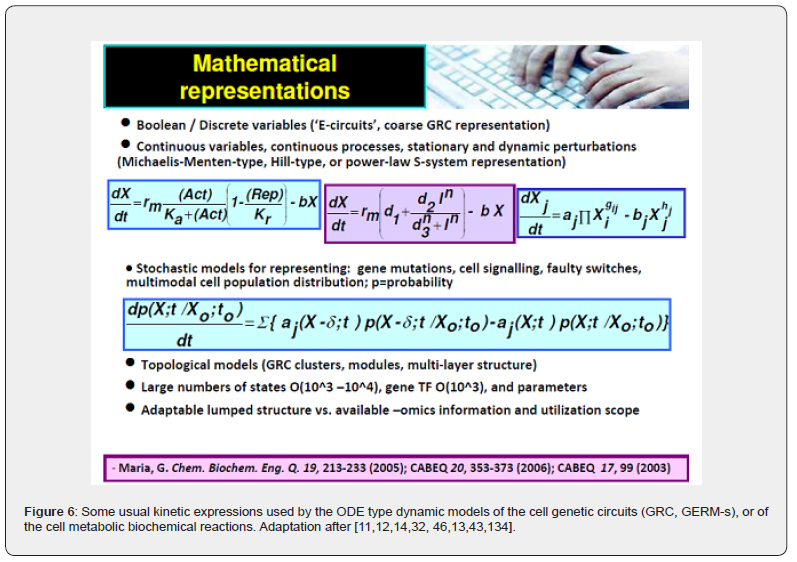
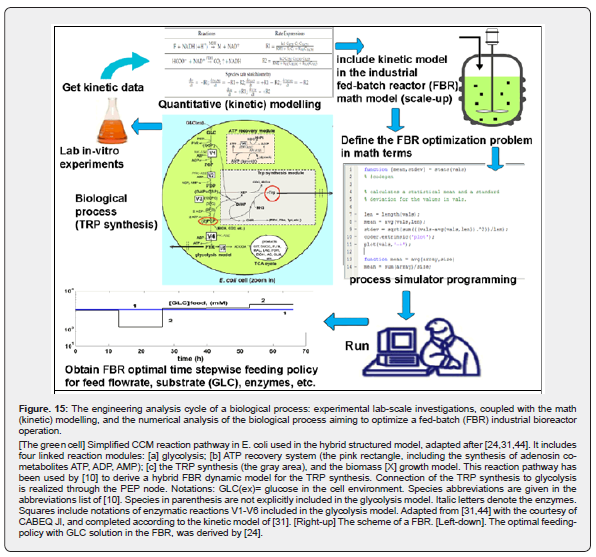
The main advantages of deterministic / continuous variable kinetic models are coming from the use of experience, concepts, math representation, rules, and algorithms of the CBE (Figure 4 and Figure 5) [11,12,14,32, 46,48,13]. The reaction rate expressions in the deterministic models of continuous variables are the usual ones of CBE that is of (extended) Michaelis-Menten, or Hill type (see Figure 6). Based on these CBE principles and rules, a large number of CCM kinetic models have been reported in the literature, such as those of [26,44,64,65,91]. A short discussion is given by [10,44]. Such a CCM-based kinetic model (Figure 2) was used by [10,24] to optimize an experimental pilot-scale FBR. In fact, the glycolysis together with the phosphotransferase (PTS)-system, or an equivalent one for GLC(glucose)-uptake, and with the pentose-phosphate pathway (PPP), and with the tricarboxylic acid cycle (TCA), all these are part of the so-called “central carbon metabolism” (CCM)(Figure 2 and Figure 3) [31].
Even if complicated and, often over-parameterized, the continuous variable dynamic deterministic ODE models (eq. 1A-B, eq. 3A-B, eq. 4A-B, eq.5-6, eq. 8 from Part-2) of the CCM metabolic pathways, or of GRC-s present a significant number of advantages, being able to reproduce in detail molecular interactions, the cell slow or fast continuous response to exo/ando-geneous continuous perturbations [40,43]. Besides, the use of ODE kinetic models presents the advantage of being computationally tractable, flexible, easily expandable, and suitable to be characterized using the tools of the nonlinear system theory [92], by accounting for the regulatory system properties, that is: dynamics, feedback / feedforward, and optimality.
And, most important, such ODE kinetic modelling approach allows using the strong tools of the classical (bio-)chemical engineering (BCE) modelling concepts summarized in (Figure 4 and Figure 5). The most important ones are the followings [11,12,14,32, 46,13,82]:
a. Fulfillment of the molecular, and elementary species (atoms types) conservation law (species differential mass balance set) [12,14,32].
b. Fulfillment of the atomic species conservation law ( atomic species mass balance).
c. The thermodynamic analysis of reactions (that is quantitative assignment of reaction directionality) [93]
d. Set equilibrium reactions by using Gibbs free energy balance analysis; set cyclic reactions; find species at quasi-steady-state to replace its differential mass balance with an algebraic equation [94]
e. Extended HSMDM–s allow improved evaluation of steady-state flux distributions (i.e. stationary metabolic reaction rates) that provide important information for metabolic engineering [94,14,32].
f. Allow application of ODE model species and/or reaction lumping rules [48,49].
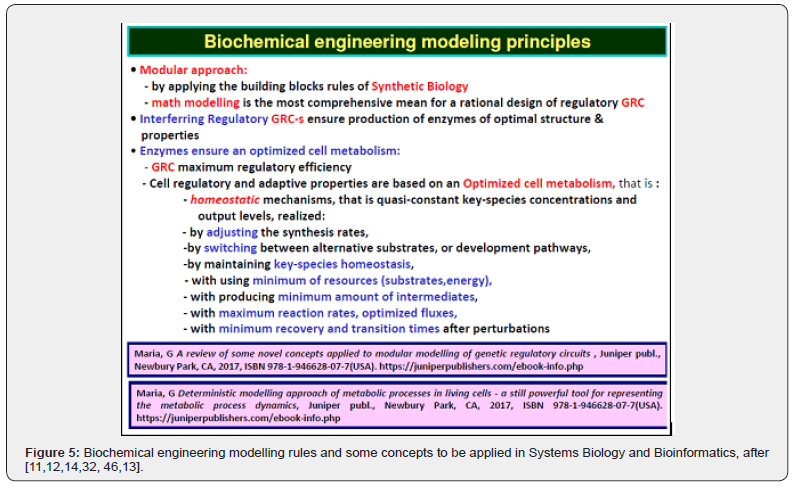

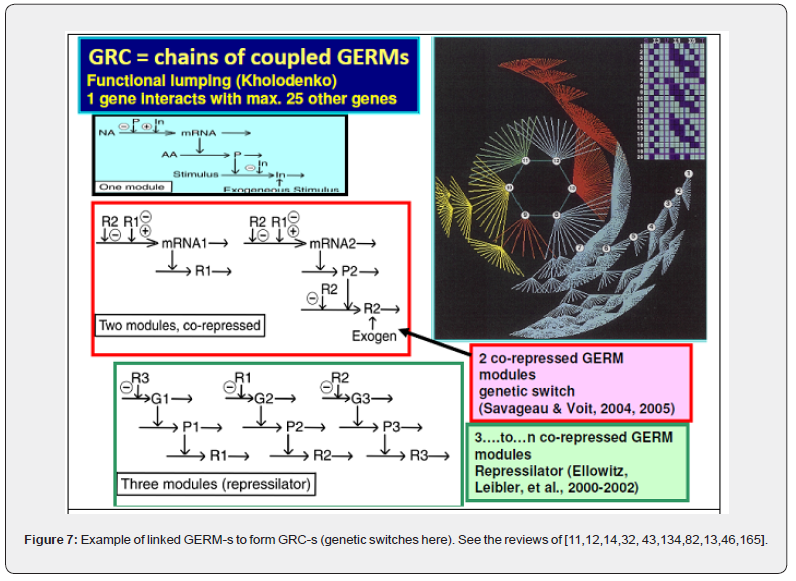
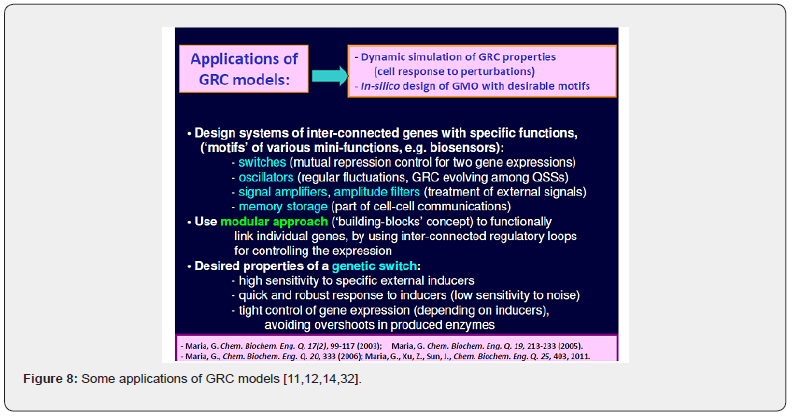
When developing deterministic models for the CCM, or for other cell metabolic processes of a micro-organism, to be further used for GMO design, an important aspect is to also include math (kinetic) models of individual GERM-s characterizing the gene expression control of the enzymes production. Also, by linking the interfering GERM modules, complex GRC regulatory chains can thus be obtained [11,12,14,32,95]. Such GRC-s refers to genetic switches (Figure 7), or operon expression (Figure 8), genetic amplifiers, etc. [11,12,14,32, 43,96,97,98,99].

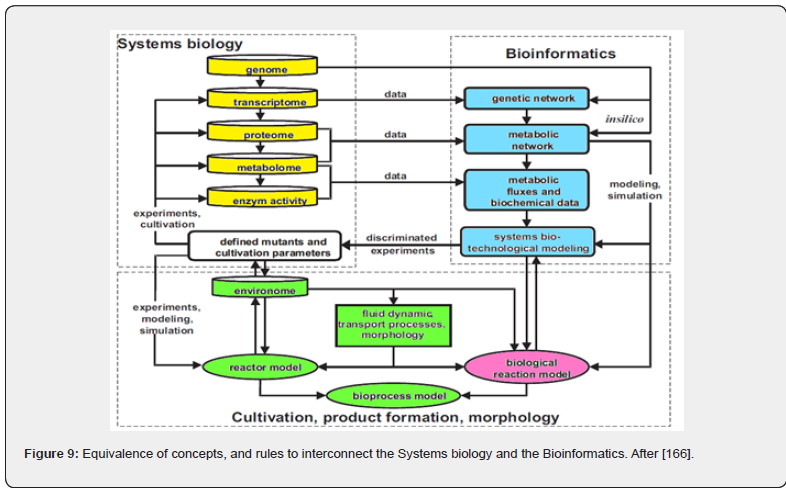
Development of ODE dynamic models to adequately reproduce such complex synthesis related to the CCM [26,64,91], but also to the GRC-s tightly controlling such metabolic processes reported significant progresses over the last decades in spite of the lack of structured experimental kinetic information, being rather based on sparse information from various sources and unconventional identification / lumping algorithms [11,12,14,32, 43,48,49,100]. However, such structured models are extremely useful for in-silico design of novel GRC-s conferring new properties/functions to the mutant cells, in response to external stimuli. This emergent field belongs to the Synthetic Biology (see below) [31,101-112]. This topics belongs to the so-called ’computational systems biology’, or simply ’bioinformatics’. In fact, the two emergent research/applicative fields are closely inter-connected, as depicted in (Figure 9), and strongly related to the CBE, NSCT, and Systems Biology principles and rules.
Systems Biology
Systems Biology is defined as “the science of discovering, modelling, understanding and ultimately engineering at the molecular level the dynamic relationships between the biological molecules that define living organisms” (Leroy Hood, Head Inst. Systems Biology, Seattle, USA). Systems Biology is one of the modern tools, which uses advanced mathematical simulation models for in-silico design of GMOs that possess specific and desired functions and characteristics. The works of [11,12,14,32, 43,48] presented short reviews about Systems Biology , by including a short history, their modern concepts, and their principles, and math/numerical-experimental tools and modelling rules, derived from those of the CBE and NSCT to construct deterministic models used by the Systems Biology to numerically simulate the dynamics of cellular metabolic processes. This involves application of the classical CBE modelling techniques (mass balance, thermodynamic principles, see the above (C1-C6) principles), and algorithmic rules, and also the NSCT principles/rules (see in the part 2 of this work, the definitions of the P.I.-s used to characterize the individual GERM–s and GRC–s self-regulatory efficiency inside the cell), and of the bioinformatics rules, briefly presented in (Figure 4, and Figure 5), and in the above (C1-C6) principles. The main principles, concepts, and rules of the CBE are also shortly reviewed by [113-117]. The metabolic pathway representation with continuous and/or stochastic variables remains the most adequate and preferred representation of the cell processes, the adaptable-size and structure of the lumped model depending on available information and the utilisation scope [11,12,14,32, 43,48,49].
The in-silico re-design of the cell metabolism is an up-to-date subject in Synthetic Biology. But in this effort, Synthetic Biology is closely assisted by the Systems Biology focus on the cell organization, the former being one of the main tools in the in-silico design of genetically modified micro-organisms (GMO) with desired characteristics, and with applications in medicine, such as therapy of diseases (gene therapy), production of new devices based on cell-cell communicators, biosensors, production of vaccines, etc. The Systems Biology also aims at understanding the dynamic interaction between components of a living system or between living systems. (http://www.erasysbio.net/). To realize these ambitious objectives, Systems Biology uses a wide range of tools, but mainly complex mathematical simulation models and numerical rules imported from the CBE, NSCT , and Bioinformatics linked to the bio–omics databanks (see below).
When developing extended structured HSMDM models, besides CBE, NSCT, and Systems Biology principles and rules (Figure 4, Figure 5, Figure 9), the Bioinformatics concepts and rules play an essential role because they make the connection with the bio-omics databanks [see the below databanks and cell simulation platforms (a-e) ], from which the most important information refers to the genome map and its correspondence to the proteome map for a certain micro-organism.
The main objective of the ’computational systems biology’ is to model the kinetics of entire living cells at the molecular level on a mechanistic base. Given the enormous complexity and unknown aspects of such systems, formulating reliable models with predictive ability remains only a dream. However, advances in genomics, transcriptomics, proteomics, metabolomics, and in the computing power provide hope that this objective might be realized within next couple of decades. Bioinformatic databases and software platforms are being constructed for modeling entire cells with massive amounts of data [72,73,103,118]. For example:

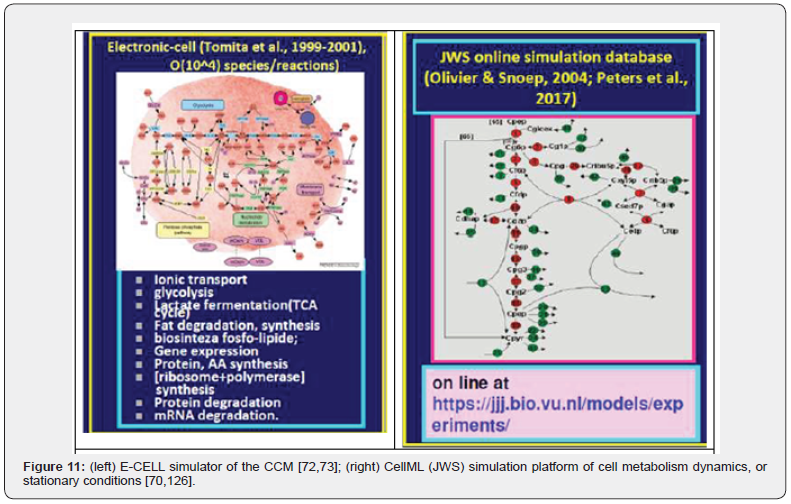
a) E-Cell software allows simulating reaction pathways within compartment-based cells using the continuous-differential modeling approach [72,73,119,120] (Figure 11-left). The computing platform objects are the followings: [compartments, compounds, genes, reactions)]. The E-cell cell simulation platform [72,73] has been used in conjunction with the EcoCyc [121], and KEGG [122,123] databases to simulate the dynamics of 127 genes/protein found in genitalium.
b) V-Cell [124,125]. The computing platform objects are the followings: [model and geometry attached to each application, and the biological interface (real images taken with the Electron Microscope)].
c) CellML, JWS (Silicon-cell) [70,126-129] is a cell modelling and simulation framework (Figure 11-right) with compartments and membranes, each of which may include species, reactions and membrane fluxes.
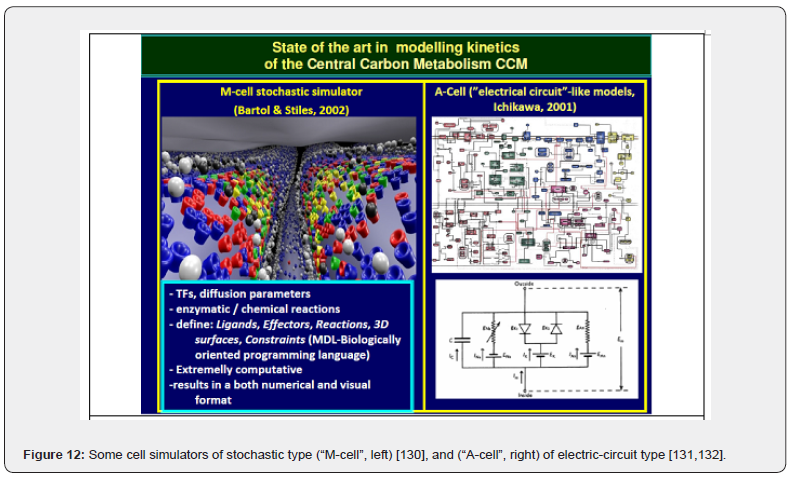
d) M-Cell simulation platform of [130] (Figure 12-left) allows simulating high-level complex cell sub-systems, such as beural communication networks, together with proteins and enzymes involved in exo-/endo-cytosis, synaptic transmission, transport and signal reception. The M-Cell simulator includes simulation of Brownian random walk, and Monte Carlo stochastic algorithms for modeling small numbers of diffusing ligands interacting with individual 3D binding sites in spatially complex environments.
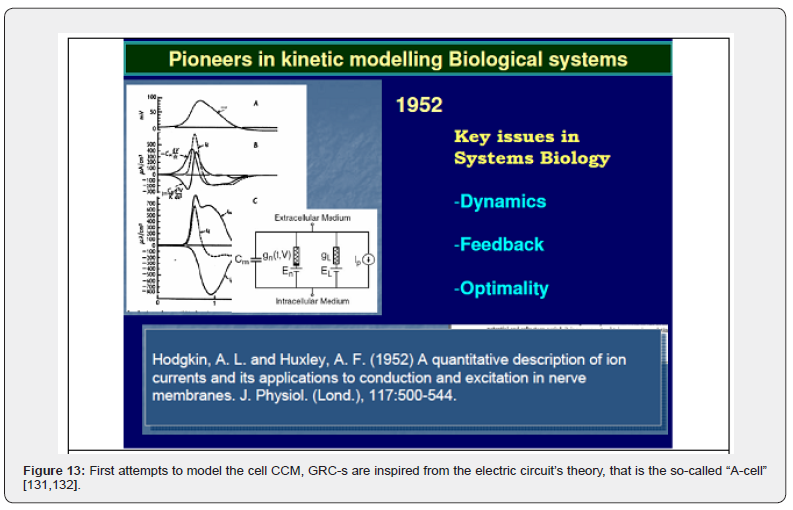
e) The A-Cell platform [131,132] (Figure 12-right; Figure 13) uses (‚electrical circuit‘ like models) to simulate biochemical reaction schemes, neurons connections, and pathways. For other bio-modelling software packages the reader is referred to the reviews of [118,131,132,133.
To easier realize the numerical simulations of such complex cell math models, specific programming languages (SBML, [118,133]), or on-line simulation platforms (JWS, [70,126]) have been developed. By using such modern computing tools, simulation of various biological systems was possible, such as [11,12,14,32, 13,43,46,134]:
• Single cell growth (e.g. Escherichia coli, Haemophilus influenzae, Mycoplasma genitalium, yeast, etc.
• Model metabolic oscillations (red-blood-cell synthesis, glycolysis, TCA (tricarboxylic acid cycle) cycle, oxidative phosphorylation, key species oscillations, etc.) [44,61,47,80,81,83,104,135]. Conditions for oscillations occurrence for various cell processes are given by [44,47,61,81,175].
• Metabolic control of protein synthesis regulation (GERM-s, GRC-s) [11,12,14,32, 43,134, 136-140].
• Modelling the central carbon metabolism (CCM) [45,64,91]. Dynamic models of some CCM-core modules and applications are given by [10,26,31,44,61] (Figure 2, Figure 3, Figure 11-left).
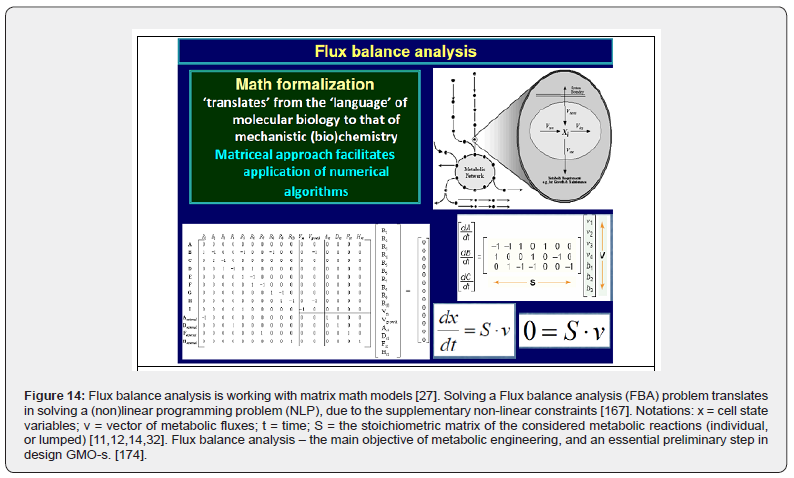
• FBA (flux balance analysis of the cell metabolism) used to design of GMO-s [95] (Figure 14). Metabolic fluxes (v(j)) are defined as the enzymatic reaction rates at the quasi-stationary-cell-conditions (QSS), that is at the cell balanced growth (homeostasis). Determination of metabolic fluxes (by using the metabolic engineering tools [27], or the dynamic HSMDM models) allowed to in-silico derive GMO-s of desired characteristics by using the so-called “gene knock-out” technique [174]. By removing a certain enzyme (and its attached encoding gene), the catalyzed reaction is also removed from the CCM. Similarly, the target reaction (flux) can be amplified / diminished by increasing / decreasing the corresponding enzyme (biocatalyst) concentration, and, correspondingly by increasing / decreasing the encoding gene plasmid concentration in the analyzed micro-organism genome. In other words, FBA, and of the metabolic engineering are the essential preliminary steps in design GMO-s. [237].
• Modelling the cell cycle [141,142].
• Modelling the drug release and cell-drug interactions [143,144].
• Modelling cellular communications, neuronal transmission
• Analysis of ‘logical essence’ of life (life minimal requirements)
At the same time, the exponential-like increase of the experimental biological information lead to development of valuable bio-omics databanks, such as those described by [11,12,14,32]:
-.- KEGG [122,123];
-.- JWS [70,126];
-.- EcoCyc [121];
-.- Roche [145], etc.
However, it is only over the last decades when Systems Biology reported notable successes due to a considerable increase in computing power of the modern computers. It is to mention here, for instance, the cell simulator platforms, and online model repository JWS of [70,126], or those developed by Rocha et al. [42], or by Tomita et al. [72,73], together with continuous expansion of the above referred bio-omics databases JWS, KEGG, EcoCyc, Roche, etc., and reported advances in the numerical algorithms used by bioinformatics, CBE, and NSCT [11,12,14,32].
Due to such favourable premises, related to the expansion of bio-omics databanks, and cell metabolism (CCM, GRC-s) dynamic models, novel works have been reported over the last decades. Among the milestone works in Systems Biology it is to mention the contributions in modelling / design of GRC-s, GERM-s, FBA, MCA of [11,12,14,32, 92,146-149]. The number of published papers in the Systems Biology area increases with two orders of magnitude from 2000 to 2007, and it is still exponentially increasing, most of them was being founded by programs of the European Science Foundation.
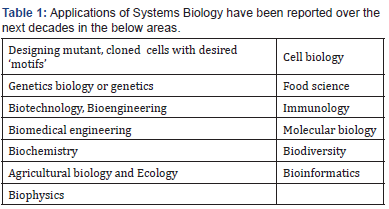
As stated by [150,151], tremendous applications of Systems Biology have been reported over the next decades in the below areas (see also [11,12,14,32]) (Table 1):
Bioinformatics
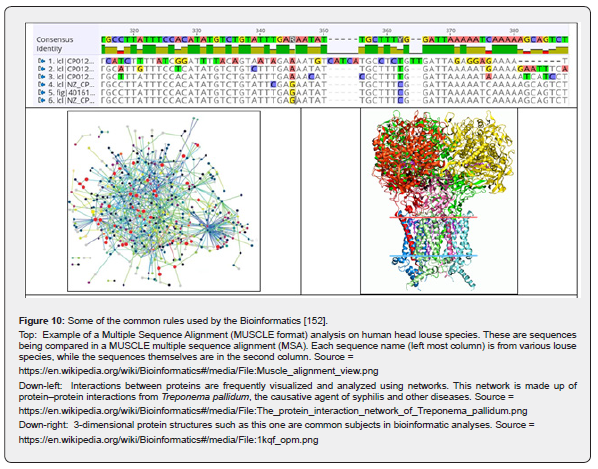
According to [152], the bioinformatics is an inter-disciplinary field that develops methods and software tools for understanding and interpret the biological data, in particular when the data sets are large and complex. As an interdisciplinary field of science, the bioinformatics combines several classic/modern disciplines, such as: biology, chemistry, physics, computer science, information engineering, mathematics, and statistics to analyze and interpret the biological data. Bioinformatics has been used rather for in-silico analyses of biological queries using a large number of computational and statistical techniques, aiming to design novel GMO-s of practical (industrial) use [41,153-162]. The most important rule of the Bioinformatics refers to the rapid, and computer-assisted genome sequentiation [152]. Some of the common rules used by the bioinformatics are given in (Figure 10) [152].
Thus, Bioinformatics includes biological studies that use computer programming as part of their methodology/rules, as well as specific analysis ‘pipelines’ that are repeatedly used, particularly in the field of genomics. Common uses of bioinformatics include identification of candidates genes and “single nucleotide polymorphisms” (SNP-s). Often, such identification is made with the aim to better understand the genetic basis of disease, unique adaptations, desirable properties (esp. in agricultural species), or differences between populations. In a less formal way, bioinformatics also tries to understand the organizational principles within nucleic acid and protein sequences, called proteomics [152].
And, “in a context of increasing calls for biology to be predictive, modelling and optimization tools of Bioinformatics (most of them imported from CBE, and NSCT ) are the only approaches biology has for making satisfactory predictions” [163]. Due to the computing facilities offered by the algorithmic rules developed by the (bio)chemical engineering and nonlinear systems biology rules (Figure 4, Figure 5, Figure 6, Figure 15), the developed cell math models use a vectorial-matriceal approach (Figure 16), with a continuous model upgrading based on dynamic experimental data recorded in a chemostat (i.e. a continuously operated bioreactor), operated under steady-state, or in a dynamic regime following an input perturbation in the substrate/enzymes/biomass concentration in the solution fed in the bioreactor [64,95].
Given these developments, as well as the seemingly inexorable advances in computing power, it is tempting to believe that reliable whole-cell models (WC) with predictive power will be forthcoming once complete sets of ’bio-omic’ information become available. However, a better theoretical understanding of cellular life, viewed holistically, may be required before we can understand how life emerges out of complex networks of molecular-level interactions between cellular components. This study represents a foyer into whole-cell kinetic modelling, in an attempt to understand holistic aspects of cell-system from a quantitative computational perspective.
Related to the WC math modelling aspect, a concerned section in the Part-2 of this work is aiming at examining some fundamental properties of living cells relevant to WC holistic math-modelling. Thus, a methodology to build-up regulatory kinetic schemes in a ’whole cell variable cell volume’ (WCVV) modelling framework, but also under stationary/perturbed environmental conditions was proposed, developed, and promoted by Maria [11,12,14,32, 17,18,43,99,100] for a generic protein/gene (P/G) pair synthesis process, its GERM regulatory module, but also for cell GRC-s, and for the whole CCM dynamic model construction. The model elements of novelty consist in accounting variable cell-volume, and constant osmotic pressure conditions. Thus, most of the drawbacks of classical (default) continuous-concentration simulators (developed for a constant volume system, WCCV) are removed. Exemplification is made for several relatively simple GRC-s from literature [11,12,14,32, 43,99,100]. The chosen examples analyse the impact of the proposed WCVV modelling approach on the MSDKM model prediction better quality, such as: (a) a more realistic prediction of regulatory scheme sensitivity to both stationary and dynamic perturbed conditions; (b) better pointed-out the cell species interconnectivity (direct, or indirect through the cell volume to which all species contributes); (c) the effect of the so-called “cell-ballast” on the cell-behaviour vs. various type of perturbations; (d) a more realistic way to design interconnected GERM-s and rules to link them in GRC-s in a common volume growing environment. Other analysis aspects, not developed in this paper, but exemplified in the Part 2 and 3&4 of this work, can be easily approached under the presented WCVV modelling hypotheses (Part 2), such as the cell system state multiplicity, and characterization of their oscillatory phenomena.
References
- Liese A, Seelbach K, Wandrey C (2006) Industrial biotransformations, Wiley-VCH, Weinheim.
- Maria G (2020A) Model-based optimization of a fed-batch bioreactor for mab production using a hybridoma cell culture, Molecules 25(23): 5648-5674.
- Scoban AG, Maria G (2016) Model-based optimization of the feeding policy of a fluidized bed bioreactor for mercury uptake by immobilized P. putida cells. Asia-Pacific J Chem Engineer 11(5): 721-734.
- Ghose TK, Fiechter A, Blakebrough N (1977-1978) Advances in Biochemical Engineering, Springer Verlag, Berlin, p. 7-10.
- Straathof AJJ, Adlercreutz P (2005) Applied biocatalysis, Harwood Academic Publ, Amsterdam.
- Jiménez-González C, Woodley JM (2010) Bioprocesses: modeling needs for processevaluation and sustainability assessment, Comput Chem Eng 34(7): 1009-1017.
- Khamseh AAG, Miccio M (2012) Comparison of batch: fed-batch and continuouswell-mixed reactors for enzymatic hydrolysis of orange peel wastes. Process Biochem 47(11): 1588-1594.
- Tsangaris DM, Baltzis BC (1996) Evaluation of batch and semi-batch reactor operation for enzymatic reactions with inhibitory kinetics. Chem Engineer Sci 51(11): 2757-2762.
- Maria G, Luta I (2013) Structured cell simulator coupled with a fluidized bed bioreactor model to predict the adaptive mercury uptake by E. coli cells. Comput Chem Engineer 58(11): 98-115.
- Maria G (2021) A CCM-based modular and hybrid kinetic model to simulate the tryptophan synthesis in a fed-batch bioreactor using modified E. coli cells. Comput Chem Engineer 153: 107450-107466.
- Maria G (2017A) A review of some novel concepts applied to modular modelling of genetic regulatory circuits. Juniper Publ, Irvine, CA, USA.
- Maria G (2017B) Deterministic modelling approach of metabolic processes in living cells - a still powerful tool for representing the metabolic process dynamics. Juniper Publ, Irvine, CA, USA.
- Maria G (2017C) Application of (bio) chemical engineering principles and lumping analysis in modelling the living systems, Current Trends in Biomedical Engineering & Biosciences. Juniper Publ, Irvine, CA, USA 1(4): 555566.
- Maria G (2018) In-silico design of Genetic Modified Micro-organisms (GMO) of industrial use, by using Systems Biology and (Bio)Chemical Engineering tools. Juniper Publ, Irvine, CA, USA.
- Maria G, Scoban AG (2017) Setting some milestones when modelling gene expression regulatory circuits under variable-volume whole-cell modelling framework. 1. Generalities, Revista de Chimie (Bucharest) 68(12): 3027-3037.
- Maria G, Scoban AG, (2018) Setting some milestones when modelling gene expression regulatory circuits under variable-volume whole-cell modelling framework. 2. Case studies, Revista de Chimie (Bucharest) 69(1): 259-266.
- Maria G, Maria C, Tociu C (2017) Comments on two novel review ebooks in deterministic modelling of metabolic processes and of genetic regulatory circuits in living cells. UPB Bull Sci 79(3): 3-19.
- Maria G, Gijiu CL, Maria C, Tociu C, Mihalachi M (2018) Importance of considering the isotonic system hypothesis when modelling the self-control of gene expression regulatory modules in living cells. Curr Trend Biomed Engineer Bio Sci 12(2): 555833.
- Maria G (2012) Enzymatic reactor selection and derivation of the optimal operation policy, by using a model-based modular simulation platform. Comput Chem Eng 36: 325-341.
- Maria G, Crisan M (2017) Operation of a mechanically agitated semi-continuous multi-enzymatic reactor by using the Pareto-optimal multiple front method. J Process Control 53: 95-105.
- Maria G (2020B) Model-based optimization of a batch reactor with a coupled bi-enzymatic process for mannitol production. Comput Chem Engineer 133: 106628-106635.
- Maria G, Peptănaru IM (2021) Model-based optimization of mannitol production by using a sequence of batch reactors for a coupled bi-enzymatic process - A dynamic approach. Dynamics-Basel 1: 134-154.
- Maria G, Renea L, Maria C (2022) Multi-objective optimization of the fed-batch bi-enzymatic reactor for mannitol production. Dynamics-MDPI 2: 270-294.
- Maria G, Renea L (2021) Tryptophan production maximization in a fed-batch bioreactor with modified E. coli cells, by optimizing its operating policy based on an extended structured cell kinetic model. MDPI Bioengineering-Basel 8(12): 210-247.
- DiBiasio D (1989) Introduction to the control of biological reactors. In: Shuler ML (ed.), Chemical engineering problems in biotechnology. Am Institute of Chem Engineer New York, pp. 351-391.
- Miskovic L, Tokic M, Fengos G, Hatzimanikatis V (2015) Rites of passage: requirements and standards for building kinetic models of metabolic phenotypes. Curr Opinion Biotech 36: 146-153.
- Stephanopoulos GN, Aristidou AA, Nielsen J (1998) Metabolic Engineering. Principles and Methodologies. Academic Press, San Diego, CA, 1998.
- Maranas C, Hatzimanikatis V (2005) Preface - Cell modelling. Comp Chem Eng 29: 405-406.
- Feist AM, Henry CS, Reed JL, Krummenacker M, Joyce AR, et al. (2007) A genome-scale metabolic reconstruction for Escherichia coli K-12 MG1655 that accounts for 1260 ORFs and thermodynamic information. Mol Sys Biol 3(121): 1-18.
- Hatzimanikatis V, Floudas CA, Bailey JE (1996) Analysis and design of metabolic reaction networks via Mixed-Integer Linear Optimization. AIChE J 42(5): 1277-1292.
- Maria G, Xu Z, Sun J (2011) Multi-objective MINLP optimization used to identify theoretical gene knockout strategies for E. coli cells. Chem Biochem Eng Quart 25: 403-424.
- Maria G (2023) Hybrid modular kinetic models linking cell-scale structured CCM reaction pathways to bioreactor macro-scale state variables. Applications for solving bioengineering problems, Juniper Publ Irvine, CA, USA.
- Bonvin D (1998) Optimal operation of batch reactors-a personal view. J Process Control 8: 355-368.
- Smets IY, Claes JE, November EJ, Bastin GP, Van Impe JF (2004) Optimal adaptive control of (bio)chemical reactors: past, present and future. J Process Control 14(7): 795-805.
- Maria G, Renea L, Peptănaru IM, Maria C (2022B) In-silico optimization of a bi-enzymatic batch reactor for mannitol.
- Srinivasan B, Bonvin D, Visser E, Palanki S (2003) Dynamic optimization of batch processes: II. Role of measurements in handling uncertainty, Comp Chem Eng 27(1): 27-44.
- Martinez E (2005) Batch-to-batch optimization of batch processes using the Stat simplex search method, Proc. 2nd Mercosur Congress on Chemical Engineering. Rio de Janeiro, Costa Verde, Brasil 20: 14-18.
- Rao M, Qiu H (1993) Process control engineering: A textbook for chemical, mechanical and electrical engineers. Gordon and Breach Science Publ, Amsterdam, 1993.
- Hartig F, Keil FJ, Luus R (1995) Comparison of optimization methods for a fed-batch reactor. Hung J Ind Chem 23: 81-160.
- Styczynski MP, Stephanopoulos G (2005) Overview of computational methods for the inference of gene regulatory networks, Comp Chem Eng 29(3): 519-534.
- Xiong J (2006) Essential bioinformatics, Cambridge University Press, Cambridge UK.
- Rocha I, Maia P, Evangelista P, Vilaça P, Soares S, et al. (2010) OptFlux: an open-source software platform for in silico metabolic engineering. BMC Syst Biol 4(45).
- Maria G (2005B) Modular-Based Modelling of Protein Synthesis Regulation. Chem Biochem Eng Quarter 19: 213-233.
- Maria G (2014A) Insilico derivation of a reduced kinetic model for stationary or oscillating glycolysis in Escherichia coli bacterium. Chem Biochem Eng Quarter 28(4): 509-529.
- Visser D, Schmid JW, Mauch K, Reuss M, Heijnen JJ (2004) Optimal re-design of primary metabolism in Escherichia coli using linlog kinetics. Metab Eng 6(4): 378-390.
- Maria G, Maria C, Renea L (2022C) Application of (bio)chemical engineering concepts and rules in Bioinformatics. Review of a CCM-based modular and hybrid kinetic model used to simulate and optimize a bioreactor with genetically modified cells. Curr Trends Biomed Eng Biosci. Juniper publ, Irvine CA, USA 20(3): 556039.
- Maria G, Gijiu CL, Maria C, Tociu C (2018A) Interference of the oscillating glycolysis with the oscillating tryptophan synthesis in the E. coli cells. Comp Chem Eng 108: 395-407.
- Maria G (2004) A review of algorithms and trends in kinetic model identification for chemical and biochemical systems. Chem Biochem Eng Quarter 18: 195-222.
- Maria G (2019) Numerical methods to reduce the kinetic models of (bio)chemical processes. Printech Publ, Bucharest, 2019 pp. 815.
- Wu WH, Wang FS, Chang MS (2011) multi-objective optimization of enzyme manipulations in metabolic networks considering resilience effects. BMC Sys Bio 5: 145.
- Maria G, Crisan M (2014) Evaluation of optimal operation alternatives of reactors used for D-glucose oxidation in a bi-enzymatic system with complex deactivation kinetics. Asia-Pacific Journal of Chemical Engineering 10 (1): 22-44.
- Crisan M, Maria G (2016) Modular simulation to check performances of various reactors for the enzymatic D-glucose oxidation. Revue Roumaine de Chimie 61(6-7): 549-556.
- Reuss M (1986) Computer control of bioreactors present limits and challenges for the future. In: Morari M, McAvoy TJ (Eds.), Proc 3rd Intl Confernce on Chemical Process Control - CPCIII. Asilomar (USA), Elsevier, Amsterdam, Netherlands.
- Banga JR, Alonso AA, Singh PR (1994) Stochastic optimal control of fed-batch bioreactors. AIChE Annual Meeting, San Francisco.
- Doran PM (1995) Bioprocess engineering principles. Elsevier Amsterdam.
- Sarkar D, Modak JM (2005) Pareto-optimal solutions for multi-objective optimization of fed-batch bioreactors using nondominated sorting genetic algorithm. Chem Eng Sci 60(2): 481-492.
- Henson MA, (2010) Model-based control of biochemical reactors. In: Levine W (Ed.), The control handbook, Taylor and Francis (2nd edition) New York.
- Henson MA, Muller D, Reuss M (2004) Combined metabolic and cell population modelling for yeast bioreactor control. In: Allgöwer F (Ed.), Proc IFAC Symposium on Advanced Control of Chemical Processes. Hong Kong.
- Bodizs L, Titica M, Faria N, Srinivasan B, Dochain D, et al. (2007) Oxygen control for an industrial pilot-scale fed-batch filamentous fungal fermentation. Journal of Process Control 17(7): 595-606.
- Ashoori A, Moshiri B, Khaki-Sedigh A, Bakhtiari MR, (2009) Optimal control of a nonlinear fed-batch fermentation process using model predictive approach. Journal of Process Control 19(7): 1162–1173.
- Maria G (2020C) In-silico determination of some conditions leading to glycolytic oscillations and their interference with some other processes in E. coli cells. Frontiers in Chemistry 8: 526679-526693.
- Roeva O, Pencheva T, Tzonkov S, Arndt M, Hitzmann B, et al. (2007) Multiple model approach to modelling of Escherichia coli fed-batch cultivation extracellular production of bacterial phytase. Journal of Biotechnology 10 (4): 592-603.
- Edwards JS, Palsson BO (2000) The Escherichia coli MG1655 in silico metabolic genotype: its definition, characteristics, and capabilities. Proc Natl Acad Sci USA 97(10): 5528-5533.
- Chassagnole C, Noisommit Rizzi N, Schmid JW, Mauch K, Reuss M et al. (2002) Dynamic modeling of the central carbon metabolism of Escherichia coli. Biotechnology and Bioengineering 79(1): 53-73.
- Usuda Y, Nishio Y, Iwatani S, Van Dien SJ, Imaizumi A et al. (2010) Dynamic modeling of Escherichia coli metabolic and regulatory systems for amino-acid production. Journal of Biotechnology147 (1): 17-30.
- Kadir TAA, Mannan AA, Kierzek AM, McFadden J, Shimizu K et al. (2010) Modeling and simulation of the main metabolism in Escherichia coli and its several single-gene knockout mutants with experimental verification. Microbial Cell Factories 19: 88.
- Ceric S, Kurtanjek Z (2006) Model identification, parameter estimation, and dynamic flux analysis of E. coli central metabolism. Chem Biochem Eng Q 20(3): 243-253.
- Tusek AJ, Kurtanjek Z (2009) Model and global sensitivity analysis of E. coli central metabolism. In: Troch I Breitenecker F. (Eds.) Proc 6th Vienna Conference on Mathematical Modelling MATHMOD Beč, Austria.
- Teusink B, Passarge J, Reijenga CA, Esgalhado E, van der Weijden CC et al. (2000) Can yeast glycolysis be understood in terms of in vitro kinetics of the constituent enzymes? Testing biochemistry. Eur J Biochem 267 (17): 5313-5329.
- Olivier BG, Snoep JL (2004) Web-based kinetic modelling using JWS Online. Bioinformatics 20 (13): 2143-2144.
- Seressiotis A, Bailey JE (1986) MPS An algorithm and database for metabolic pathways synthesis. Biotechnol Lett 8: 837-842.
- Tomita M, Hashimoto K, Takahashi K, Shimizu T, Matsuzaki Y, et al. (1999) E-Cell: Software environment for whole cell simulation. Bioinformatics 15(1): 72-84.
- Tomita M (2001) Whole-cell simulation a grand challenge of the 21st Trends in Biotechnology 19 (6): 205-210.
- Slepchenko BM, Schaff JC, Macara I, Loew LM (2003) Quantitative cell biology with the Virtual Cell, Trends in Cell Biology, 13(11): 570-576.
- Schmid JW, Mauch K, Reuss M, Gilles ED, Kremling A et al. (2004) Metabolic design based on a coupled gene expression-metabolic network model of tryptophan production in Escherichia coli. Metabolic Engineering 6(4): 6364-377.
- Costa RS, Machado D, Rocha I, Ferreira EC (2009) Large scale dynamic model reconstruction for the central carbon metabolism of Escherichia coli. In: Distributed Computing, Artificial Intelligence Bioinformatics Soft Computing, and Ambient Assisted Living Proc IWANN conf Salamanca Spain.
- Costa RS, Machado D, Rocha I, Ferreira EC (2010) Hybrid dynamic modeling of Escherichia coli central metabolic network combining Michaelis–Menten and approximate kinetic equations. BioSystems 100(2): 150-157.
- Machado D, Zhuang KH, Sonnenschein N, Herrgård MJ (2015) Current challenges in modeling cellular metabolism. Frontiers in Bioengineering and Biotechnology 26(3): 193.
- Kiparissides A, Koutinas M, Kontoravdi C, Mantalaris A, Pistikopoulos EN et al. (2011) Closing the loop in biological systems modeling -From the in silico to the in vitro. Automatica 47(6): 1147-1155.
- Maria G, Mihalachi M, Gijiu CL (2018C) In silico optimization of a bioreactor with an E. coli culture for tryptophan production by using a structured model coupling the oscillating glycolysis and tryptophan synthesis. Chemical Eng Res and Design 135: 207-221.
- Maria G, Mihalachi M, Gijiu CL (2018B) Model-based identification of some conditions leading to glycolytic oscillations in E. coli cells. Chemical and Biochemical Engineering Quarterly 32(4): 523-533.
- Maria G, Gijiu CL, Maria C, Tociu C, Mihalachi, M (2018D) Importance of considering the isotonic system hypothesis when modelling the self-control of gene expression regulatory modules in living cells. Current Trends in Biomedical Engineering & Biosciences 12(2): 29-48.
- Maria G, Mihalachi M, Gijiu CL (2018E) Chemical engineering tools applied to simulate some conditions producing glycolytic oscillations in E. coli cells. UPB Sci Bull Series B Chemie 80(2): 27-38.
- Maria G (2009B) A whole-cell model to simulate the mercuric ion reduction by coli under stationary and perturbed conditions. Chemical and Biochemical Engineering Quarterly 23 (3): 323-341.
- Maria G (2010) A dynamic model to simulate the genetic regulatory circuit controlling the mercury ion uptake by E. coli cells. Revista de Chimie 61(2): 172-186.
- Maria G, Renea L, Maria C (2023B) Pareto optimal operating policies of a batch bi-enzymatic reactor for mannitol production. Computers and Chemical Eng 133(2): 106628.
- Edwards K, Edgar TF, Manousiouthakis VI (1998) Kinetic model reduction using genetic algorithms. Comp& Chem Eng 22: 239-246.
- Martinez EC, Beltramini LJ (1990) Lumping upon timescales: Modeling upon topological factors. Chem Eng Sci 45(8): 2103-2108.
- Maria G (2005) Relations between apparent and intrinsic kinetics of programmable drug release in human plasma, Chemical Engineering Science 60(6): 1709-1723.
- Dorka P (2007) Modelling batch and fed-batch mammalian cell cultures for optimizing MAb productivity. MS diss University of Waterloo Canada.
- Kurata H, Sugimoto Y, (2018) Improved kinetic model of Escherichia coli central carbon metabolism in batch and continuous cultures. Journal of Bioscience and Bioengineering 125(2): 251-257.
- Heinrich R, Schuster S (1996) The regulation of cellular systems New York Chapman & Hall 1996.
- Haraldsdottir HS, Thiele I, Fleming RMT (2012) Quantitative assignment of reaction directionality in a multicompartmental human metabolic reconstruction. Biophys J 102(8): 1703-1711.
- Zhu Y, Song J, Xu Z, Sun J, Zhang Y, et al. (2013) Development of thermodynamic optimum searching (TOS) to improve the prediction accuracy of flux balance analysis. Biotechnology and Bioengineering 110(3): 914-923.
- Blass LK, Weyler C, Heinzle E (2017) Network design and analysis for multi-enzyme biocatalysis. BMC Bioinformatics 18(1): 366.
- Hlavacek WS, Savageau MA (1997) Completely uncoupled and perfectly coupled gene expression in repressible systems. J Mol Biol 266(3): 538-558.
- Elowitz MB, Leibler S (2000) A synthetic oscillatory network of transcriptional regulators. Nature 403(6767): 335-338.
- Voit EO (2005) Smooth bistable S-systems. IEE Proc Syst Biol 152(4): 207-213.
- Maria G (2007) Modelling bistable genetic regulatory circuits under variable volume framework. Chemical and Biochemical Engineering Quarterly 21(4): 417-434.
- Maria G (2006) Application of lumping analysis in modelling the living systems -A trade-off between simplicity and model quality. Chemical and Biochemical Engineering Quarterly 20: 353-373.
- Heinemann M, Panke S (2006) Synthetic Biology - putting engineering into biology, Bioinformatics 22: 2790-2799.
- Salis H, Kaznessis Y (2005) Numerical simulation of stochastic gene circuits, Computers & Chemical Engineering 29: 577-588.
- Kaznessis YN (2006) Multi-scale models for gene network engineering. Chemical Engineering Science 61: 940-953.
- Atkinson MR, Savageau MA, Myers JT, Ninfa AJ (2003) Development of genetic circuitry exhibiting toggle switch or oscillatory behavior in Escherichia coli Cell 113(5): 597-607.
- Klipp E, Nordlander B, Krüger R, Gennemark P, Hohmann S (2005) Integrative model of the response of yeast to osmotic shock, Nature Biotechnology 23: 975-982.
- Klipp E (2009) Timing matters, FEBS Letters 583: 4013-4018.
- Klipp E, Liebermeister W, Wierling C, Kowald , Lehrach H, et.al (2009) Systems biology A textbook Wiley-VCH, Weinheim.
- Chen MT, Weiss R (2005) Artificial cell-cell communication in yeast Saccharomyces cerevisiae using signalling elements from Arabidopsis thaliana, Biotechnol 23: 1551-1555.
- Tian T, Burrage K (2006) Stochastic models for regulatory networks of the genetic toggle switch, Proc Natl Acad Sci USA 103(22): 8372-8377.
- Sotiropoulos V, Kaznessis YN (2007) Synthetic tetracycline-inducible regulatory networks: computer-aided design of dynamic phenotypes, BMC Syst Biol 9: 1-7.
- Tomshine J, Kaznessis YN (2006) Optimization of a stochastically simulated gene network model via simulated annealing, Biophys J 91: 3196-3205.
- Zhu R, Ribeiro AS, Salahub D, Kauffman SA (2007) Studying genetic regulatory networks at the molecular level: delayed reaction stochastic models J Theor Biol 246(4): 725-745.
- Sherlock JP (2016) Chemical engineering matters, report of the IChemE technical vice president London, UK.
- Woinaroschy A, Ofiteru DI, Lavric V (2009) Exploratory investigation of bioprocesses sustainability improvement by multicriteria-multilevel optimization, Environmental Engineering and Management Journal 8(3): 521-526.
- Woinaroschy A (2016) A paradigm-based evolution of chemical engineering, Chinese Journal of Chemical Engineering 24: 553-557.
- Gani R (2004) Computer-aided methods and tools for chemical product design, Eng Res Des 82 (A11): 1494-1504.
- Gani R, Bałdyga J, Biscans B, Brunazzi E, Charpentier JC, et al. (2020) A multi-layered view of chemical and biochemical engineering, Chemical Engineering Research and Design 155: A133-A145.
- Hucka M, Bergmann FT, Drager A, Hoops S, Keating SM, et al. (2015) Systems Biology Markup Language (SBML) Level 2 Version 5: Structures and Facilities for Model Definitions Integrative bioinformatics 12(2): 271.
- Ballet P, Zemirline A, Marce L (2001) Cellular automata, reaction-diffusion and multiagents systems for artificial cell modeling, Departement d'Informatique, Universite de Bretagne Occidentale (France).
- Kinoshita A, Nakayama Y, Tomita M (2001) In silico analysis of human erythrocyte using E-Cell system, ICSB 2001 - 2nd Int. Conf. on Systems Biology, Nov. 4-7, 2001, California Institute of Technology, Pasadena (CA).
- EcoCyc (2005) Encyclopedia of Escherichia coli K-12 genes and metabolism, SRI Intl., The Institute for Genomic Research, Univ. of California at San Diego.
- KEGG (2006) A collection of manually drawn pathway maps representing our knowledge of the molecular interaction, reaction and relation networks for a large number of cells and micro-organisms, Kanehisa Laboratories, Kyoto.
- Kanehisa M, Furumichi M, Sato Y, Ishiguro-Watanabe M, Tanabe M (2021) KEGG: integrating viruses and cellular organisms, Nucleic Acids Research 49(D1): D545-D551.
- NRCAM (2002) The National Resource for Cell Analysis and Modeling. Virtual cell modeling and simulation framework, The University of Connecticut Health Center, Richard D. Berlin Center for Cell Analysis & Modeling, Farmington (USA).
- Moraru I, Schaff JC, Slepchenko BM, Loew LM (2002) The Virtual Cell: An integrated modeling environment for experimental and computational cell biology, Annals of the New York Academy of Sciences 971: 595-596.
- Peters M, Eicher JJ, Van Niekerk DD, Waltemath D Snoep JL (2017) The JWS online simulation database. Bioinformatics 33(10): 1589-1590.
- Westerhoff HV (2006) Engineering life processes live: the Silicon cell. ESCAPE-16 Conference, Garmisch-Partenkirchen, July 9-13, 2006, Germany.
- CellML Group (2002) CellML: Related biological languages, Bioengineering research group, The University of Auckland New Zealand.
- Hedley WJ, Nelson MR, Bullivant DP, Nielson PF (2001) A short introduction to CellML, Philosophical Transactions of the Royal Society of London A 359: 1073-1089.
- Bartol TM, Stiles JR (2002) MCell: A general Monte Carlo simulator of cellular microphysiology, The Salk Institute, Computational Neurobiology Lab.
- Ichikawa K (2000) A-Cell: a platform for the construction of biochemical reaction and electrical equivalent circuit models in a biological cell and a neuron, Fuji Xerox Co. Ltd., Kanagawa (Japan).
- Ichikawa K (2001) A-Cell: graphical user interface for the construction of biochemical reaction models, Bioinformatics 17(5): 483-484.
- CRGM (2002) Centro di Ricerca in Genetica Molecolare, Istituto di Istologia ed Embriologia Generale, University of Bologna.
- Maria G (2014B) Extended repression mechanisms in modelling bistable genetic switches of adjustable characteristics within a variable cell volume modelling framework, Chemical & Biochemical Engineering Quarterly 28(1): 35-51.
- Guantes R, Poyatos F (2006) Dynamical principles of two-component genetic oscillators, PLoS Computational Biology 2(3): 188-197.
- Chen T, He HL, Church GM (1999) Modeling gene expression with differential equations, Pacific Symposium of Biocomputing Hawaii pp29-40.
- Drengstig T, Jolma IW, Ni XY, Thorsen K, Xu XM, et al. (2012) A Basic Set of Homeostatic Controller Motifs, Biophys J 103(9): 1-11.
- Komasilovs V, Pentjuss A, Elsts A, Stalidzans E (2017) Total enzyme activity constraint and homeostatic constraint impact on the optimization potential of a kinetic model. Biosystems 162: 128-134.
- Matsuura T, Tanimura N, Hosoda K, Yomo T, Shimizu (2017) Reaction dynamics analysis of a reconstituted Escherichia coli protein translation system by computational modeling. PNAS 114(8): E1336-E1344.
- Zhang R, Ehigie JO, Hou X, You, X, Yuan C (2017) Steady-State-Preserving simulation of genetic regulatory systems, Computational and Mathematical Methods in Medicine pp1-16.
- Morgan JJ, Surovtsev IV, Lindahl PA (2004) A framework for whole-cell mathematical modelling. Jl Theor Biology 231(4): 581-596.
- Surovtsev IV, Morgan J, Lindahl PA (2007) Whole-cell modeling framework in which biochemical dynamics impact aspects of cellular geometry, Jl Theor Biol 244(1): 154-166.
- Maria G, Berger D, Nastase S, Luta I (2012A) Modelling alternatives of the irinotecan release from functionalized mesoporous-silica supports, Microporous and Mesoporous Materials 149(1): 25-35.
- Maria G, Stoica AI, Luta I, Stirbet D, Radu GL (2012B) Cephalosporin release from functionalized MCM-41 supports interpreted by various models, Microporous and Mesoporous Materials 162(11): 80-90.
- Michal G (2014) Roche co. report, Basel CH, Source.
- Torres NV, Voit EO (2002) Pathway analysis and optimization in metabolic engineering, Cambridge University Press, Cambridge (MS) 2002.
- Cornish-Bowden A (2016) Biochemical evolution- The pursuit of perfection, Garland Science, New York.
- Brazhnik P, de la Fuente A, Mendes P (2002) Gene networks – how to put the function in genomics, Trends in Biotechnology 20(11): 6.
- Savageau MA (2002) Alternatives designs for a genetic switch: Analysis of switching times using the piecewise power-law representation. Math. Biosciences 180: 237-253.
- Nandy SK (2017) BioXYZ, what is XYZ ?, Trends Biomedical Eng. & Biosci3(2): 555606.
- Sutherland W (2005) Optimization- this beguilingly simply idea allows biologists not only to understand current adaptations, but also to predict new designs that may yet evolve, Nature
- Wikipedia (2022) Bioinformatics, last accessing.
- Silva MR (2006) Bioinformatics tools for the visualization and structural analysis of metabolic networks, PhD Thesis, TU Braunschweig Germany.
- Janowski SJ (2013) VANESA - A bioinformatics software application for the modeling, visualization, analysis, and simulation of biological networks in systems biology applications, PhD thesis, Faculty of Technology at the Bielefeld University, Germany.
- Eden P (2003) Course on: Bioinformatics and Computational Biology, Faculty of Medicine, Lund University Sweden.
- Lesk AM (2002) Introduction to Bioinformatics, Oxford Univ. Press Oxford.
- Baxevanis AD, Ouellette BF (2001) Bioinformatics - A Practical Guide to the Analysis of Genes and Proteins, Wiley.
- Lengauer T (2002) Bioinformatics - From genomes to drugs, Wiley .
- Baldi P, Brunak S (2001) Bioinformatics, The Machine Learning Approach, MIT Press, Cambridge, Massachusetts.
- Isaev A (2004) Introduction to mathematical methods in Bioinformatics, Springer.
- Eidhammer I, Jonassen I, Taylor WR (2004) Protein Bioinformatics - An algorithmic approach to sequence and structure analysis, Wiley.
- Luscombe NM, Greenbaum D, Gerstein M (2001) What is bioinformatics? An introduction and overview, Yearbook of Medical Informatics 2001(1): 83-99.
- Banga J (2008) Optimization in computational systems biology. 6-th Simon Stevin lecture on optimization in engineering, Leuven-Heverlee, NL. Also in BMC Systems Biology 2: 47.
- Moulijn JA, Makkee M, van Diepen A (2001) Chemical process technology, New York, Wiley.
- Maria G, Gijiu CL, Crişan M, Maria C, Tociu C (2019) Model-based re-design of some genetic regulatory circuits to get Genetic Modified Micro-organisms (GMO) by using engineering computational tools (a mini-review), Current Trends in Biomedical Engineering & Biosciences 18(3): 555988.
- Hempel DC, (2006) Development of biotechnological processes by integrating genetic and engineering methods. Life Sci 6: 443-447.
- Price ND, Reed JL, Palsson BO (2004) Genome-scale models of microbial cell: evaluating the consequences of constraints, Nature Reviews Microbiology 2(11): 886-897.
- Maria G, Crisan M, Maria C (2016) Parameter estimation of the (Bio)chemical process kinetic models, Printech Publ., Bucharest p528.
- Basso A, Serban S (2020) Overview of Immobilized Enzymes’ Applications in Pharmaceutical, Chemical, and Food Industry. In: Guisan, J, Bolivar, J, López GF, Rocha MJ (eds), Immobilization of Enzymes and Cells. Methods in Molecular Biology, 2100, Humana, New York.
- Buchholz K, Hempel DC (2006) From gene to product (Editorial), Life Sci 6: 437-437.
- Górecka= E, Jastrzębska M (2011) Immobilization techniques and biopolymer carriers, Biotechnol Food Sci 75(1): 65-86.
- Nedovic V, Willaert R (2005) Applications of cell immobilisation technology, Springer verlag, Amsterdam.
- Maria G, Luta I, Maria (2013) Model-based sensitivity analysis of a fluidised-bed bioreactor for mercury uptake by immobilised Pseudomonas putida cells, Chemical Papers 67(11).
- Schuetz R, Kuepfer L, Sauer W (2007) Systematic evaluation of objective functions for predicting intracellular fluxes in E. coli, Molecular Systems Biology 3: 119.
- Yang X, Mao Z, Huang J, Wang R, Dong H, et al.(2022) The necessity of considering enzymes as compartments 1 in constraint-based genome-scale metabolic models, BioRxiv - The preprint server for biology, Cold Spring Harbor Lab (USA).






























