An Update on Genome Editing with the Utilization of CRISPR/Cas 9 System for Evaluation and Treatment of Human Diseases - A Systemic Review
Kulvinder Kochar Kaur1*, Gautam N Allahbadia1 and Mandeep Singh2
1Scientific Director, Centre for Human Reproduction, Punjab, India
2Consultant Neurologist, Swami Satyanand Hospital, Punjab, India
Submission:March 15, 2022;Published:May 04, 2022
*Corresponding author:Kulvinder Kochar Kaur, Scientific Director, Centre For Human Reproduction 721, G.T.B. Nagar, Jalandhar-144001, Punjab, India Curr
How to cite this article: Kulvinder K K, Gautam N. A, Mandeep S. An Update on Genome Editing with the Utilization of CRISPR/Cas 9 System for Evaluation and Treatment of Human Diseases - A Systemic Review. Curr Trends Biomedical Eng & Biosci. 2022; 20(5): 556046. DOI:10.19080/CTBEB.2022.20.556046
Abstract
The CRISPR/Cas9-techique again emerged in 2012 with the generation of the Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)-based gene editing that represents a modulation tool that has been obtained from the defense system of the some bacteria, against viruses in addition to plasmids. This is an economical , simple approach that has got utilized in a lot of experimental models inclusive of cell lines, various laboratory animals, plants, as well as human Clinical trial. The CRISPR/Cas9 system is constituted of guiding the Cas 9 nuclease for generation of site-directed double stranded DNA break with the utilization of small RNA molecules for directing. It is an event that results in permanent manipulation of the genomic target sequence which can heal the injury that occurs in the DNA. Thus here we conducted a systematic review utilizing search engine pubmed, google scholar and other sutilizing the MeSH terms like zinc finger nucleases; TALENs; CRISPR/Cas9 system; DSB; genome editing; trans-encoded small cr RNAs(tracr RNAs); single guide gRNA; Protospacer Adjacent Motif (PAM); endonuclease; HNH domain; Ruv C-like nuclease domain; insertions along with deletions (indels); dead Sp- Cas9 (d Sp- Cas9); n Sp- Cas9 (n Cas9)]. Hirshsprung disease; Megacystis-Microcolon-Intestinal Hyperperistalsis Syndrome (MMIHS); β-haemoglobinopathies; Sickle Cell Disease (SCD); β thallasaemia, is Human Papilloma Virus (HPV); human immunodeficiency virus(HIV); Hepatitis B virus(HBV); cancers; neurological diseases; from 1990 to 2021 till date. We found 2050 articles out of which we selected 141 articles for this review. No meta-analysis was done. Here we present approaches as well as manipulations of enzyme Cas9 for removal of target cuts away the various applications of CRISPR/Cas9 system for looking besides activation as well as repression. Furthermore, we outline the therapeutic aspects besides the latest updates in their utilization in various human diseases.
Keywords: Juju dhau; ICRISPR/Cas9 system; Tracr RNAs; PAM; Hirshsprung disease; HPV; HIV; Mitochondrial diseases’ breast cancer
Introduction
For estimation of the function of a gene, a gene can get inactivated by homologous recombination, or via its blockade of its messenger RNA via RNA interference [1]. This strategy is applicable, in case of cultured cells by their transfection or in living organisms by transgenesis [1]. More Recently, advancement in case of genome editing aid in its manipulation of any type of gene at its originating locus in a different kind of species along with tissues that is inclusive of cultured cells as well as animal organs. Genome editing is a robust tool with regards to biomedical research, besides aiding in rectification of certain inherited diseases.
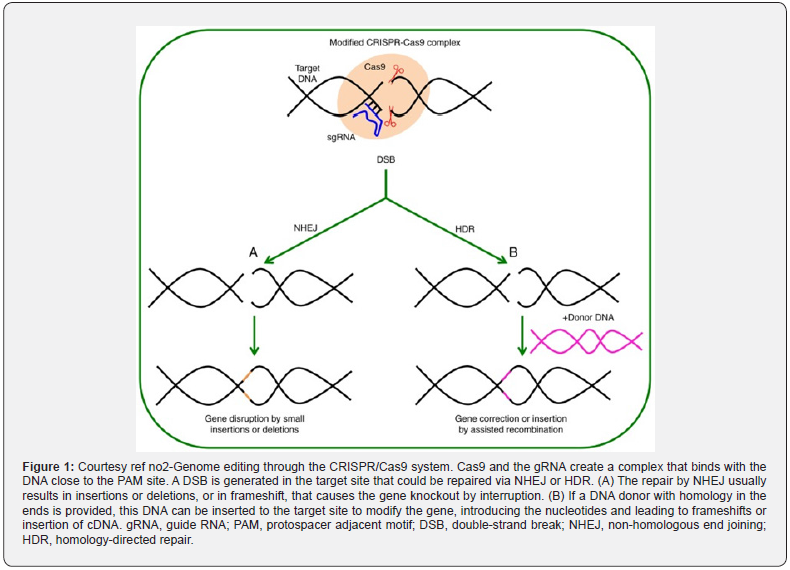
Genome editing is dependent on the utilization of markedly particular in addition to programmable nucleases, that generate particular alterations in the areas of attraction in the genome via introduction of double strand breaks (DSB), which get later healed by cellular modes. These healing modes are inclusive of the i) non homologous end joining (NHEJ), which carries a susceptibility to mistakes. II) homology directed repair (HDR) which is free of these mistakes (Figure 1) [2]. Healing aids in the development of insertions in addition to deletions or substitutions in the target region [3,4]. These mutations might stop, remove or rectify the abnormalities in the genes. These latter probabilities aids in the capacity of rectification of the mistakes that are inherited in DNA,besides resulting in disease. The programmable nucleases are basically mega nucleases [5], Zinc finger nucleases transcription activator like effectors nucleases (TALEN) in addition to the CRISPR/Cas9 system (that implicates the Clustered regularly Interspersed short Palindromic Repeats nuclease(s) correlated with CRISPR [6-8]. Commonly, the CRISPR/Cas9 system constituents, are a guide RNA, which gives direction to the Cas9 nuclease for generation of a DSB in a particular site of the genome. More recently, in the past decade greater interest, besides its acceptance over the other systems, in view of it being easy, rapid, along with effectiveness for the manipulation of the endogenous genes in any cell or target tissue, in case the organisms that possess the maximum toughness with regards to the treatment.
Methods
Thus here we conducted a systematic review utilizing search engine pumped, Google scholar and others utilizing the MeSH terms like zinc finger nucleases; TALENs; CRISPR/Cas9 system; DSB; genome editing; trans-encoded small cr RNAs (tracr RNAs); single guide gRNA; protospacer adjacent motif (PAM); endonuclease; HNH domain; RuvC-like nuclease domain; insertions along with deletions (indels); dead Sp- Cas9 (d Sp- Cas9); n Sp- Cas9(n Cas9). Hirshsprung disease; megacystis-microcolon-intestinal hyperperistalsis syndrome(MMIHS); β-haemoglobinopathies, sickle cell disease (SCD); β thallasaemia, is human papilloma virus (HPV); human immunodeficiency virus (HIV); Hepatitis B virus (HBV); cancers; neurological diseases from 1990 to 2021 till date.
Results
We found a total of 2050 articles out of which we selected 141 articles for this review. No meta-analysis was done
CRISPR/Cas9system
The invention of CRISPR/Cas9 system
The eubacteria in addition to archaea contain a defense system that is meant for getting acclimatized, via RNA, for recall besides, destruction of the exogenous DNA, besides RNA. This gives adaptive immune response against the invading plasmids along with viruses [9]. The observation of this system has been seen in 50% of the bacterial genome which are sequenced as well as in 87% of the archaeal genome [9,10]. It is further significant in a broad range of extra functions, that is inclusive of replication divisions [11], acclimatization to high temperature, chromosomal reorganizations, besides, healing of DNA [12-14].
The invention of CRISPR occurred in case of Escherichia coli genome in 1987 in the forms of repeated fragments of 29 nucleotides (nt) in length that are Interspersed with different sequences of recurrent fragments of 32 nt [15]. With the invoked interest in the CRISPR system along with its correlated Cas genes resulted in the invention of akin short recurrent Palindromic sequences of 24-40 nt in various groups of bacteria as well as archaea. These recurrent sequences are distant by individual varying sequences of 20-58 nt [9,16]. The correlated genes (Cas) got isolated usually intricately with a CRISPR locus, which pointed that there was a functional correlation [17]. In the beginning it was posited that the function of CRISPR locus was that it participates in cellular DNA healing in addition to replicon partitioning events; nevertheless, the initial proof with regards to the significance of CRISPR/Cas system belonging to the adaptive prokaryotic immune system was documented in 2005 via the finding that most of the sequences were inserted among same repeats got obtained via the invading phage along with plasmid genomes [18]. Furthermore, in 2007 the insertion of new spacers got illustrated in a CRISPR/Cas locus of Streptococcus thermophiles [19], whereas the transcription altering to small mature CRISPRRNAs( cr RNAs) which directed the Cas complex of Escherichia coli, got experimentally corroborated in 2008 [20]. Subsequently in 2010, Cas complex of Streptococcus thermophilus was illustrated to generate a single DSB at a particular position in case of target DNA [21], that was followed by the documentation with regards to the maturation of cr RNAs needs trans-encoded small cr RNAs(tracr RNAs), Cas 9 in addition to RNAse III in Streptococcus pyogenes [22]. The proof of function in a heterologous system was revealed in2011 when it got demonstrated, that the CRISPR/ Cas system of Streptococcus thermophilus once transported to Escherichia coli, yielded heterologous immunity against plasmid along with phage infection [23]. Simplifying CRISPR/Cas 9 of Streptococcus pyogenes system got attained via replacementnt of tracr RNAs in addition to cr RNAsin2012 by a synthetic single gRNA to guide Cas 9 towards its target in addition to carry out cleavage [8]. Ultimately in 2013 the utilization of CRISPR/Cas 9 system (type II Streptococcus pyogenes) was detailed in the form of a genome editing tool for the initiation of site particular DSB in addition to following mutagenesis in plants, mouse, besides, human cells in addition to clinical trials [24,25].
Working of CRISPR/Cas 9 system in the form of Genome editing tool
The type II CRISPR/Cas 9 system is the system whose utilization with regards to genome editing with the utilization of Cas 9 endonuclease of Streptococcus pyogenes whose properties have been well studied. In case of endogenous system, the mature cr RNAs unites with a tracr RNAs (ie small RNA which is complementary to the CRISPR sequences)for generation of a tracr RNA: cr RNA complex, that directs the Cas 9 to a target region. Hence, CRISPR/Cas 9 conducts sequence particular cleavage via simple crosstalk of cr RNA by base pairing at the target region. Subsequent to the union of the target region, the two DNA strands get cleaved through the nuclease domain of Cas 9, that is a HNH domain which cleaves the complementary target strand to the gRNA in addition to Ruv C-like nuclease domain, that cleaves the non target strand [26]. The gRNA that got fashioned by Jinek et al. [8], in 2012 was a chimerical RNA that possesses all the necessary, constituents of cr RNAs in addition to tracr RNAs for directing Cas 9. Subsequently a lot of variations of CRISPR/Cas 9 have got generated, that recalls the sequences of 18-24 of the gRNA, in addition to the 2-4nt of protospacer adjacent motif (PAM) in target regions [4,27]. Hence CRISPR/Cas 9 can be guided towards a particular sequence of22-29nt that is distinctive in maximum genomes, despite noticing that CRISPR/Cas 9 possesses a lot of tolerance for non particular mating of base pairs among gRNA, in addition to its complementary target sequence. There exists a specificity which possesses sensitivity toward amounts, position as well as distribution of incorrect crosstalk [4,8,24]. Like the CRISPR/Cas 9 of Streptococcus pyogenes (sp Cas 9) has tolerance for up till 6 non balances of base pairs at the target region [8]. The genome editing brought about by CRISPR/Cas 9 is based on the development of the DSB in addition to the following event of DNA healing. The DSB formed by the CRISPR/Cas 9 initiates the event of cell healing in DNA, in the form of NHEJ, that possesses is susceptible to going incorrect as well as hence possess the capacity of generation of mutation, that implicates small insertions along with deletions (indels) in case of target region that can bring a pause or delete the function of the genes or genomic target elements (like controlling areas). One separate healing event which can further get stimulated is the HDR error free, that has the possibility of rectification of innate disease, resulting in mistakes of DNA (genes or controlling elements [28].
PAM sequence in addition to off target cuts
How specific CRISPR/Cas 9 is in addition to complementarity of the gRNA/ target sequence, needs a PAM sequence which resides just subsequent to the target sequence. The dependence on the PAM sequence for the cleavage of the DNA, limits the speed of the cleavage region in the genomes, hence target areas are observed more commonly for small PAM sequences in contrast to longer ones; thus subsequently off target cut areas possess, lesser chances to be present for the longer ones, in contrast to shorter ones. The isolation of PAM sequences differ among separate microorganisms, as well as these sequences have got documented: 5’NGG-3’ in Streptococcus pyogenes (sp Cas9) [8]. 5’NGGNG-3’ or 5’NNAGAAW-3’in Streptococcus thermophiles (St1 Cas 9) [21,29]. 5’NNGRRT-3’ or 5’NNGRR(N)-3’ in Staphylococcus aureus (Sa Cas9) [30]. 5’NNNRRT-3’or 5’NNNNGMTT-3’ In Neisseria meningitides (NmCas9) [31], as well as 5’NGG-3’ in Francisella Novivida (FnCpf1) where N point to nucleotides [32], R to purines, A or G, M to nucleotide A or C, as well as W to weak bonds A or T [4,10,27,33]. Despite, the DSB action of CRISPR/Cas 9 is dependent on the complementarity of the target sequence with the gRNAs of 20nt in length, the system aids cleavage at genomic areas that are partly complementary to the gRNA, since the gRNA lets mismatch pairing among the DNA in addition to gRNA [4]. These off target cuts of CRISPR/Cas 9 is remain one of the largest problems that at present are unwanted as well as not solved (resulting in unwanted outcomes) as well as vary among separate target areas in view if the variability of nucleotide sequence in addition to with regards to the genome. The utilization of CRISPR/Cas 9 system has got done with success with regards to gene editing, in addition to for the regulation of various biological systems, nevertheless, little proof is existent with regards to the consequences of the off target genome editing. Actually, just an occasional assay about genotoxicity that is dependent on cell in culture which aid in quantification, stratification in addition to aids in avoidance of biological adverse actions of gene editing in a particular target cell population. Despite, a lot of gene editing studies have arrived at the phase with regards to clinical trials, clinical proof that demonstrated that gene editing possess the capacity of treatment of various conditions is minimal [23].
Various methods have got posited for maximizing the gRNA fashioning besides reduction of the off target cuts to arrive at an acceptable in addition to specificity that is essential for safety with regards to treatment applications [34]. Isolation of the ideal gRNA of the different applicants for a specific target area besides the placement of probable off target can be validated via various bioinformatics tools [35]; like CRISPR direct [36], E- CRISPR [37], WU- CRISPR [38], CRISPR- gRNA design tool (http://www. atum.bio/eCommerce.cas9/input), sg RNA Designer [39], sg RNA Scorer Scorer 2.0 [40], CRISPR-scan [41], CRISPR-ERA [42], CCtop [43], CRISPOR [44], Breaking –Cas [34], CHOP- CHOP [45], CRISPR-Design [48], CRISPR-Multi target [46], GT- Scan [47], ge- CRISPR [47], CRISPR- Designer [48], Cas- Designer [49], Cas-OF Finder [50], COSMID [51], DESKGEN Guide Picker [52], as well as CRISPR- gene analyzer [53]. All these software programmes possess various properties in addition to applications. The ideal choice would be based on the kind of organism (eukaryotic or prokaryotic) for which the gRNA fashioning has to be carried out, the different PAM has to get utilized, the size of the DNA target, besides the different types of currently documented Cas-like nuclease [34].
Approaches along with Manipulation of The Enzyme Cas9 for Reduction of Off- Target Cuts to Least
The fashion of the differences in classical Sp- Cas9 system has been evaluated with regards to the regulation of its output, like utilization in the form, of an inducible system for interim expression of the nuclease [54]. An alternative approach for reduction of action of off- target is to exchange the Sp- Cas9 enzyme for mutant n Sp- Cas9 (n Cas9), that breaks a single guide strand via inactivation of a nuclease domain, Ruv C HNH. Here for carrying out a DSB, two n Cas9 are required for targeting the opposite strands of DNA intricately placed (separated just via<100bp), with every Cas9 directed via its own sgRNA which possess the capacity to cleave just the strand that is complementary to sgRNA [33,55].
On utilization of 2 n Cas9 is carried out for the performance of DSB off- target action reduction takes place by 50-1500 fold [33,56], specificity by amelioration of the helicase action (e Sp- Cas9) for interference of off- target regions without disruption with specificity in addition to action [57]. Fusion of the nucleaseFok1 with dead Sp- Cas9 (d Sp- Cas9) further resulted in escalation of specificity; here dSp- Cas9 has absence of its cleavage action via inactivation of its 2 domains of the nuclease, namely Ruv C as well as HNH domains. As a consequence the fusion generates RNA-directed Fok1 nuclease, hence the action with regards to excision for the DSB’s will be on the basis of the 2 bounds of the 2 gRNAs to DNA which would comprise of a well defined spacing in addition to orientation, that aids in the dimerization of monomers Fok1- dSp- Cas9 for the generation of a catalytically active Fok1 dimer, that results in reduction in the probability that a target sequence is appropriate repeatedly in the genome as well as thus result in escalation of specificity [45,46]. A separate manipulation, of the binary system Split-Sp Cas9 that causes utilization of the nuclease lobe as well as the α-helical domain with independence. These 2 exist naturally in the enzyme Cas9 by themselves solely or as an attachment to the DNA via the gRNA. The domains do not crosstalk by themselves however, gRNA results in the recruitment of all of them in a tertiary complex which recapitulates the action of Cas9 along with catalyzing the region particular DNA cleavage. The utilization of manipulated RNA delete the action of Cas9, that breaks the dimer, that aids in the generation of an inducible besides, the dimerization system that can be adjusted with regards to genome editing applications. This system has got evaluated both in vivo, in addition to in vitro in case of mouse models [59]. Of the other manipulations till date, the Sp Cas9-cytidine deaminase can be observed that is a fusion of dSp- Cas9 with the enzyme cytidine deaminase. This effect of cytidine deaminase, causes transformation of cytosine(C) to uracil (U), that act on a single stranded DNA that can get accessed in the tertiary complex, among Cas9 gRNA, in addition to the target DNA for introducing point mutation [60].
Newer variations of the CRISPR system
Other studies have isolated more enzyme belonging to the CRISPR family that were inclusive of enzymes that got encoded by Cas genes that were smaller in contrast to Sp Cas9 (4.2Kb) like Sa Cas9, St1 Cas9 in addition to Nm Cas9 (3.2, 3.4 as well as 3.2kb, respectively [28,33]. These enzymes would promote its getting packed into viral vectors [28,33]. Other more recently discovered enzyme known as Cpf1 possessing shorter crRNA sequence can get utilized rather than Sp Cas9 [32]. Furthermore, the more recently invented C2c2 (Cas13a) as well as C2c6 (Cas13b) that possess the capacity of cleaving RNA [61]. Cas13b had been generated in the form of RNA base editing technique, whose utilization as a tool for editing RNA with the utilization of catalytically inactive Cas13b that had got fused to the adenosine deaminase domain of ADAR2 for Programmable adenosine to inosine replacement in case of transcripts [61].
More applications of CRISPR/Cas9
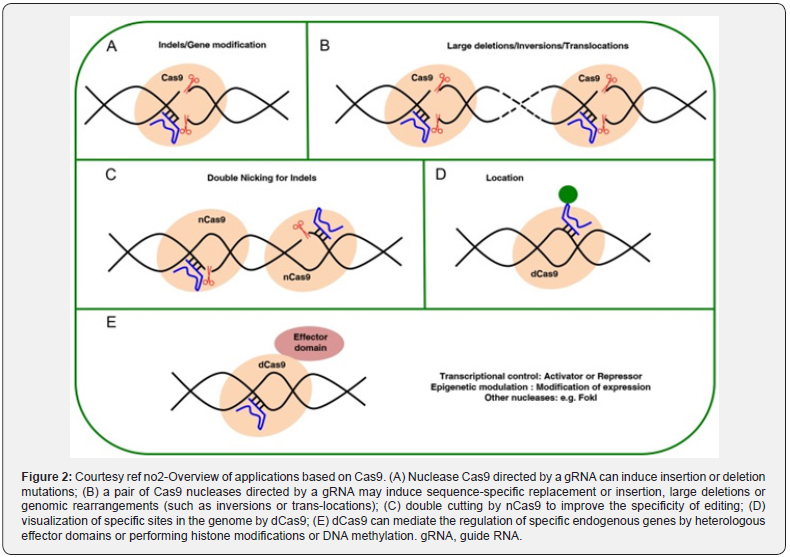
Apart from the utilization of CRISPR/Cas9 system for the development of indels (Figure2), manipulated version have got generated for various applications, like activation or suppression via controlling of gene expression by fusion of hetrologous domains with the, manipulation of epigenetic signatures on histones (i.e. alteration of methylation patterns) or transcriptional activators or repressors [62]. Utilization of CRISPR/Cas9 has been carried out regarding selective labeling of the genome, like for monitoring of dynamic chromatin events [63].
Disease models as well as CRISPR/Cas9 system
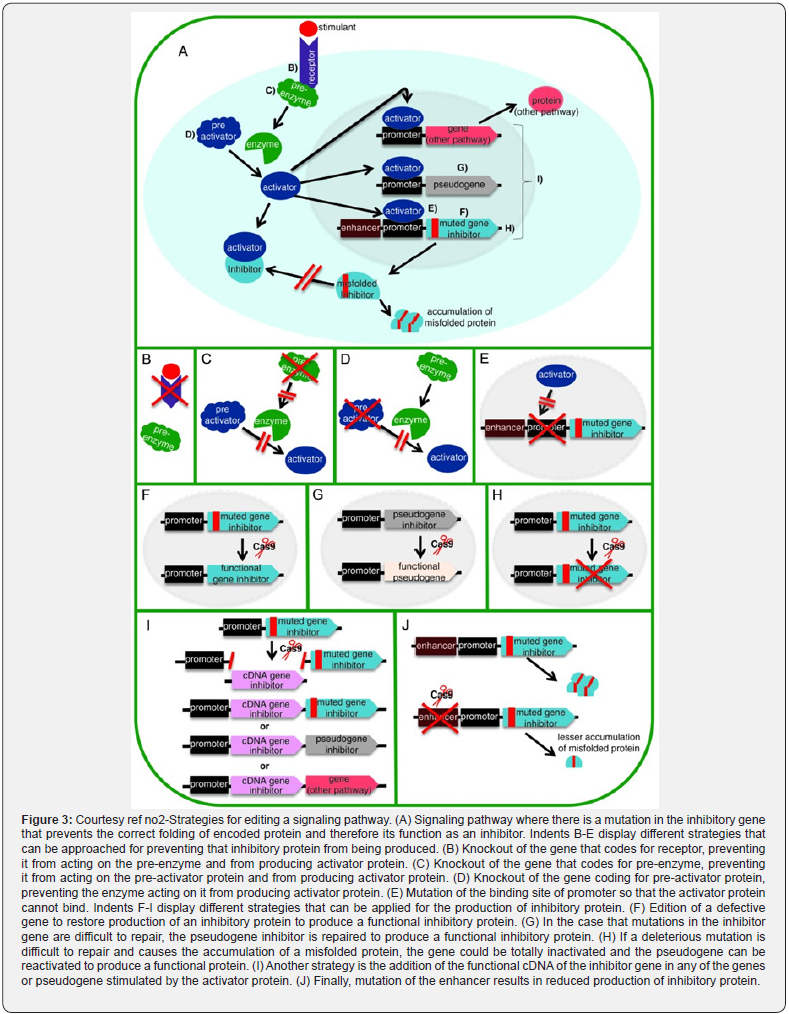
The CRISPR/Cas9 system possesses a miraculous capacity of treatment of various diseases where the etiology of genetic impairment, is clear, or for evaluation of these diseases via the generation of cell/animal models. Treatment which is dependent on genome editing can result in the re-installment of gene function or compensate for the mutation. Single nucleotide polymorphism (SNP) editing has got targeted with the utilization of various approaches, like i) gene knock out that results in disease [64], ii) adding a mutation that confers protection [65], iii) or introduction of a transgenic that possess the capacity of treatment [66]. In case the etiology of the disease is a virus, cleavage of the viral DNA can get carried out [67-70]. Figure 3 illustrates some of the approaches that have got detailed earlier, as well as whose utilization can be done as a strategy for various diseases. The idea of this figure is for expansion of the treatment probabilities of CRISPR/Cas9 system. If facing problems with regards to the repairing of a mutation In view of genomic aspect, there might be the existence of a pseudogene whose activation might aid In the replacement of the gene having undergone mutation [71]. Nevertheless, in case the etiology of the disease is a protein resulting in injury to the organism via its aberrant properties, (like misfolding as well as collection in a tissue, like amyloidosis), down regulation of its expression at various points of its pathway where it gets expressed [72].
A. Pulmonary as well as Gastrointestinal diseases as well as CRISPR/Cas9
The homozygous Δ 508 mutation in the cystic fibrosis transmembrane conductance regulator (CFTR) gene rectification was done with the utilization of CRISPR/Cas9, in vitro in case of intestinal stem cells of cystic fibrosis patients [73]. Edited stem cells were differentiated into intestinal organoids in addition to demonstrated a functional, CFTR product [73]. CRISPR/Cas9 has further been illustrated to have probable advantageous actions, in patients that need a liver transplantation, like drug treatment resistant metabolic liver conditions [69]. Utilization of CRISPR/ has been conducted for repression of genes, hence reprogramming metabolic pathways, that aids in attaining a benign phenotype in the treatment of hereditary tyrosinaemia in case of mice [60]. With the aid of deletion of 4-hydroxy-phenylpyruvatedioxygenase (HPD) gene with the idea of hampering the second step in tyrosine catabolism, the modulated hepatocytes (FAH-/- HPD- /-). The replacement was about to get finished in many mice (92-99%) in 8wks. The HPD mutation resulted in escalation of catabolism of tyrosine, that prevented the collecton of tyrosine in addition to toxic metabolites, like fumaryl acetoacetate as well as succinylacetone in hepatocytes, that in turn causes robust liver injury with escalation of, chances of Hepatocellular carcinoma (HCC) [69].
Furthermore, utilization of CRISPR/Cas9 system can be done with regards to the treatment of Hirshsprung disease in addition to megacystis-microcolon- intestinal hyperperistalsis syndrome (MMIHS), where some mutations were isolated via whole -exome sequencing, in case of patients in which these diseases had got diagnosed [74,75]. In case of Hirshsprung disease 4 genes (DENND3, NCLN, NUP98 along with TBATA) have got correlated with the neuronal events that get shared by the central nervous system(CNS) as well as enteric nervous system (ENS), this function got validated in vivo via a gene knockout by CRISPR/ Cas9 [74,75]. utilization of a zebrafish model was done to see that the knockout of the genes detailed earlier caused a deletion of function in addition to break in the generation of the ENS (deletion of enteric neurons) resulting in akin Hirshsprung phenotype in vivo [74]. The LMOD1 gene is implicated in generation of normal smooth muscle-cyto-skeletal-contractile coupling, as well as mutation results in MMIHS. Mice that are knockout of LMOD1 gene demonstrated, an akin phenotype of this syndrome (protein amounts as well as pathology that agreed with MMIHS), besides, demonstration of the starting of bladder distension at18. 5d of embryonic generation till it approached the abdominal cavity, whereas the histological evaluation observation was the early initiation of thinning of the detrusor muscle of the bladder of LMOD1-/- mice. These outcomes pointed to a part of the LMOD1 gene in the generation of normal smooth muscle-cyto-skeletalcontractile coupling [75].
5.3B. CRISPR/Cas9 as well as Haematological Diseases
β-haemoglobinopathies which are inclusive of sickle cell disease (SCD) as well as β thallasaemia, occur secondary to mutations in the β-globin (HBB) gene, that affects millions of people all over the world. Utilization of CRISPR/Cas9 in addition to donor sequences have been done by various studies for attainment of homologous recombination in the HBB gene in Induced pluripotent Stem Cells (iPSCs) in addition to haematopoietic Stem Cells (HSC) [76,77]. Like rectification of the Glu6Val mutation with efficiency that is the etiological factor for SCD was attained with the utilization of pluripotent Stem Cells that had been obtained from patients, besides, differentiation to erythrocytes, that led to the expression of the mRNA of adult β-globin (HbA) along with validation of the intact transcriptional controlling of the manipulated HBB alleles [76]. The other genome editing approach for the therapy of haemoglobinopathies is the intercalation of a mutation for the development, of a benign genetic disorder. Once there is hereditary persistent fetal haemoglobin (HPFH), a benign genetic disorder, the mutation ameliorates the transfer of γ globin to -β-globin, resulting in a large concentrations of fetal haemoglobin expression (HbF) all through the life, that can aid in relief of the Clinical signs of β thallasaemia or SCD. In an earlier study utilization of CRISPR/Cas9 for simulation of the HPFH mutation in case of promoters of the HBG1 as well as HBG2 genes in human blood progenitor cells generated red blood cells (RBCs). The progenitor cells that had got edited resulted in generation of RBCs with escalation of HbF amounts which were enough for hampering the pathological hypoxia of RBCs that got observed in SCD [78]. A separate approach for the escalation of HbF amounts was the CRISPR/Cas9 mutation of the BCL11 erythroid escalator, that has been a corroborated suppressor of HbF in addition to the treatment target for the β-haemoglobin conditions [79]. Moreover, β thallasaemia that was secondary to the expressionsuppression point mutations or deletions in the β-globin gene, efficient rectification of the HBB mutations by CRISPR/Cas9 was attained in iPSCs cells obtained, from a patient that differentiated in erythroblasts that possessed the restoration of the expression of β-haemoglobin [77].
Lastly CRISPR/Cas9 has got utilized with regards to the treatment of a lot of other haematological diseases like auto alloimmune bleeding conditions, that is inclusive of fetal in addition to neonatal allo immune thrombocytopenia besides posttransfusion purpura for conversion of Leu33+ megakaryocytes – like DAM1 cells as well as iPSC towards the Pro33 allotypes, that is implicated in the generation of human plateletsalloantigen Ia as well asa Ib epitopes [80]. Moreover, the utilization of this technology has been carried out with regards to the treatment of Fanconi aaemia by rectification of point mutation in patient obtained fibroblasts [81], in addition to haemophilia for the rectification of factor VIII deficiency in mice [82]. Noticeably utilization of CRISPR/Cas9 has been done for the development of mutant pigs in the vWTF gene for working as an animal model for von Willebrand disease, or for promotion of bleeding before meat processing in the context of meat industry [83].
C. Viral diseases as well as CRISPR/Cas9
Viruses represent obligate intracellular pathogens which possess the capacity of infecting cells via particular receptors as well as require cellular constituents of the host for their reproducibility. On entrance within the cells, the Viral genome gets replicated, transcribed in addition to translation for finishing its life cycle [84]. A lot of genomes of the DNA Viruses besides the retro viruses get incorporated into the cellular genome. Robust disease can result secondary to infection with lot of mortality, morbidity along with followed by transmission to other human beings. Reduction of some of the viral infections can get attained by vaccination immunity, whereas it is not feasible with regards to others. The viral infections whose requirements is greater concentration in view of their kind besides the social influence are those whose etiology is human papilloma virus (HPV), human immunodeficiency virus (HIV), as well as the Hepatitis B virus (HBV).
Expression of HPV 16 variants of the oncoproteins E6 as well as E7, that are closely correlated with the generation besides sustenance of the malignant phenotypes which can cause cervical cancer [69], that is the second commonest etiology of cancer all through the world [68]. Either solely or in combination CRISPR/ Cas9 utilization has been carried out with the other therapies for In vitro and in vivo synergistic therapeutic studies to tackle HPV 16 in addition to 18 causative factors of cervical cancer [68,69,85]. Targeting of HPV 16 as well as HPV 18 oncogenes E6 besides E7 by CRISPR/Cas9 in case of cervical cancer cell lines, that was inclusive of HeLa as well as SiHa cells resulted in cell cycle arrest with ultimate demise of these malignant cells [85]. Further in a study by Zhen et al. [69], mutation of the HPV 16 oncogenes E6 as well as E7 hampered the tumor growth in vivo that illustrated that therapy with CRISPR/Cas9 acts as a treatment in combination with cisplatin, which is one of the 1st line therapies that is maximum utilized in the form of a chemotherapy agent [69].
As far as C-C chemokine receptor 5(CCR5) acts in the form of a necessary co- receptor for HIV-1 virus, hence deletion of CCR5 receptor function confers protection against this viral infection. Utilization of CRISPR/Cas9 was done with the idea of editing the CCR5 gene in CD4+cells as well as in human iPSCs(hiPSCs) cells [65,86]. These mutant hiPSCs differentiation into macrophages which developed resistance to trophic CCR5 for HIV-1 virus [65]. The continuous existence of a covalently closed circular DNA (ccc DNA ) of HBV works as the biggest hurdle for anti viral treatment for chronic hepatitis B getting eradicated. For obtaining a complete cure total deletion of the continuous existence of ccc DNA the requirements would be the total elimination of hepatocytic viral load [67]. In case of a lot of studies with the utilization of a particular gRNA against the HBV, significant reduction, in the generation of HBV as well as core proteins inHuh7 occurred subsequent to CRISPR/Cas9 system utilization [67]. HepG2 [87], HepG2. 2. 15 [87,88], HepG2-H1. 3 cell, lines that got transfected with a HBV expression vector [86]. Moreover, in a mouse model CRISPR/Cas9 system cleaved the intrahepatic plasmid that possessed the genome as well as promoted its elimination in vivo which caused a decrease of the antigen surface concentrations. This pointed that the CRISPR/Cas9 system possessed the capacity of impairment of HBV, both In vitro and in vivo, that suggested that it probably possesses the capacity of eradication of continuous existence of this viral infection [67,88]. Besides its utilization in the treatment of this viral infection, CRISPR/Cas9 system has further been utilized for the evaluation of cellular modes which result in carcinogenesis [89].
With regards to hepatitis C virus (HCV) which is the etiological factor for chronic hepatitis, in addition to Hepatocellular carcinoma (HCC), utilization of CRISPR/Cas9 has got done for the isolation of key host constituents for HCV infection [90]. Apart from that Epstein Barr virus (EBV) causes generation of continuous infection, all through life in 90-95% adults. Despite, not resulting in disease in healthy carriers, this infection is correlated with lymphoid as well as epithelial neoplasms, like Burkitts lymphoma, Hodgkin’s disease as well as nasopharyngeal carcinoma [91]. CRISPR/Cas9 system utilization has got done In vitro against Raji cell lines that illustrated a significant decrease in proliferation in addition to viral loads, besides resurrection of the apoptosis pathway In cells following therapy [92]. The other approach for this virus was the elimination of the promoter area (558nt), that is 1 of the 2 clusters, which codes for22 differentiatially expressed premicroRNAs at the time of latent EBV infection, besides is thought to be implicated in epithelial cells conversion [91]. This area was removed in particular human epithelial cell lines with latent EBV infection, inclusive of nasopharyngeal carcinoma C666- 1 cells. This was evaluated for checking if it is needed for infection besides conversion of epithelial cells, that caused the deletion of BART miRNA expression as well as action. The observation was that pBART deletion was lesser in contrast to those of WT virus as estimated in Raji cells, which pointed the significance of miR BART in the viral infection of epithelial cells, besides it would evoke interest to evaluate if they are specifically needed for the infection in addition to conversion of epithelial cells [91].
Immunotherapy has been demonstrated to be significantly efficacious in certain kinds of cancer caused by viruses. Gene therapy implicates involves insertion or modification of a therapeutic gene, to rectify the inappropriate gene products that cause/may cause diseases. Utilization of both these types of therapy have been done as alternative methods for prevention of cancers caused by oncoviruses. In this review, Araujode Almeida et al. [93], summarized the recent studies with regards to immunotherapy and gene therapy inclusive of the topics of oncolytic immunotherapy, immune checkpoint inhibitors, gene replacement, antisense oligonucleotides, RNA interference, clustered regularly interspaced short palindromic repeats Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)-based gene editing, transcription activator-like effector nucleases (TALENs) in addition to custom treatment for Epstein– Barr virus, human T-lymphotropic virus 1, hepatitis B virus, human papillomavirus, hepatitis C virus [93].
Moreover, HIV-1/AIDS remains a world wide public health problem. The world health organization (WHO) documented that by the end of 2019, 38 million people were living with HIV-1 all over the world, of which only 67% were accessing antiretroviral therapy (ART). Despite great success in the clinical management of HIV-1 infection, ART does not eliminate the virus from the host genome. Rather, HIV-1continues to be latent as a viral repertoire in any tissue that possesses resting memory CD4+ T cells. The removal of these residual proviruses which can reseed fullblown infection upon treatment interruption continues to be the major barrier towards curing HIV-1. Innovative strategies have recently been generated to excise or disrupt the virus from the host cells (e. g., gene editing with the CRISPR-Cas system) to permanently shut off transcription of the virus (block-and-lock and RNA interference strategies), or to reactivate the virus from cell reservoirs so that it can be eliminated by the immune system or cytopathic effects (shock-and-kill strategy). Here, Moranghuino in addition to Valente reviewed each of these approaches, with the major concentration placed on the block-and-lock strategy [94] (Figure 4).

D. Vector diseases as well as CRISPR/Cas9 system
As far as diseases that get transmitted by a vector utilization of CRISPR/Cas9 has been done for the evaluation of function of gene as well as for what drives the gene with the objective of eradication of significant diseases that get transmitted by a vector, like chikungunya, dengue, yellow fever along with malaria [95,96]. For evaluation of the function of particular genes studies have got carried out on the Plasmodium species (spp) genome, the parasite that is the etiological factor for malaria [95]. In view of the parasite residing in the RBCs the effectiveness of transfection is lesser due to the need for it to get across four membranes, (namely the RBC membrane, parasitophorous membrane, parasite cytoplasmic membrane, parasite nuclear membrane). Nevertheless, utilization of CRISPR/Cas9 technology, 100% effectiveness of gene elimination 22-45% effectiveness with regards to tagging, besides -25% effectiveness in nucleotides replacement got attained, that represents an important advancement as far as newer studies with regards to the parasite [96]. Moreover with the use of CRISPR/ Cas9 the primary Vector mosquito Aedes aegyptii was evaluated as it is implicated in transmission of lots of viruses. Here Kristler et al. [95], utilized it for evaluation of genetic as well as neurological reasons for the innate chemosensory behavior for attaining stable as well as specific loss of function mutations in 5 genes [95].
CRISPR/Cas9 modulated gene drive implicates, stimulation of inheritance that is biased, of specific alleles like gene knockouts, gene replacement, besides gene getting transformed with the idea of alteration of the total population of the, organism. The significance of this strategy lies in various spp are deleterious with regards to humans actions that is inclusive of humans health or agriculture [97]. CRISPR/Cas9 might get, utilized for Site specific selfish genes drive as tools of for the regulation and with aid of genetic engineering drive a harmful trait like imbalance of sex ratio, reduction of fertility as well as chemical sensitivity [98]. Like in Anopheles gambiae i.e. the major malaria mosquito vector, CRISPR/Cas9 system that was utilized with the idea of conferng female infertility phenotype by impairment, of 3 target genes i.e. AGAP005958, AGAP011377 as well as. AGAP007280 [98]. This study illustrated that causation of sole impairment of AGAP007280 gene with gene drive by CRISPR/Cas9 was of success with regards to targeting female reproduction [99].
E. Cancer as well as CRISPR/Cas9 system
Cancer constitutes a diseases group with the properties of genetic as well as epigenetic changes that are existent in Oncogenes in addition to tumour suppressor genes. Experiments with regards to modulation of normal as well as cancerous cell genome is key with the context of remodeling the disease, along with systematic evaluation of the genes that are implicated, in the start, propagation as well as treatment reaction of cancer. The rapid modeling of the genetic processes has assumed significant importance in view of finding the significance of genetic changes that are existent in human tumor as well as whose observations have been found via large scale sequencing of the of the genome of the cancer. The reasons why these are essential is to observe the so called passenger mutation which are presumed to have no direct, impact on the tumorigenic event, as well as those which possesses the capacity of directly or indirectly cause stimulation of mutations which facilitate the conversion of normal cells into cancer cells by causing mutation of oncogenes (i.e. facilitation of gain of function mutations) as well as /or inactivation of tumor suppressor genes (loss of function mutations) [100].
Via the CRISPR/Cas9 techique, it was found that NANOG and NANOGP8decreases the malignant potential of prostate cancer. Gene knockout of NANOG and NANOGP8 in human prostate cancer DU145 cells ameliorated the malignant potential significantly, that was inclusive of sphere generation anchorage independent growth, migration capacity besides drug resistance in contrast to the parent DU145 cells. Despite, the cell proliferation not being hampered, in vivo, in case of mice that were immunodeficient there was a significant reduction in the tumorigenic capacity [101]. For evaluation of influence of BCR-ABL fusion on the leukaemic event in the Boff-p210 cell lines, a haematopoietic cell line, that possesses independence of interleukin-3 (IL3) as well as cause expression of BCR-ABL p210, with utilization of CRISPR/ Cas9 for removal of the p210, Oncoprotein. This caused the deletion of the capacity of Boff-p210 cell line to grow in the lack of IL3, besides illustrated a significant, escalation of apoptosis amounts [102]. In case of immunodeficient mice the edited BCRABL p210 cells, generated smaller tumors in contrast to those that had got initiated from the parent Boff-p210 cells [102]. Moreover, a single clone of edited BCR-ABL cells with a distinctive frameshift mutation (that avoided the p210 Oncoprotein) could not generate tumors, a kin to the outcomes in the Baf/3 parent line [102].
Before CRISPR earlier animal models were developed for evaluation of anti cancer agents, in addition to unknown drug resistance mode used to be costly, besides time consuming [100]. For obtaining a flexible along with efficacious approach for evaluation of somatic changes of loss of function, besides their impact on tumorigenesis, CRISPR/Cas9 utilization was done for the impairment of somatic genes via unique removal of PTCH1 or triple elimination of TRP53, PTEN, as well as NF1 genes in the mouse brain, that caused the generation of medulloblastoma as well as glioblastoma respectively [103]. In certain cases Oncogenic chromosomal reorganization do a central act in the etiopathogenesis of human cancers, like the Oncogenic fusion among echinoderm, microtubule- correlated protein like 4 (EMLA4) gene as well as anaplastic lymphoma kinase (ALK) gene. The Oncogene that resulted i.e. EMLA4- ALK is seen in a subset of human non small lung cancer (NSCLC), in addition to that this is of clinical significance since this confers sensitivity to inhibitors of ALK. Utilization of CRISPR/Cas9 was done for the development of a mouse model of lung cancer that was driven through ALK inhibitors [104]. At present Clinical protocols with regards to cancer have got originated for the evaluation of application of the CRISPR/Cas9 technique in case of lung, prostate, renal esophageal as well as bladder cancer in addition to neoplasms that are correlated with HBV, as well as EBV. https://clinical trials.gov/ ct2/results?cond=term-crispr&entry=&state=&city=&dist=).
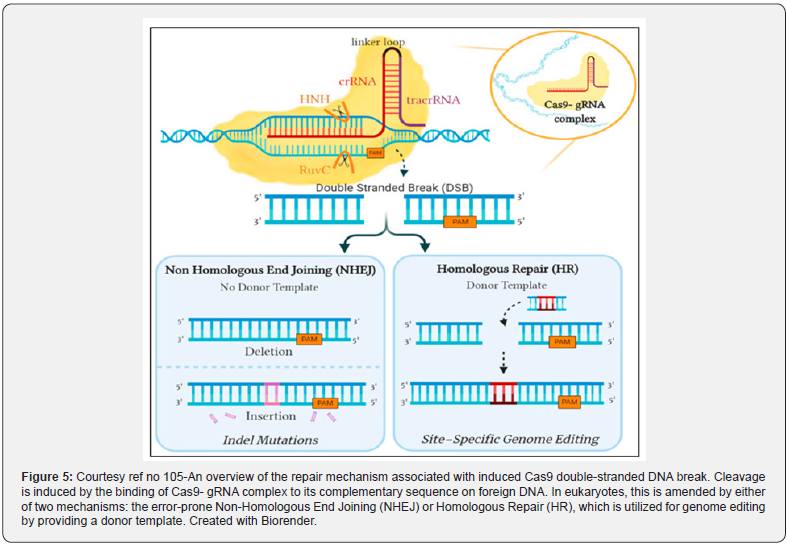
Further Breast cancer is one of the commonest types of cancer worldwide, besides is one of the leading causes of demise in women. In view of possession of multiple abnormal genes, that aid in the heterogenic properties, in addition to the multi-step pathway of tumor generation. Despite the fact that several disease initiating mutations have been isolated, treatment is usually directed at alleviating symptoms instead of rectification of the mutation in the DNA sequence. The Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)/Cas9 is a groundbreaking tool that is being utilized for the identification as well as corroboration of genomic targets bearing tumorigenic potential. CRISPR/Cas9 supersedes its gene-editing predecessors via its unparalleled simplicity, efficacy in addition to being edconmical. In this review, Ahmad et al. [104], gave an overview of the CRISPR/Cas9 mode besides detailing genes that were edited with the utilization of this system for the treatment of breast cancer. In addition, they shed light on the administration approaches, involving both viral as well as and non-viral, whose utilization might be used to deliver the system and the barriers associated with each. Overall, the present review give new insights into the potential therapeutic applications of CRISPR/Cas9 for the advancement of breast cancer treatment [105] (Figure 5).
F. Auto immune as well as inflammatory diseases & CRISPR/Cas9
The broad range of rheumatic diseases vary from occasional monogenic auto inflammatory diseases, besides the complicated autoimmune diseases [106]. The monogenic auto inflammatory diseases possesses the properties of escalation of IL1. Usually some of these syndromes occur secondary to mutations in the NLRP3 gene with autosomal inheritance as well as differing penetrance [106]. In the complicated polygenic autoimmune diseases like rheumatoid arthritis (RA) a lot of genic factors, besides environmental triggers are implicated, like the heritability of RA is believed to be-65% [107]. A lot of these factors diseases are probably the clinical targets for CRISPR/Cas9, where this technique utilization for in vivo as well as in vitro functional, genomics studies to find the single besides combination role of SNP that are isolated by correlated studies at the genomic level via the fast generation of cellular as well as animal models. Besides that, this technique application can be done for the personalized therapy in the form of a tool for correction of mutations with regards to the approaches that have got adapted for every subject.
From this aspect utilization of CRISPR/Cas9 has got done for the generation of rat chondrosarcoma cell lines, which shows stable expression of Cas9 for this study of complicated crosstalk which is controlling function, differentiation of chondrocytes homeostasis, besides, evaluation of genes correlated with cartilage degeneration studies [108]. CRISPR/Cas9 possess the capacity of vitro, as well as in vivo, escalation of, functional demonstration in the part of the genes in the inflammatory, besides bone constituents of these diseases. From this angle CRISPR/Cas9 strategie’s application in a murine macrophage cell lines to illustrate that the RasGRP3 gene restricts the inflammatory reaction by activation of Rap1, which represented a small GTPase [109]. Another study illustrated that the miRNA-155 caused a proinflammatory reaction in addition to proosteoclastogenic action via the CRISPR/Cas9 stimulation of the miR155 binding site to the SHIP1 gene, that is a negative controller of inflammation [110]. Additionally, CRISPR/Cas9 technique might further integrate a cell therapy approach, like in case of rheumatoid arthritis (RA), a disease possessing the properties of impairment of reaction towards proinflammatory cytokines like IL1 as well as tumor necrosis factor alpha (TNFα), utilization of CRISPR/Cas9 was done with the idea of programming the murine stimulated iPSCs possessing the capacity of reaction towards an inflammatory stimulus with robust as well as autonomously controlled generation of the anti cytokines. Both TNFα as well as IL1 represent two of the maximum robust controllers of the CCL2 promoter gene expression. In case a gene that possesses antagonistic action to TNFα or IL1 is kept under the regulation of CCL2 promoter, it would react to cytokine amounts as well as result in a self controlled inflammatory event. This aids in the regulation of the cell expression by delivery of biological drugs as treatment modality [66].
G. Primary immunodeficiencyies as well as CRISPR/ Cas9
Primary immunodeficiency (PID) represents a group of heterogenous as well as occasional diseases, where part of the immune system work is impaired or does not possess any functional, capacity. There are various etiological genetic impairments of which over 230 have got isolated with a lot of different presentations. Of the different PID diseases, severe combined immunodeficiencies (SCID) that causes a blocking of T cell generation with an extra primary or secondary impairment in B cells, as well as natural killer (NK) cells, might or might not be influenced. The commonest kind of SCID is the X chromosomes correlated syndromes (X - SCID), that occurs secondary to mutations in the gene that get encoded by the γ receptor of IL2 (i.e. the IL2RGgene) [111].
PID that is inclusive of SCID can get treated by allogenic haematopoietic stem cells (HSC) transplantation. Nevertheless, histocompatibility issues might be the challenges, besides the chances of acquiring transmitted diseases [112]. For overcoming these issues, utilization of patients own HSCs via insertion of a copy of the functional, gene via a viral vector. Despite, the success visualized in clinical trials, robust complications might take place in view of viral infection that is intricately placed with oncogenes as well as hence their activation [113]. An alternative that offers great advantages is the utilization of homologous repair that possesses definite advantages over the classical gene therapy. In a different way in some immunodeficiencies, there occurs loss of function or gain of function in some genes that possesses a strict association with regulation of expression. The examples cited are the X-linked-agammaglobulinemia as well as the signal transducers and activators of transcription3 (STAT3) loss of function in addition to STAT3 along with STAT1 gain of function, or PI3K-δsyndrome. In ideal circumstances these diseases need the rectification of the existent gene for reinstatement of normal function or control that might get attained with CRISPR/Cas9 [114]. It is significant to possess knowledge that the effectiveness of treatment of gene editing is considerably different based on various factors on the cell kind, senescence status, as well as cell cvcle status of the target. Other parameters, which impact the effectiveness of treatment are cell aptitude, implying the possibility of arriving at a therapeutic modulation, threshold, besides the efficacy of administration of programmable nuclease system to the target tissue, that is believed to be efficacious just in case of programmable nuclease reaches with safety in addition to effectiveness to the nucleus of the target cell. Lastly the specificity of editing is one more significant parameter, that pointed to only editing the target DNA without influencing any more genes [115]. In view of haematopoietic cells being the commonest target in immunological diseases autologous haematopoietic stem cell transplantation is a good grafting procedure of cells that are corrected ex-vivo already by CRISPR/Cas9 that replaces total or partial compartment of the haematopoietic stem cell [114]. More recently, the CRISPR/Cas9 technique got applied for the generation of an in vitro model of Janus kinase deficiency. Elimination of T cell blocking generation was done with the utilization of CRISPR/ Cas9 modulated editing inion [116]. Chronic granulomatous disease (CGD) possesses the properties of robust in addition to continuous infection secondary to absence of an antipathogenic oxidative burst that gets conducted by the phagocytic cells to restrict besides remove bacterial as well as fungal growth. Utilization of CRISPR/Cas9 was done in induced pluripotent Stem Cells (iPSCs) that had been obtained, from a patient with CYBB for correction of a single mutation in the CYBB gene intron besides for resurrection of oxidative function in phagocytes [117]. Furthermore, utilization of CRISPR/Cas9 was done for removal of GT dinucleotide in the NCF1 Ggene, that encodes human p47phox protein in an acutemyeloid leukaemia cell line (PLB-985) for generation of a CGD cell model which acts as per the genetic background of the disease to be able to utilize them for preclinical vector evaluation[118]. Furthermore, utilization of CRISPR/Cas9 was done for the generation of multigene knockout models which had not been prone for effective genetic manipulation for the evaluation of immune system, like rabbits with the knocking of 1-5 genes simultaneously like (IL2RG, RAG1, RAG2, TIKII, as well as ALB) with effectiveness of treatment varying from 100% (for 1 gene) to 33.5% (for 5 genes) by microinjection into pronuclear stage embryos cytoplasm [119]. Rabbits possessing the knocking of IL2RG as well as RAG1, genes represent significant animal model of immunodeficiency that possess properties of lacking T, B as well as NK cells [119]. The rest of models implicate the golden syrian hamsters possessing the knocking of the modulates signal transducers and activators of transcription2 (STAT2)gene for evaluation of viral infections of the KCNQ1 gene for cardiovascular evaluation of the PP1R2C gene meant for transgenic migration [120]. Additionally, of B cell deficient pigs possessing the gene knockout of the gene that encodes the JH area of human immuogloulin (IgM)heavy chain that is key for the generation in addition to differentiation of B cells, caused the generation of piglets that had absence of antibody generating of B cells. The development of B cell deficient mutants represent the initial step in the generation of human antibody repositories in large animal models [121].
H. CRISPR/Cas9 as well as other immune diseases
Utilization of CRISPR/ Cas9 for the evaluation of allergic diseases approach is done for the evaluation of some genes like the MUC 18 gene, alias CD146 or melanoma cell adhesion molecule. Via the CRISPR/Cas9 the knockout of the MUC 18 gene was performed in primary human airway epithelial cells that represent s the initial defense against the environmental factors, like pathogens besides pollution. This mediated, this resulted in a reduction of reaction against IL 18 subsequent to the stimulation of the toll like receptor agonist that pointed that there is existence, of a part of MUC 18 gene in reaction to bacterial as well as viral stimulation [122]. Moreover, utilization of CRISPR/ Cas9 was done in X-linked hyper IgM syndrome for the rectification of the mutations of the CD40 ligand which addresses another use of CRISPR/ Cas9 [123]. Moreover, utilization of CRISPR/ Cas9 was done for inducing the recombination of IgH chain class alterations in the sub classes in which we want in murine as well as human B cells. Furthermore, its utilization was done for the generation of Fab fragments rather than the whole IgH molecule in case of mouse hybridoma cells with aim of greater scrutiny of Ig sub classes as well as innovative strategies for the accessible generation of Fab fragments for scientific as well as treatment utilization [124].
Monkeys represent a markedly significant animal models for the generation of treatment approaches in view of thei rintricate similarities with humans [125]. Successful utilization of CRISPR/ Cas9 for the development, of monkeys that possess the knockout of NROB1, PPAR-γ as well as RAG1 genes [126], that had the capacity to work as models of X-linked adrenal hyperplasia congenital as well as hypogonadotropic hypogonadism [127], lipodystrophy metabolism [128], insulin sensitivity, obesity in addition to inflammatory diseases [129]. Moreover, monkey knockout models of RAG gene possesses capacity of utilization in re generation medicine, allografts as well as xenografts transplantation, besides the experiments that are correlated with reconstitution that are linked to the immune system [130].
CRISPR/Cas9 as well as other diseases
Application of CRISPR/Cas9 system has been done in cellular as well as animal models for evaluation, besides looking for therapies for various neurological conditions like Parkinson’s diseases [131]. Mutations in the PARK2 or PINK1 genes resulted in early initiation of Parkinson’s diseases in the form of an autosomal recessive condition in humans [132]. The PARK2 gene encodes a protein known as parkin, that is a constituent of the multi protein complex E3 ubiquitin ligase, whereas the PINK1 gene encodes a for the PTEN induced putative kinase1 that is a mitochondrial serine /threonine kinase protein. Furthermore, utilization of CRISPR/ Cas9 has also been done in amyotrophic lateral sclerosis, huntington’s disease, schizophrenia as well as autism [133-135]. Furthermore, it has got utilized in movement diseases, like duchenne muscular dystrophy, in metabolic disease, like type1 Diabetes, hyper cholesteremia, in inherited diseases that influence vision, like retinitis pigmentosa [136-139], among lot of other diseases.
CRISPR/Cas9 System as well as Mitochondrial Genome Editing
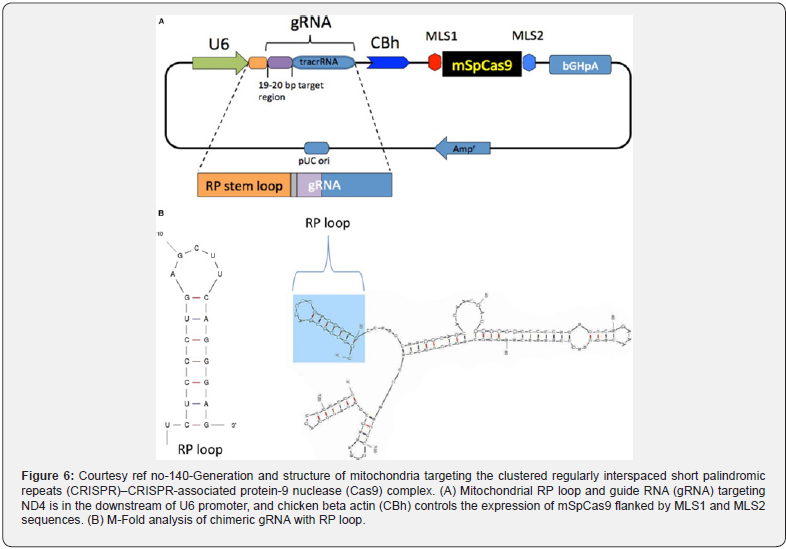
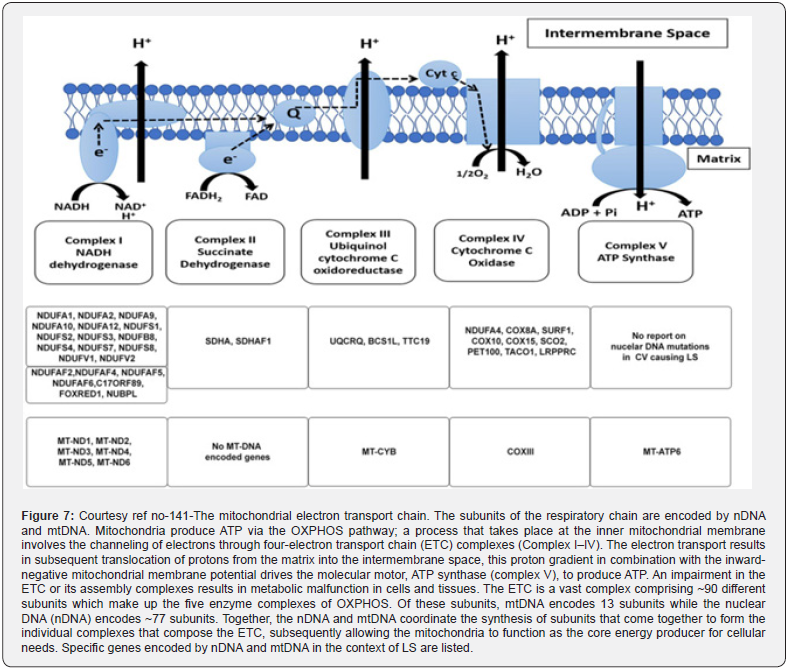
Gene editing of the mitochondrial genome using the CRISPR-Cas9 system is very tough, basically in view of the lower efficiency of delivering the guide RNA in addition to the Cas9 enzyme complexes into the mitochondria. In this study, Hussein et al. [140], possessed the capacity of carrying out gene editing in the mitochondrial DNA by appending an NADH-ubiquinone oxidoreductase chain 4 (ND4) targeting guide RNA to an RNA transport- obtained stem loop element (RP-loop) and expressing the Cas9 enzyme with a preceding mitochondrial localization sequence. They found a mitochondrial colocalization of RP-loop gRNA with a significant decrease of ND4 expression in the cells carrying a 11205G variant in their ND4 sequence coincidently causing a reduction of the mtDNA amounts. This proof-of-concept study pointed that a stem-loop element added sgRNA can be transferred to the mitochondria and functionally crosstalk with Cas9 to modulate sequence- particular mtDNA cleavage. With the utilization of this innovative strategy of targeting the mtDNA, their outcomes yield greater proof that the CRISPR-Cas9-mediated gene editing might probably be beneficial for the treatment mitochondrial-associated diseases [140] (Figure 6). Further Bakare et al. reviewed the role of mitochondrial genome using the CRISPR-Cas9 system in Leigh syndrome [141] (Figure7).
Conclusions
There is anticipation that CRISPR/Cas9 technique would soon get corroborated via ex vivo Clinical protocols that have already got initiated in humans to evaluate against different kinds of cancer. Meanwhile, it is totally applicable for experiments on cell therapies, awa further requirements, are to robustly generate them as a scientific tool in significantly variable biological systems. This would aid in the essential betterment for further biomedical applications. Thus avoidance of cancer generation secondary to onco-viruses, therapy for various haemoglobinopathies and as advances take further trying for therapy of various neurological awa immunodeficiency diseases. With greater experience mitochondrial genome using the CRISPR-Cas9 system might become feasibility in mitochondrial diseases.
References
- Im W, Mon J, Kim M (2016) Applications of CRISPR/Cas9 for Gene editing in hereditary movement disorders. J Mov Disord 9: 136-143.
- Rodriquez- Rodriquez D, Ramirez-Solis R, Garza-Elizondo MB, Garza- Rodriquez MD, Barrera-Saldana HA (2019) Genome editing: A perspective on the application of CRISPR/Cas9 to study human diseases (Review). Int J Mol Med 43: 1559-1574.
- Gaj T, Gersbach CA, Barbas CF III (2013) ZPN, TALEN, and CRISPR/Cas-based methods for Genome engineering. Trends Biotechnol 31: 397-405.
- Sander JD, Joung JK (2014) CRISPR/Cas systems for editing, regulating and targeting genomes. Nat Biotechnol 32: 347-55.
- Epinat JC, Arnould S, Chames P, Rochaix P, Desfontaines D, et al. (2003) A novel engineered mega nucleases induces homologous recombination in yeast and mammalian cells. Nucleic Acid Res 31: 2952-2962.
- Urnov FD, Rebar EJ, Holmes MC, Zhang HS, Gregory MD (2010) Genome editing with engineered zinc finger nucleases. Nat Rev Genet 11: 636-646.
- Miller JC, Tan S, Qiao G, Barlow K, Wang J, et al. (2011) ATALE nucleases architecture for efficient Genome editing. Nat Biotechnol 29: 143-148.
- Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, et al. (2012) A programmable dual RNA guided DNA endonuclease in adaptive bacterial immunity. Science 337: 816-821.
- Horvath P, Barrangou R (2010) CRISPR/Cas-the immune system of bacteria and archaea. Science 327: 167-170.
- Singh V, Braddick D, Dhar PK (2017) Exploring the potential of Genome editing in CRISPR/Cas technology. Gene 599: 1-18.
- Mojica FJ, Ferrer C, Juez G, Rodriguez-Valera F (1995) Long stretches of short tandem repeats are present in the largest replicons of the Archaea Haloferax mediterranei and Haloferax volcanii and could be involved in replicon partitioning. Mol Microbiol 17: 85-93.
- Richle MM, Bennett AF, Long AD (2001) Genetic architecture of Escherichia Coli. Proc Natl Acad Sci USA 98: 525-530.
- De Boy RT, Mongolia EF, Emerson JB, Nelson KE (2006) Chromosomal evolution in Thermotogales: large scale inversions and strain diversification of CRISPR sequences. J Bacteriol 188: 2364-2374.
- Makarova KS, Aravind L, Grishin NV, Rogozin IB, Koonin EV (2002) A DNA repair system specific for thermophilic Archaea and bacteria predicted by Genomic context analysis. Nucleic Acid Res 30: 482-496.
- Ishimo Y, Shingawa H, Makino K, Amemura M, Nakata A (1987) Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversions in Escherichia Coli and identification of the gene product. J Bacteriol 169: 54329-54333.
- Mojica FJ, Diez-Villasenor C, Soria E, Juez G (2000) Biological significance of a family of regularly spaced repeats in the Genomes of Archaea, bacteria and mitochondria. Mol Microbiol 36: 244-246.
- Jansen R, Embden JD, Gaastra W, Schouls LM (2002) Identification of genes that are associated with DNA repeats in prokaryotes. Mol Microbiol 43: 1565-1575.
- Mojica FJ, Diez-Villasenor C, Garcia-Martinez J, Soria E (2005) Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. J Mol Evol 60: 174-182.
- Barrangou R, FremauxC, Devau H, Richards M, Boyaval P, et al. (2007) CRISPR provides acquired resistance against viruses in prokaryotes. Science 315: 1709-1712.
- Brouns SJ, Jore MM, Lundgren MM, Weistra ER, Slijkhuis RJ, et al. (2008) Small CRISPR- RNAs guide antiviral defense in prokaryotes. Science 321: 960-964.
- Garneau JE, Dupuis ME, Villion M, Romero DA, Barrangou R, et al. (2010) The CRISPR/Cas bacterial immune system cleaves bacterio phage and plasmid DNA. Nature 468: 67-71.
- Deltcheva E, Chylinski K, Sharma CM, Gonzalez K, Chao Y, et al. (2011) CRISPR RNA maturation by small RNA and host factor trans-encoded small RNase III. Nature 471: 602-607.
- Sapranuskas R, Gasiunas G, Fremaux C, Barrangou R, Horvath P, et al. (2011) The Streptococcus thermophilus CRISPR/Cas system provides immunity in Escheriria Coli. Nucleic Acid Res 39: 9275-9282.
- Mali P, Yang L, Esvelt KM, Aach J, Guell M, et al. (2013) RNA guided human Genomic engineering via Cas9. Science 339: 819-823.
- Cyranoski D (2016) Chinese scientists topioneer first human CRISPR trial. Nature 535: 479-479.
- Wright AV, Nunez JK, Doudna JA (2016) Biology and applications of CRISPR systems: harnessing nature’s toolbox for Genomic engineering. Cell 164: 29-44.
- Zhang F, Wen Y, Guo X (2014) CRISPR/Cas for genome editing: progress, implications and challenges. Hum Mol Genet 23: R40-R46.
- Canver MC, Bauer DE, Orkin SH (2017) Functional interrogation of non-coding through CRISPR genome editing. Methods 121-122: 118-129.
- Gasiunas G, Barrangou R, Horvath P, Siksnys V (2012) Cas 9 ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc Natl Acad Sci USA 109: E2579-E2580.
- Ran FA, Cong L, Yan WX, Scott DA, Gootenberg JS, et al. (2015) In vivo Genome editing using Staphylococcus aureus Cas 9. Nature 520: 186-191.
- Hou Z, Zhang Y, Propson NE, Howden SE, Chu LF, Sontheimer EJ, et al. (2013) Efficient Genome engineering in human pluripotent stem cells using Cas 9 from Neisseria meningitides. Proc Natl Acad Sci USA 110: 15644-15649.
- Chen F, Ding X, Feng Y, Seebeck T, Jiang Y, et al. (2017) Targeted activation of diverse CRISPR/Cas systems for mammalian genome editing via proximal CRISPR targeting. Nat Commun 8: 14958.
- Murovec J, Pric Z, Yang B (2017) New variants of CRISPR RNA genome editing enzymes. Plant Biotechnol J 15: 917-926.
- Oliveros JC, Franch M, Tabas-Madrid D, San Leon D, Montoliu L, et al. (2016) Breaking -Cas-interactive design of guide RNAs for CRISPR/Cas experiments for ENSEMBL genomes. Nucleic Acid Res 44: W267-W271.
- Kaur K, Gupta AK, Rajput A, Kumar M (2016) ge CRISPR-an integrated pipeline for the prediction and analysis of sg RNAs genome editing efficiency for CRISPR/Cas systems. Sci Rep 6: 30870.
- Naito Y, Hino K, Bono H, Ui-Tei K (2015) CRISPR-direct: software for designing CRISPR/Cas as guide RNAs with reduced off- target sites. Bioinformatics 32: 1120-1123.
- Heigwer F, Kerr G, Boutros ME (2014) E- CRISP: Fast CRISPR target site identification. Nat Methods 11(2): 122-123.
- Wong N, Liu W, Wang X (2015) WU- CRISPR: characteristics of, functional guide RNAs for CRISPR/Cas, system. Genome Biol 16: 218.
- Doench JG, Hartenian E, Graham DB, Tutova Z, Hegde M, et al. (2014) Rational design of highly significant sgRNA for CRISPR/Cas mediated gene inactivation. Nat Biotechnol 32: 1262-1267.
- Chari R, Mali P, Moosburner M, Church GM (2017) sgRNA Scorer 2.0: a species independent model to predict CRISPR/Cas activity. ACS Synth Biol 6: 902-904.
- Moreno-Mateos MA, VeJinar CE, Beaudoin JD, Fernandez JP, Mis EK, et al. (2015) Rational CRISPR-scan: Designing highly efficient sgRNA for CRISPR/Cas targeting in vivo. Nat Methods 12: 982-988.
- Liu H, Wei Z, Dominguez A, Li Y, Wang X, et al. (2015) CRISPR-ERA: A Comprehensive design toolfor CRISPR mediated genome editing, repressionand activation. Bioinformatics 31: 3676-3678.
- Stemmer M, Thumberger T, DelSolKeyer M, Wittbrodt G, Mato JL (2015) CCtop: An intuitive, flexible, and reliable CRISPR/Cas prediction tool. PLoSOne 10: e0124633.
- Haeussler M, Schonig K, Eckert H, Eschstruth A, Mianne J, et al. (2016) Evaluation of off- target and on target scoring algorithms and integration into the guide RNA selection tool CRISPOR. Genome Biol 17: 148.
- Montagiue TG, Cruz JM, Gagnon JA, Church GM, Valen E (2014) CHOP- CHOP: A CRISPR/Cas 9 and TALEN web tool for genome editing. Nucleic Acid Res 42: W401-W407.
- Prykhozhij SV, Rajan V, Gaston D, Berman JN (2015) CRISPR-Multi target: A web tool to find common and unique CRISPR single guide RNA targets in asset of similar sequences. PLoSOne 10: e0119372.
- O’Brien A, Bailey TL (2014) GT- Scan: identifying unique genomic targets. Bioinformatics 30: 2673-2675.
- Hsu PD, Scott DA, Weinstein JA, Ran FA, Konermann S, et al. (2013) DNA targeting specificity of RNA guided Cas 9 nucleases. Nat Biotechnol 31: 1827-1832.
- Park J, Bae S, Kim JS (2015) Cas-Designer: A web basedtool for choice of CRISPR/Cas9 target sites. Bioinformatics 31: 4014-4016.
- Bae S, Park J, Kim JS (2014) Cas-OFFinder: A fast and versa-title algorithm that searches for potential off- target sites of Cas9 RNA guided endoucleases. Bioinformatics 30: 1473-1475.
- Cradicvk TJ, Qiu P, Lee CM, Fine EJ, Bao G (2014) COSMID: A web based tool for identifying and validating CRISPR/Cas9 offtarget sites. Mol Ther Nucleic Acids 3: e214.
- Hough SH, Kancleris K, Brody L, Humphryes-Kirilov N, Wolansky J, et al. (2017) Guide picker is a comprehensive design tool for visualizing and selective guides for experiments. BMC Bioinformatics 18: 167.
- Guell M, Yang L, Church GM (2014) Genome editing assessment using CRISPR- genome analyzer (CRISPR-GA). Bioinformatics 30: 2968-2970.
- Cao J, Wu L, Zhang SM, Lu M, Cheung WM, et al. (2016) An easy and efficient inducible CRISPR/Cas9 platform with improved specificity for multiple gene targeting. Nucleic Acid Res 44: e149.
- Ran FA, Hsu PD, Lin CY, Gootenberg JS, Konermann S, et al. (2013) Double nicking by RNA guided CRISPR/Cas9 for enhancing genome editing specificity. Cell 154: 1380-1389.
- Sato T, Sakuma T, Yokonishi T, Katagiri K, Kamimura S, et al. (2015) Genome editing in mouse Spermatogonial stem cell lines using TALEN and Double nicking CRISPR/Cas9. Stem Cell Reports 5: 75-82.
- Slaymaker IM, Gao I, Zetsche B, Scott DA, Yan WX, et al. (2016) Rationally engineered, Cas9 nucleases with improved specificity. Science. 351: 84-88.
- Tsai SQ, Wyvekens N, Khayter C, Foden JA, Thapar V, et al. (2014) Dimeric CRISPR RNA- guided Fok1- nucleases for highly specific Genome editing. Nat Biotechnol 32: 569-576.
- Wright AV, Sternberg SH, Taylor DW, Staahl BT, Bardales JA, et al. (2015) Rational design of a split- Cas 9 enzyme complex. Proc Natl Acad Sci USA 112: 2984-2989.
- Komor AC, Kim YB, Packer MS, Zuris JA, Liu DR (2016) Programmable editing of a target base in genomic without double stranded DNA cleavage. Nature 533: 420-424.
- Cox DB, Gootenberg JS, Abudayyeh OO, Franklin B, Kellner MJ, et al. (2017) RNA editing with CRISPR/Cas13. Science 358: 1019-1027.
- Konermann S, Brigham MD, Trevino AE, Joung J, Abudayyeh OO, et al. (2015) Genome scale transcriptional activation by an engineered CRISPR/Cas9 complex. Nature 517: 583-588.
- Qin P, Parlak M, Kuscu C, Bandaria J, Mir M, et al. (2017) Live cell imaging of low and non repetitive chromosome loci using CRISPR/Cas9. Nat Commun 8: 14275.
- Panowicz FP, Barzi M, Legras X, Hubert L, Mi T, et al. (2016) Reprogramming metabolic pathways in vivo with CRISPR/Cas9 Genome editing to treat hereditary tyrosinaemia. NatCommun 7: 12642.
- Kang H, Minder P, Park MA, Mesquitta WT, Torbett BE, et al. (2015) CCR5 disruption in Induced pluripotent stem cells using CRISPR/Cas9 provides selective resistance of immune cells to CCR5 tropic HIV-1 virus. Mol Ther Nucleic Acids 4: e268.
- Brunger JM, Zutschi A, Willard VP, Gerssbach CA, Guilac F (2017) Genome engineering of stem cells for autonomously regulated closed loop delivery of biological drugs. Stem Cell Reports 8: 1202-1213.
- Lin SR, Yang HC, Kuo YT, Liu CJ, Yang TY, et al. (2014) The CRISPR/Cas9 system facilitates clearance of the intrahepatic HBV template in vivo. Mol Ther Nucleic Acids 3: e186.
- Zhen S, Hua L, Takahashi Y, Narita S, Liu YH, et al. In vitro and in vivo growth suppression of human papilloma virus16 positive cervical cancer cells by CRISPR/Cas9. Biochem Biophys Res Commun 450: 1422-1426.
- Zhen S, Lu JJ, Wang LJ, Sun XM, Zhang JQ, et al. In vitro and in vivo synergistic therapeutic effects of cisplatin with human papilloma virus16 E6/ E7 CRISPR/Cas9 on cervical cancer cell lines. Transl Oncol 9: 498-504.
- Das D, Smith M, Wang X, Morgan IM (2017) The deacetylase SIRT1 regulates the replication properties of human papilloma virus 16 E1 and E2. J Virol 91: e0012-e00117.
- Merling R, Kuhns D, Sweeney C, Wu X, Burkett S, et al. (2016) Gene edited psedogene resurrect ion corrects p47 phox deficient chronic granulomatous disease. Blood Adv 1: 270-278.
- Paquet D, Kwart D, Chen A, Sprout A, Jacob S, et al. Efficient introduction of specific homozygous and heterogenous mutations using CRISPR/Cas9. Nature 533: 125-129.
- Schwank G, Koo BK, Sasselli V, Dekkers JF, Heo I, et al. (2013) Functional repair of CTFR by CRISPR/Cas9 in intestinal stem cells organoids of cystic fibrosis patients. Cell Stem Cell 13: 653-658.
- Gui H, Schreimer D, Cheng WW, Chauhan RK, Antinolo G, et al. (2017) Whole-exome sequencing coupled with unbiased functional analysis reveals low Hirshsprung disease genes. Genome Biol 18: 48.
- Halim D, Wilson MP, Oliver D, Brosens E, Verheij JB, et al. (2017) Loss of LMOD1 impairs smooth muscle-cyto- contractility and causes megacystis-microcolon-intestinal hyperperistalsis syndrome in humans and mice. Proc Natl Acad Sci USA 114: E2739-E2747.
- Ou Z, Niu X, He W, Chen Y, Song B, et al. (2016) The combination of CRISPR/Cas9 and iPSC technology in the gene therapy of ß thallasaemia in mice. Sci Rep 6: 32463.
- Xu P, Tong Y, Wang TT, Cheng L, Wang BY, et al. (2015) Both TALENs and CRISPR/Cas9 directly target the HBBIVS2-654(C>T) mutation in ß thallasaemia -derived iPSC. Sci Rep 5: 12065.
- Traxler EA, Yao Y, Wang YD, Woodard KJ, Kurita R, et al. (2016) A genome editing strategy to treat ß-haemoglobinopathies that recapitulates a mutation associated with a benign genetic condition. Nat Med 22: 987-990.
- Canver MC, Smith EC, Sher F, Pinello L, Sanjana NE, et al. (2015) BCL11A enhancer dissection by Cas9-mediated in situ saturating mutagenesis. Nature 527: 192-197.
- Zhang N, Zhi H, Curtis BR, Rao S, Jobaliya C (2016) CRISPR/Cas9 mediated conversion of human plateletsallo antigen allotypes. Blood 127: 675-680.
- Osborn MJ, Gabriel R, Webber BR, Defeo AP, Mc Elroy AN, et al. (2015) Fanconianaemia gene editing by the CRISPR/Cas9 system. Hum Gene Ther 26: 1114-1126.
- Zhang H, Mc Cartney N (2016) CRISPR/Cas9 technology and its application in haematological disorder. BrJ Haematol 175: 208-225.
- Hai T, Teng F, Guo R, Li W, Zhou X (2014) Onestep generation of knockouts of pigs by zygote injection of CRISPR/Cas9 system. Cell Res 24: 372-375.
- Puschnik AS, Majzoub K, Ooi YS, Carette JE (2017) A CRISPR toolbox to study virus host interaction. Nat Rev Microbiol 15: 351-364.
- Kennedy EM, Kornepati AV, Goldstein M, Bogerd HP, Poling BC, et al. (2014) Inactivation of the human papilloma virus E6 or E7 gene in cervical carcinoma cell lines by using a bacterial CRISPR/Cas9 RNA guided endonuclease. J Virol 88: 11965-11972.
- Wang W, Ye C, Liu J, Zhang D, Kimata JT, et al. (2014) CCR5 gene disruption via lentiviral vectors expressing Cas9 and a single guided RNA renders cells resistant to HIV-1 infection. PLoSOne 9: e115987.
- Zhen S, Hua L, Liu YH, Gao LC, Fu J, et al. (2015) Harnessing the Clustered regularly Interspersed short Palindromic Repeats (CRISPR) CRISPR- associated Cas9 system to disrupt the Hepatitis B virus. Gene Ther 22: 404-412.
- Karimova M, Beschorner N, Dammermann W, Chemintz J, Inderbirken D, et al. (2015) CRISPR/Cas9 nickase-mediated disruption of Hepatitis B virus openreading frame S and X. Sci Rep 5: 13734.
- Xu Y, Qi Y, Lou J, Yang J, Xie Q, et al. (2017) Hepatitis B virus X protein stimulates proliferation, wound closure and inhibits the apoptosis of Huh7 cells via CDC42. Int J Mol Sci 18: 3E586.
- Ren Q, Li C, Yuan P, Cai C, Zhang L, et al. (2015) A dual-reporter system for real time monitoring and high throughput CRISPR/Cas9 library screening of the hepatitis C virus. Sci Rep 5: 8865.
- Yuen KS, Chan CP, Wong NH, Ho CH, Ho TH, et al. (2015) CRISPR/Cas9 mediated genome editing of Epstein Barr virus in human cells. J Gen Virol 96: 626-636.
- Wang J, Quake SR (2014) RNA guided endonuclease provides a therapeutic strategy to cure latent herpes viridae infection. Proc Natl Acad Sci USA 111: 13157-13162.
- Araujode Almeida DM, de Almeida Ribeiros CR, Vasquez Raposo J, de Paula VS (2021) Immunotherapy and gene therapy for oncoviruses infections. Viruses 13: 822.
- Moranghuino I, Valente ST (2020) Block and lock: New horizons for cure for HIV. Viruses 12(12): 1443.
- Kristler KE, Vosshall LB, Mathews BJ (2015) Genome engineering with CRISPR/Cas9 in the mosquito Aedes aegyptii. Cell Rep 11: 51-60.
- Zhang C, Xiao B, Jiang Y, Zhao Y, Li Z, et al. (2014) Efficient editing of malaria parasite Genome using CRISPR/Cas9 system. M Bio 5: e01414-e01414.
- Burt A (2003) Site specific selfish genes as tools of for the control and genetic engineering of, natural populations. Proc Biol Sci 270: 921-928.
- Webber BL, Raghu S, Edward OR (2015) Opinion-Is CRISPR based gene drive a biocontrol silver bullet or global conservation threat? Proc Natl Acad Sci USA 112: 10565-10567.
- Hammond A, Galizi R, Kyrou K, Simoni A, Siniscalchi C, et al. (2016) A CRISPR/Cas9 gene drive system targeting female reproduction in the malaria mosquito vector Anopheles gambiae. Nat Biotechnol 34: 78-83.
- Sanchez-Rivera FJ, Jacks T (2015) Application of the CRISPR/Cas9 system in cancer biology. Nat Rev Cancer 15: 387-395.
- Kawamura N, Nimura K, Nagano H, Yagamuchi S, Nanomura N, et al. (2015) CRISPR/Cas9- mediated gene knockout of NANOG and NANOGP8 decreases the malignant potential of prostate cancer cells. Oncotarget 6: 22361-22374.
- Garcia-Tunon I, Hernandez-Sanchez M, Ordonez JI, Alinso-Perez V, Alamo-Quijada M, et al. (2017) The CRISPR/Cas9 system efficiently reverts the tumorigenic ability of BCR/ABL in vitro and in a xenograft model of chronic myeloid leukaemia. Oncotarget 8: 26027-26040.
- Zuckermann M, Hovestadt V, Knobbe-Thomsen CB, Zapatka M, Northcott PA, et al. (2015) Somatic CRISPR/Cas9 mediated tumor suppressor disruption enables versatile brain tumors modeling. Nat Commun 6: 7391.
- Maddalo D, Manchado E, Concepcion CP, Bonetfi C, Vidigal JA, et al. (2014) In vivo engineering of Oncogenic chromosomal rearrangements with the CRISPR/Cas9 system. Nature 516: 423-427.
- Ahmad M, Hope Daoud G, Mohammad A, Harati R (2021) New insights into the therapeutic applications of CRISPR/Cas9 genome editing in breast Cancer. Genes 12(5): 723.
- Pathak S, Mc Dermot MF, Savic S (2017) Autoinflammatory diseases, update on classification, diagnosis and management. J Clin Pathol 70: 1-8.
- Kim K, Bang SY, Lee HS, Bae SC (2017) Update on the genetic architecture of rheumatoid arthritis. Nat Rev Rheumatol 13: 13-24.
- Yang M, Zhang L, Stevens J, Gibson G (2014) CRISPR/Cas9 mediated generation of stable chondrocytes cell lines with targeted gene knockouts: analysis of an aggrecan knockout cell line. Bone 5: 4657.
- Tang S, Chen T, Yu Z, Zhu X, Yang M, et al. (2014) Ras GRP3 limits toll like receptor –triggered response in macrophages by activating Rap1 small GTPase. Nat Commun 5: 46573.
- Jing W, Zhang X, Sun W, Hou X, Yau Z, et al. (2015) CRISPR/Cas9 mediated genome editing of miRNA-155 inhibits proinflammatory cytokines production by RAW264 7 cells. Biomed Res Int 7: 326042.
- Ott de Bruin LM, Volpi S, Musunuru K (2015) Novel genome editing tools to model and correct primary immunodeficiencies. Front Immunol 6: 250.
- Cowan MJ, Neven B, Cavazanna-Calvo M, Fischer A, Puck J (2008) Haematopoietic stem cells transplantation for severe combined immunodeficiencies. Biol Blood Marrow Transplant 14(1Suppl1): S73-S75.
- Kohn DB, Kuo CY (2017) New frontiers in the therapy of primary immunodeficiency: From gene addition to gene editing. J Allergy Clin Immunol 139: 726-732.
- Goodman MA, Moradi-Manesh D, Malik P, Rothenberg ME (2017) CRISPR/Cas9 in Allergic and immunological diseases. Expert Rev Clin Immunol 13: 5-9.
- Wang M, Glass ZA, Xu Q (2017) Non viral delivery of genome editing nucleases for gene therapy. Gene Ther 24: 144-150.
- Chang CW, Lai YS, Westin E, Khodadadi-Jamayran A, Pawlik KM, et al. (2015) Modeling human severe combined immunodeficiency and correction by CRISPR/Cas9- enhanced gene targeting. Cell Rep 12: 1668-1677.
- Flynn R, Grundmann A, Renz P, Hanzler W, James WS, et al. (2015) CRISPR mediated genomic and phenotypic correction of a chronic granulomatous disease mutation in human iPS cells. Exp Haematol 43: 838-848.
- Wrona D, Siler U, Reichenback I (2017) CRISPR/Cas9 generated p47 (phox) deficienct cell line for chronic granulomatous disease gene therapy vector development. Sci Rep 7: 44187.
- Yan Q, Zhang H, Yang H, Zou Q, Tang C, et al. (2014) Generation of multigene knockout rabbits using the Cas9/ gRNA system. Cell Regen (Lond) 3: 12.
- Fan Z, Li W, Lee SR, Meng Q, Shi B, et al. (2014) Efficient gene targeting in golden Syrian hamsters by CRISPR/Cas9 system. PLoSOne 9: e109755.
- Chen F, Wang Y, Yuan Y, Zhang W, Ren Z, et al. (2015) Generation of B cell deficienct pigs by highly efficient CRISPR/Cas9 mediated gene targeting. J Genet Genomics 42: 437-444.
- Chu HW, Rios C, Huang C, Wesolowska-Andersen A, Burchard EG, et al. (2015) CRISPR/Cas9 mediated gene knockout in primary human airway epithelial cells reveals a pro inflammatory role for MUC 18. Gene Ther 22: 822-829.
- Kuo CY, Hoban MD, Joglekar AV, Kohn DB (2015) Site specific gene correction of defects in CD40 ligand using the CRISPR/ Cas9 genome editing platforms. J Allergy Clin Immunol 135: A17.
- Cheong TC, Çompagno M, Chiarle R (2014) Editing of mouse and humanimmuogloulin genes by CRISPR/Cas9 system. NatCommun 7: 10934.
- Rodriguez-Sanchez P, Guindon J, Ruiz M, Tejero ME, Hubbard JE, et al. (2016) The edocannabinoid system in the baboon (Papio spp) as a complex framework for developmental pharmacology. Neurotoxicol Teratol 23: 30.
- Niu Y, Shen B, Cui Y, Chen Y, Wang J, et al.(2014) Generation of gene modified cynomolgous monkey via Cas9/ RNA- mediated gene targeting in one cell embryos. Cell 156: 836-843.
- Kang Y, Zheng B, Shen B, Chen Y, Wang J, et al. (2015) CRISPR/Cas9 mediated Dax1 knockout in the monkey recapitulates human AHC-HH. Hum Mol Genet 24: 7255-7264.
- Kim S, Huang LW, Snow KJ, Ablamunits V, Hasham MG, et al. (2007) A mouse model of conditioned lipodystrophy. Proc Natl Acad Sci USA 104: 16627-16632.
- Croasdell A, Duffney PF, Kim N, Lucy SH, Sime PJ, et al. (2015) PPARγ and the innate immune system mediate the resolution of inflammation. P PAR Res 2015: 5489691
- Huang J, Guo X, Fan N, Song J, Zhao B, et al. (2014) RAG1/2 pigs with severe combined immunodeficiency. J Immunol 193: 1496-1503.
- Chen Y, Xiong M, Dong Y, Habarman A, Cao J, et al. (2016) Chemical control of grafted human iPSC-derived neurons in a mouse model of Parkinson’s disease. Cell Stem Cells 18: 817-826.
- Zhou X, Xin J, Fan N, Zou X, Huang J, et al. (2015) Generation of CRISPR/Cas9 mediated gene targeted pigs via somatic cell unclear transfer. Cell Mol Life Sci 72: 1175-1184.
- Wang L, Yi F, Fu L, Yang J, Wang S, et al. (2017) CRISPR/Cas9 mediated targeted gene correction in amyotrophic lateral sclerosis patients iPSC. Protein Cell 8: 365-378.
- Merienne E, Vachey G, de Longprez L, Meunier C, Zimmer V, et al. (2017) The self-inactivating Kam/Cas9 system for the editing of CNS disease genes. Cell Rep 20: 2980-2991.
- Page SC, Hamersky GR, Gallo RA, Rannals MD, Calcaterra NE, et al. (2018) The schizophrenia and autism associated gene transcription factor 4 regulates the columnar distribution of layer2/3 prefrontal pyramidal neurons in an activity dependent manner. Mol Psychiatry 23: 304-315.
- Long C, Amoasii L, Mireault AA, Mc Anally JR, Li H, et al. (2016) Postnatal genome editing partially restores dystrophin expression in a mouse model of muscular dystrophy. Science 351: 400-403.
- Ackermann AM, Zhang J, Heller A, Briker A, Kaestner KH (2017) High fidelity Glucagon-Cre-ER mouse line generated by CRISPR/Cas9 assisted gene targeting. Mol Metab 6: 236-244.
- Jarrett KE, Lee CM, Yeh YH, Hsu RH, Gupta R, et al. (2017) Somatic genome editing with CRISPR/Cas9 generates and corrects a Metabolic disease. Sci Rep 7: 44624.
- Yu W, Mookherjee S, Chaitankar V, Hiriyaanna S, Kim JW, et al. (2017) Nrl knockdown by AAV-delivered CRISPR/Cas9 prevents retinal de generation in mice. Nat Commun 8: 14716.
- Hussein SR, Yalvac ME, Khoo B, Eckhart S, Mc Laughlin KJ (2021) Adapting CRISPR-Cas9 system for targeting mitochondrial genome. Front Genet 12: 627050.
- Bakare AB, Lesnefsky EJ, Iyer S (2021) Leigh syndrome: a tale of two genomes. Front Physiol 12: 693734.






























