Fermentative Kinetic Modeling for Protein Biomass from Sugar Diversification
Omar Pérez1, Juan E. Miño2, Agustín García1 and Erenio González1
1Department of Chemical Engineering, Marta Abreu Central University of Las Villas, Cuba
2Faculty of Engineering, National University of Misiones (UNAM), Argentina
Submission: February 26, 2019; Published: March 26, 2019
*Corresponding author: Omar Pérez, Department of Chemical Engineering. Marta Abreu Central University of Las Villas. Road to Camajuaní. Km 5 ½. Santa Clara, Cuba
How to cite this article: Omar P, Juan E. M, Agustín G, Erenio G. Fermentative Kinetic Modeling for Protein Biomass from Sugar Diversification. Curr Trends Biomedical Eng & Biosci. 2019; 18(5): 555998. DOI: 10.19080/CTBEB.2019.18.555998
Abstract
An analysis of the models proposed in the scientific literature is carried out to describe the kinetics of fermentation from a substrate conformed by cane juice and supplemented with final honey and nutrients that are inoculated with autochthonous yeasts of the medium. It is proposed the adjustment of the models through the interactive simulation of processes with minimum integral error of the sums of the differences between the experimental values and those predicted by the models. Initial total reducing sugars concentrations of 45 and 62 g/L were considered maintaining the pH between 4.4 and 5.2 and the temperature at 36 C. The best adjustment to the experimental values was obtained for the free inhibition model of Teisiser, while the Monod model presents deficiencies to describe the first and last part of the growth curve. The parameters of Teiissier at s0=45g/L are μm =0,21 and ks = 11,21 g/l while at s0 = 62 g/l are um=0,25h-1 and ks =11,21g/l. As a result of the simulation it was determined that there are no significant effects of inhibition by substrate or product and that the Malthos model of cell growth accurately describes this process.
Keywords: Kinetics; Models; Fermentation; Protein biomass; Sugar cane
Introduction
The production of protein biomass by fermentation of by-products of sugar cane treatment is low investment and energy requirements. For these reasons it is very attractive for sugar diversification and can be developed from intermediate juices of the sugar process, final honey and hydrolyzed bagasse [1,2]. When it uses effluent juice from the filters, it considerably increases the quality of the sugar and the energy efficiency in its manufacture. This material is rich in protein content and intended for animal feed, mainly pork, with adequate conversion rates and yield in weight of the animals [3]. When the process is submerged, the difficulties inherent to solid-state fermentation are avoided. However, a heterogeneous three-phase system is presented whose phenomenological description is complex but very necessary.
In order to apply adequate design and scaling methods of these bioreactors for this process, it is necessary to apply effective modeling of the behavior of these equipments. The global model of the bioreactor implies obtaining the heat and mass transfer models, the hydrodynamics of the equipment and especially the kinetic model of the microorganism population, in the selected culture conditions [4]. This kinetic model allows to determine the cell growth rate and the influence of external factors as well as the simulation and optimization of the operation of the equipment.
Materials and Methods
Kinetic models for protein biomass
This fermentation generates an amount of protein biomass (X) and ethanol associated with metabolism (P), through the consumption of an amount of total reducing sugars (S). The proposed models are based on the kinetics of enzymatic catalysed reactions. Starting from the specific growth rate of protein biomass (),μ Monod was proposed as the primary model, defined as:
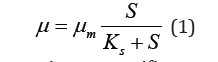
Where mμ is the maximum specific growth rate of biomass and KS is the growth limiting concentration S, for which u=um/2 is reached.
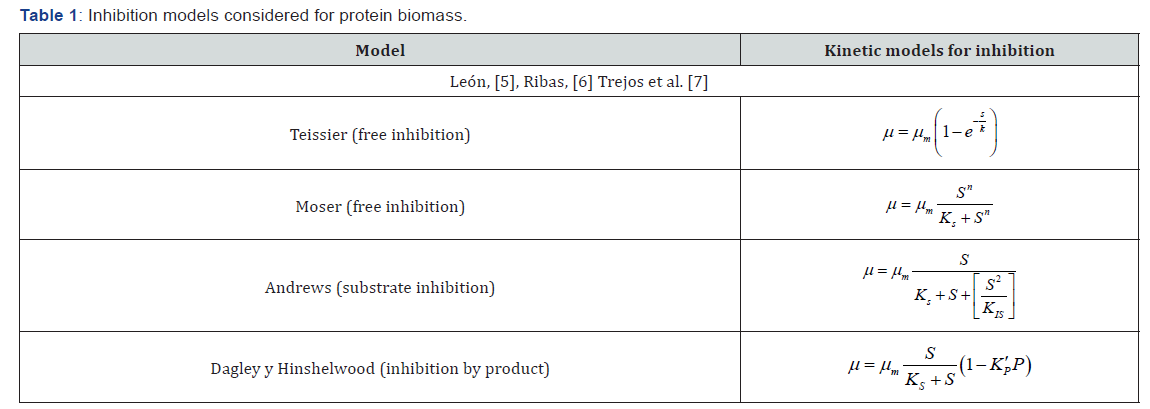
As the Monod model is not always adequate to describe this process, fermentative kinetic models were proposed, considering free inhibition, by substrate and by product. Table 1 shows the models considered for inhibition over .μ Table 2 shows the models considered for cell growth under discontinuous conditions [5-7].
Analytical and experimental procedure
The expression of velocity and the equation of cell growth best adapted to experimental discontinuous fermentation data for protein biomass were determined. For the adjustment, interactive simulation of processes using the Hooke and Jeeves method was applied in the optimization of the integral error results of the differences between the values calculated by the model and the experimental data, defined as:
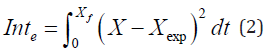
The differential equations were solved by Runge - Kutta numerical integration from fourth order to fixed step. The dynamic experimental behaviour of X, S and P at pH between 4,4 and 5,2 and 36 °C was determined. A substrate formed by 80 % of cane juice and 20 % of final honey was used. It was inoculated with autochthonous cane juice yeasts. It was fermented at initial substrate concentration ( ) 0 S of 45 and 62 g/L and aeration of 1 air -medium L L min. 1,27 g/L of ammonium phosphate and 2,36 g/L of urea were added as nutrients.
The cell, substrate and product (ethanol) concentration was determined everyone hour during 10 hours of discontinuous fermentation.
Results and Discussion
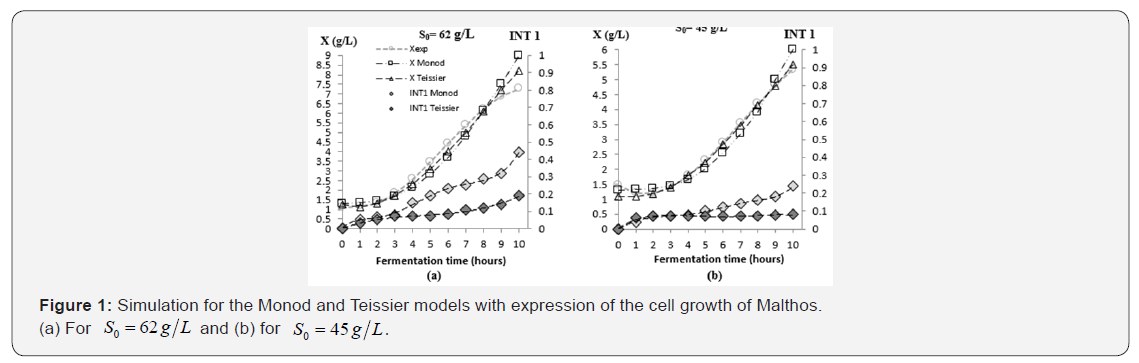

The simulation results are shown in Figure 1 a & b. In (a) to S 0= 45 g /L and in (b) to S0 = 62 g/ L. They considered: the Monod model according to equation 1, the Teisiser model according to Table 1, expression of the cell growth of Malthos according to Table 2 and expression of INT1 of equation 2.
The Monod model fits acceptably in the first part of the logarithmic phase but has limitations in the adaptation stage and in the last part of the logarithmic phase. The best fit is achieved with the Teissier model, also considering cell growth by the Malthos model. Table 3 summarizes the parameters and results of the adjustment to the experimental data at both 0S
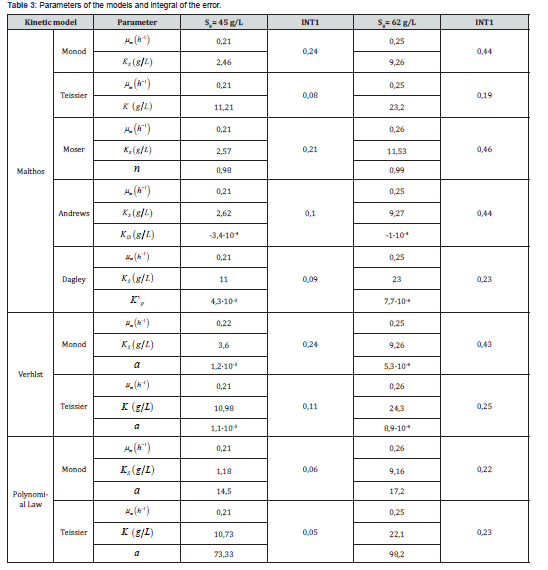
The Andrew model does not improve INT1 with respect to Teissier, while the order of the KIS value indicates the little influence of substrate inhibition. This is due to the fact that concentrations that had already been selected as suitable for this process in previous works were considered experimentally [8]. The Dagley model shows a good fit, but very small values of the inhibition constants per ’P K product. The Malthos model achieves similar results as the more complex solution models, such as Verhlst and Polynomial Law, because there are no important inhibition mechanisms in this process and the logarithmic phase predominates in the studied conditions.
Conclusion
Fermentative kinetics for protein biomass from a substrate made up of cane juice supplemented with final honey and nutrients using microorganisms native to the environment, is adequately described through Teissier and Malthos models of cell growth, resulting in the same:
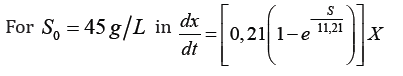
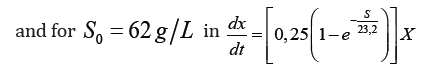
The effects of inhibition of cell growth by substrate and metabolism products are not noticeable. Expressions of cell growth, different from that of Malthos, would generate greater complexity in solving the differential equations, without achieving significant improvements in the fit of the models.
References
- Albernas Y (2013) Procedimiento para la síntesis y el diseño óptimo de plantas discontinuas de obtención de bioetanol empleando bagazo de caña de azúcar. Thesis in Option to the Scientific Degree of Doctor in Technical Sciences. Marta Abreu, Central University of Las Villas, Department of Chemical Engineering, Santa Clara.
- Mesa L (2010) Estrategia investigativa para la tecnología de obtención de etanol y coproductos del bagazo de la caña de azúcar. Thesis in Option to the Scientific Degree of Doctor in Technical Sciences, Marta Abreu, Central University of Las Villas, Department of Chemical Engineering, Santa Clara.
- García RA, Herrera CN, Marín CA, Días RI (1999) Producción de biomasa proteica a partir de jugo de caña. Respuestas. UFPS, Pp 32-34.
- Flores HAM, Flores MM, Reyes RJ, Astorga Z (2013) Diseño y modelado de un bioreactor tipo batch y continuo para aplicaciones de control automático. Congreso Nacional de Control Automático 2013. Ensenada, Baja California, Mexico, Octubre 1: 16-18.
- León MA (1999) Desarrollo de Modelos Cinéticos para Bioprocesos: Aplicación a la Producción de Xantano. MEMORIA para optar al GRADO de DOCTOR. Madrid, Spain.
- Ribas M, Hurtado RN, Díaz de los Ríos M, Doménech F, Sabadí R, et al. (2006) Modelación matemática y simulación de procesos fermentativos. Ingeniería química Nº. 438, Pp. 118-125.
- Trejos VM, Fontalvo AJ, Gٌ ómez G (2009) Descripción Matemática Y Análisis De Estabilidad De Procesos Fermentativos. DYNA, 76(158): 111-121.
- García RA, Herrera CN, Marín CA, Gómez RL (2005) Tecnología simplificada de producción de biomasa proteica. Centro Azúcar 32(1): 57-61.






























