Applying Social Network Analysis to Understand the Percentages of Keywords within Abstracts of Journals: A System Review of Three Journals
Tsair Wei Chien1,2, Yang Shao3 and Willy Chou4,5*
1Research Departments, Chi-Mei Medical Center, Taiwan
2Department of Hospital and Health Care Administration, Chia-Nan University of Pharmacy and Science, Tainan, Taiwan
3Department of Electronics and Information Engineering, Tongji Zhejiang College, China
4Department of Sports Management, Chia Nan University of Pharmacy and Science, Taiwan
5Rehabilitation Department, Chi-Mei Medical Center, Taiwan
Submission: May 02, 2018; Published: July 02, 2018
*Corresponding author: Willy Chou, Department of Sports Management, College of Leisure and Recreation Management, Chia Nan University of Pharmacy and Science, Tainan, Taiwan, Email: willyufan0101@ms22.hinet.net
How to cite this article: Tsair Wei Chien, Yang Shao, Willy Chou. Applying Social Network Analysis to Understand the Percentages of Keywords within Abstracts of Journals: A System Review of Three Journals. Curr Trends Biomedical Eng & Biosci. 2018; 16(1): 555926. DOI: 10.19080/CTBEB.2018.16.555926.
Abstract
Background:Academic literature suggests keywords that are retrieved from a paper’s title and abstract represent important concepts in that study. The percentage of keywords within an abstract (PKWA) is required to investigate.
Objective:To compare the PKWA in journals of medical informatics and the keyword network relationship in order to develop a self-examining policy for the journal.
Methods:Selecting 5,985 abstracts and their corresponding keywords in three journals (JMIR, JAMIA, and BMC Med Inform Decis Mak.) published between 1995 to 2017(April) on the US National Library of Medicine National Institutes of Health (Pubmed.org), we computed the PKWA for each journal by using MS Excel modules and compared the percentage differences across journals and years via a two-way ANOVA. Social Network Analysis (SNA) was performed to explore the relations of keywords in journals.
Results:The PKWA are 48.81, 41.59, and 56.84 for the three journals, respectively. A statistically significant difference (p<0.05) is found in the percentages among journals selected. In contrast, no differences (p>0.05) are found (1) between years (2016 and 2017) and (2) in interaction effects between journals and years. Three journals display significantly different patterns in network keywords and major cohesion measures.
Conclusion:It is required to apply the computer module when inspecting whether keywords are within abstracts. The cohesion measure provides journal editors with a method of examining keywords within an abstract for a paper under review.
Keywords: Cohesion measure; Journal; Social network analysis; Abstract
Background
Authors are required to provide three to ten keywords that represent the main content of the article when submitting it to a journal [1-5]. Keywords or short phrases published with an abstract can assist indexers in cross-indexing the article. However, few studies have investigated whether keywords are substantially associated with the abstract and what percentages of keywords truly exist in the corresponding abstract.
Meanwhile, we have seen some computer scientists placing high hopes in machine-learning algorithms, data mining and artificial intelligence. All of these methods are based on recently developed technologies of Natural Language Processing (NLP) and Text Prediction to process natural spoken language, to read unstructured data in Big Biomedical Data (BBD), to comprehend the intent of physicians, to quantify research information, and even to create a structured database [6-16]. Furthermore, informal patient data on the Web is increasing, accessible, inexpensive, available in real-time, and seems likely to cover a significant proportion of the population. Accordingly, extracting the intent of authors from unstructured journal papers may be possible and reachable in the near future. The keywords suitable for use in an index should be examined on the matter.
In literature, keywords retrieved from a paper’s title and abstract as important words to a study can help readers to find the article. We expect keywords are specific enough to represent the manuscript content. To answer whether each keyword appears in the accompanying abstract requires analysis. The Percentage of Keywords (PKW) within an abstract for a paper can be used to compare journals.
In search of keywords “internet OR Internet” to Pub-med on 2017/04/24, we have seen 84,069 published papers, in which 2,073 articles are subject to J Med Internet Res. What keyword in papers is most closely associated with “internet” is still unknown. An apocryphal story often told to illustrate data mining concepts is about beer and diaper sales,which were strongly correlated [17-19]. We are interested in using Social Network Analysis (SNA) [20-22] to analyze keywords related to a journal’s aims and scope as some studies reporting co-authorship relations within and between papers [23-25].
The SNA approach [26-28] is used to define facilities as the “nodes” of a keyword network connecting to another node (e.g., a square box) with a relation represented as an edge (e.g., an arrow line) [20,24]. For instance, a string of 4 3 5 denotes that the keyword 4 associated with another keyword 3 accounts for 5 times (with a weight 5) within a specific period, displayed graphically as γ4γ3. Several algorithms and measures have been applied to SNA. When the aim is to investigate the status of an actor in the network, the centrality measures should be applied [24]. This means that an actor is analyzed generally by its centrality [29,30].
In this study, we selected three journals (i.e., J Med Internet Res. [JMIR], J Am Med Inform Assoc. [JAMIA], and BMC Med Inform DecisMak. [MIDM]) from the category of medical informatics to compare their PKWA and their centrality (which takes into account three measures of Degree centrality, Closeness centrality and between’s centrality for the published papers in journals).
Our aims are to
a) Compare the PKWA among journals.
b) Show the pattern of a journal according to the keywords’ association and compute the macro cohesion measure.
c) Apply SNA to identify whether author’s papers target the journal’s scopes and aims according to the minor cohesion measure of the journal.
d) Evaluate the equality of centrality for a journal using Ferguson’s delta coefficient [31-34].
Methods
Data sources
Selecting 5,985 abstracts and their corresponding keywords in three journals (JMIR, JAMIA, and MIDM) published between 1995 and 2017 (April 15th) from the US National Library of Medicine National Institutes of Health (Pubmed.com), we computed the PKWA by using an Excel module made by the author. All papers with one keyword or more are included. We can see in Multimedia 1 showing both JMIR and JAMIA start from 2013 and MIDM from 2016 instead. The Figure 1 presents the study flowchart.

Compare the PKWs among journals
We demonstrated two ways to show each journal’s PKWA: (i) the MML (Method of Maximum Likelihood) [35] with a diagram comparison and (ii) the mean comparison using a two-way ANOVA across journals and years. The former was employed to select the maximal count number determined as the PKWA across all possible PKWAs (from 0 to 1.0 by an interval of 0.1 and the nil representing no keyword in an article) for each journal. The latter was used to compute the total count within the respective abstract over the total count across all abstracts for a specific journal in 2016 and 2017, due to the minimal overlapping years being limited to the MIDM PKW that were available, Percentages of key words within an abstract across years and journals.
Pattern of a journal’s keywords
To select two keywords with the strongest association for ease of display, i.e., with a large number of counts simultaneously listed in an article, we extracted the top 100 pairs with the highest linkage count using Pajek SNA software [22] to draw the visualized representations. The wider and darker linkage line between two keywords (i.e., called the edge between nodes in SNA) is shown, the stronger the association will be. The larger bubble represents the higher probability of a keyword’s occurrence in the journal. Any node with an identical color means it belongs to a similar category of the keyword occurrence number. We chose the weighted degree centrality measure to draw the keyword pattern and selected the separate component algorithm to plot the drawing. For detailed information, interested readers are advised to refer Extracting data using an author-made MS Excel module http://www.healthup.org.tw/marketing/course/marketing/ JMIR_abstract.mp4
Cohesion measure to examine papers’ targeting of a journal’s scope
There are three centrality measures usually applied to SNA [24]:
1. Degree centrality of a node is defined as the total number of edges that are adjacent to this node. This measures how many linkages directly connect keywords to their neighbours in the network. Closeness centrality focuses on how close an actor is to all other actors. It is measured as a function of mean geodesic/shortest distances [36].
2. Closeness centrality thus extends the description of degree centrality with a focus on that a keyword is relatively most close to all the other authors.
3. Between’s centrality expresses an operationalization of centrality on the basis of specifying how often a node is found on the shortest route between each pair of nodes in the network.
Due to different scaling scores across all three measures, we standardized them following ~N(0,1). The cohesion measure for examining the extent of any paper targeting a journal’s scope is obtained by averaging the above mentioned three standardized centrality measures. A higher cohesion measure means a stronger keyword association with the journal’s features. For detailed information, interested readers are recommended to consult Computing major cohesion measure for a journal using Pajek SNA software. http://www.healthup.org.tw/marketing/course/ marketing/jmir_pajek.mp4
Ferguson’s delta coefficient to evaluate the equality of centrality to a journal
Ferguson’s delta [31-34] is an index of discrimination measured by the proportion of discriminations (i.e., the degree of uniform distribution). It is reported that a normal distribution would be expected to have a discrimination of delta>0.90. We applied it to examine whether journals have an identical delta coefficient. A higher value means a more uniform distribution among the journal papers in cohesion measures.
Results
Comparison of the PKWs among journals
Summarizing data from Multimedia 1, we examined the top point on the line chart foreach journal in Figure 2 (i.e., 30% for JMIR, 80% for MIDM, and nil for JAMIA) and found that JAMIA has many articles without any keyword in this period from 2013 to 2017(April). If ignoring the nil portion (e.g., non-research articles such as perspectives, reviews, editorials, etc), JAMIA’s PKW is 30% equal to JMIR using the MML approach.

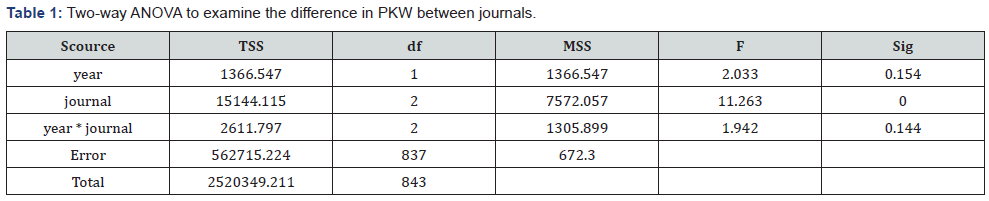
From Table 1, we can see that a significant difference exists among the journals, but there is no difference between years (i.e., year 2016 and 2017). The PKWA means are 56.84% (MIDM), 48.82 (JMIR), and 41.59(JAMIA), respectively.
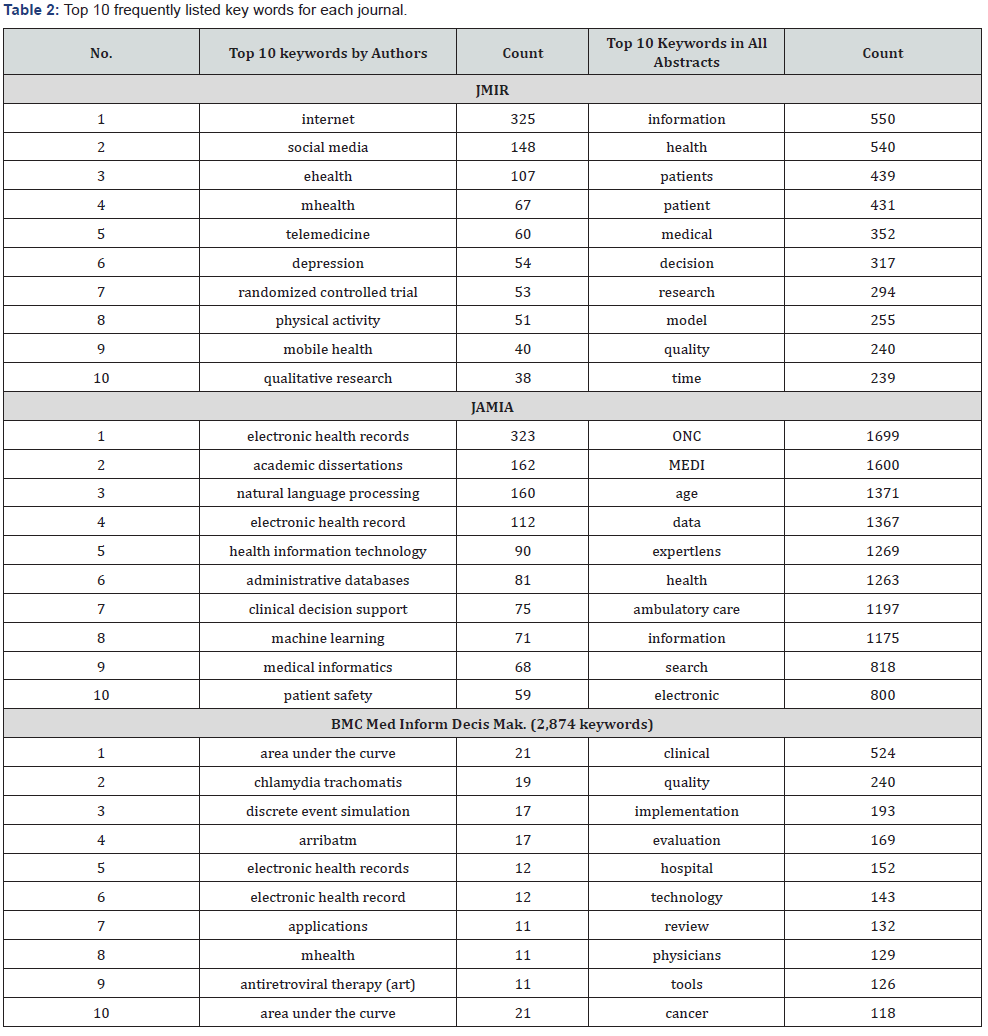
The most frequently used keywords listed by authors in papers (with keywords in the period from 2013 to 2017) are internet (JMIR), electronic health records (JAMIA), and area under the curve (MIDM), see Table 2. Relatively, the most frequently used keywords are information (JMIR), ONC (JAMIA), and clinical (MIDM) when applying journal keywords (2,051 in JMIR, 2,688 in JAMIA, and 1,246 in MIDM) to search abstracts of all papers from the beginning of the journal article publication.
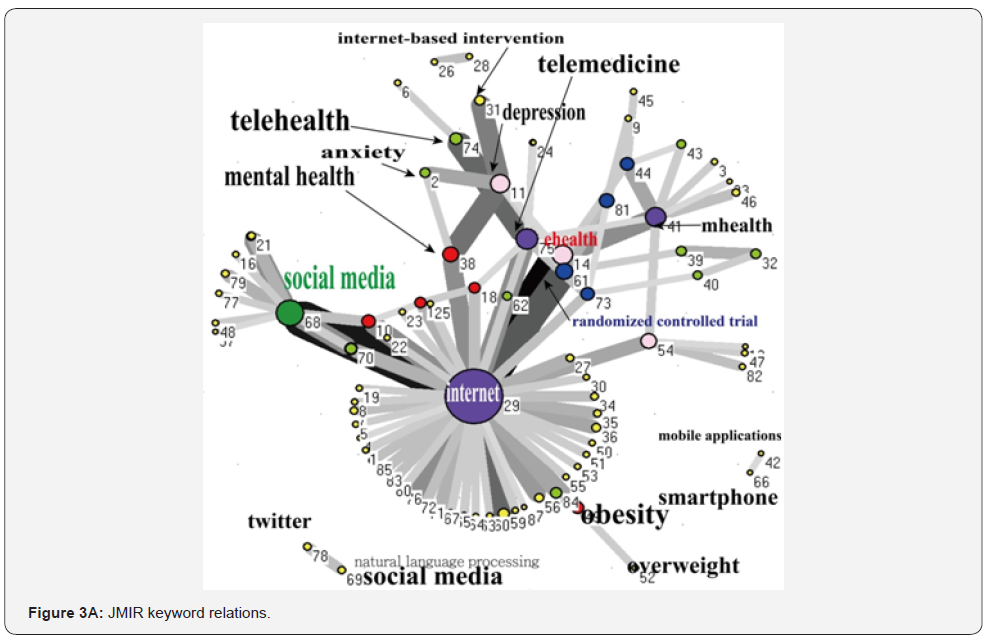
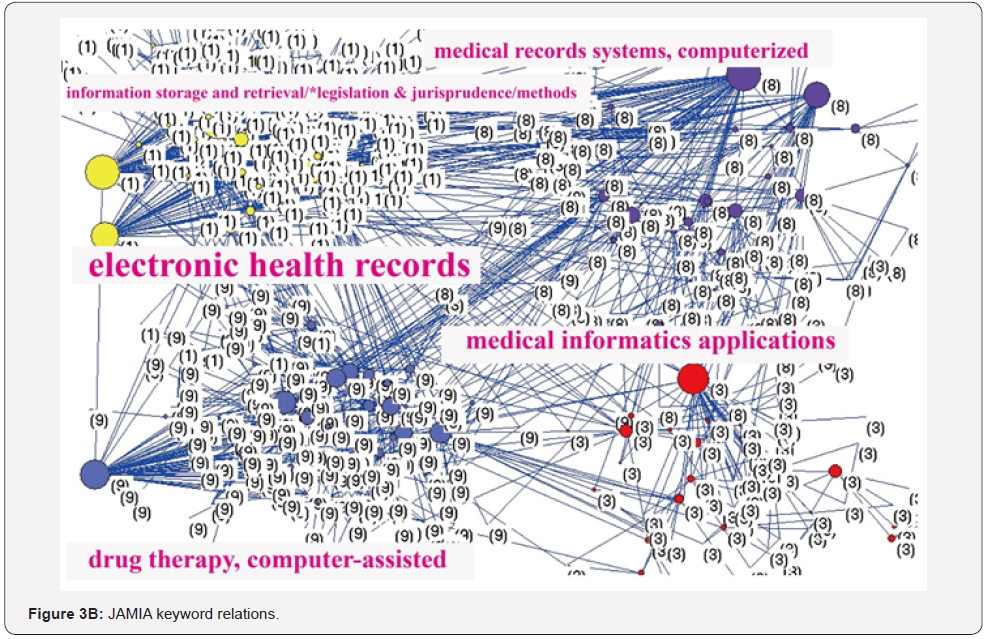
Pattern of the journals’ keywords
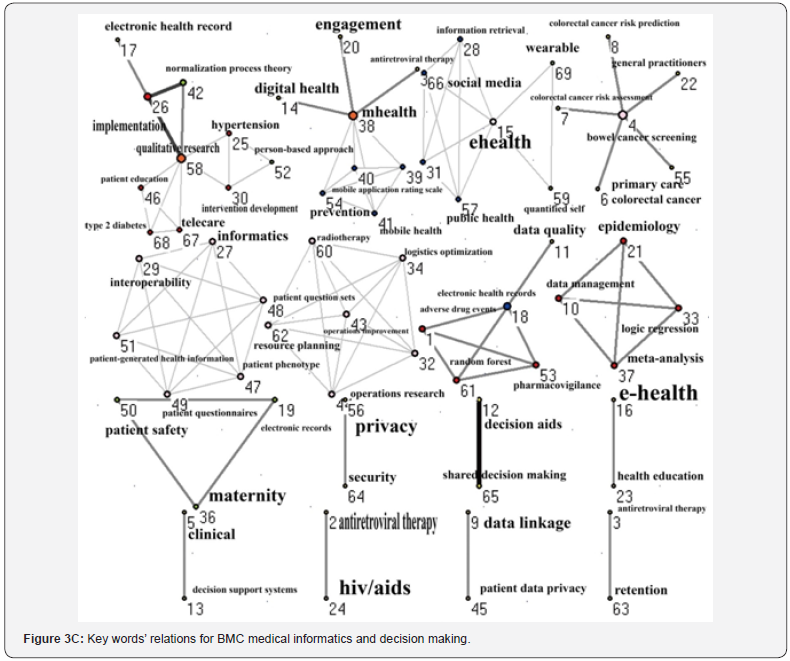
We traced the keyword patterns of the three journals. We can see that internet and electronic health records present a significant core category in networks of JMIR in Figure 3A and JAMIA in Figure 3B. MIDM, on the other hand, has not shown any core category in its network, see Figure 3C. The closest association pairs between two keywords are internet and social media for JMIR in Figure 3(A), electronic health records and health information technology policy for JAMIA in Figure 3B, and decision aids and shared decision making for MIDM in Figure 3C.
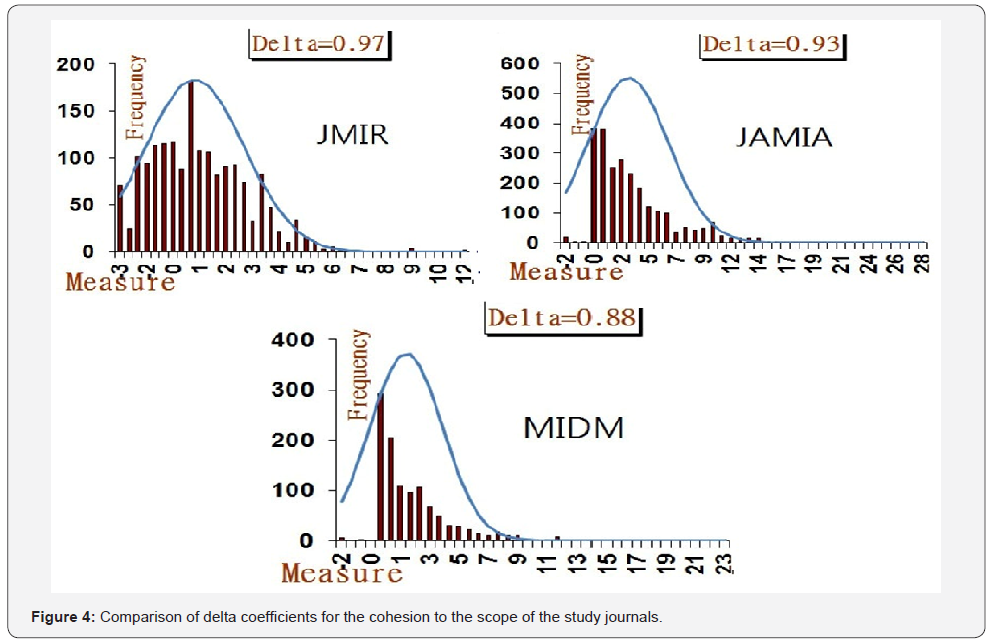
Measure targeting each journal’s scope and explanation of Ferguson’s delta coefficient
Excluding those cases without any keyword in a paper, the macro cohesion measures [=mean of standardized centrality = (Weighted all degree + Closeness + Between’s)/3] are 7.73 (SD=0.24) for JMIR, 4.47 (SD=0.25) for JAMIA, and 1.01 (SD=0.21) for MIDM, respectively, indicating that JMIR earns the greatest cohesion measure. The Ferguson’s delta coefficients are 0.86 for JMIR, 0.90 for JAMIA, and 0.97 for MIDM, respectively, implying that JAMIA suffers from less equality in the macro cohesion measure, see Figure 4.
Discussion
Key findings
The journal with the most cohesion is JMIR with a measure of 7.73 (SD=0.24). Both JMIR and MIDM earn a high Ferguson coefficient (0.96 and 0.97). Although MIDM gains the highest PKWA among the three journals, its keyword count begins in 2016, later than its two counterparts, which start in 2013.
What this adds to what was known
Many studies reported co-authorship relations within and between papers using SNA [23-25]. The association between beer and diaper sales [17-19] can be easily found by the SNA approach. However, we have not seen any paper using keywords in papers to investigate journal cohesion tendency and it’s PKWA, though keywords are required to be extracted from a paper’s title and abstract to help readers interested in its topic to find the article in the future
Through this study, we suggest that the journal editor’s assistant be able to (i) objectively measure the extent of paper cohesion in accordance with the journal scope and aims, as in Figure 4, (ii) efficiently examine keywords emerging in each paper’s abstract, and (iii) graphically depict journal’s keyword associations, as in Figure 3.
What it implies and what should be changed
Machine-learning algorithms and data mining have incorporated artificial intelligence based on Natural Language Processing (NLP) and Text Predictions to interpret natural spoken language [6-16], which could be applied to an article and its abstract. Before reaching this milestone, we are looking forward to seeing more papers that analyze keywords among similar journals using SNA.
In statistics, Exploratory Data Analysis (EDA) is an approach to analyzing data in order to summarize their main characteristics, often with visual methods. Thus, EDA discovers what the data can tell us beyond formal modelling or hypothesis testing [37]. The information shown in Figure 3 can help us know the journal image using the keyword SNA. Furthermore, journal editors and reviewers will focus more efforts on keywords and its PKW in the future. As a result, the journal’s aims and scope will be obviously recognized from its keywords’ alignment with the abstracts and titles of its contents.
Readers may be curious about the relations between centrality measures. We conducted a small study on correlations among Degree, Closeness and between’s [38]. The Closeness centrality (i.e., corr.≈0.30) is less correlated to the other two measures. The Degree is closely associated to between’s (i.e., corr.>0.90). For simplicity’s sake, we can select either Degree or between’s as a measure in the future, see Multimedia 3. In addition, the keyword is a noun instead of an adjective. We see some, such as medical and clinical abbreviations and acronyms, were found in Table 2. Journal editors and reviewers should put more emphasis on keyword correction and are suggested to use the checking system of Me SH term [39] in the future.
Strengths of this study
We present two videos in Multimedia 2 and 3 to interested readers: (i) how to extract data from such internet cloud databases as the US National Library of Medicine National Institutes of Health (Pubmed.org), and (ii) how to proceed with the cohesion measure using Pajek SNA software. Future researchers are suggested to mimic this approach on other journals’ keywords using SNA, which is somewhat different from search and extraction methods in literature [40].
We used SNA to analyze keyword associations in journals, which is different from others applying to health report issues [21,41]. In Figure 3, we can see that JMIR is dominated by the keyword “internet” and JAMIA by “electronic health record” because the closest association pairs are centred by the keywords “internet” and “electronic health records” for the both journals. As for MIDM, no special term was to dominate the journal, indicating that EDA is very different from initial data analysis (IDA) [37], which focuses more narrowly on checking the assumptions required for model fitting and hypothesis testing, handling missing values, and transforming variables as needed. EDA thus encompasses IDA to help us in policy making.
Limitations and future studies
This study has several limitations. First, all data were extracted from Pubmed.com. Some keywords were originally incorrectly saved in the dataset, such as comma, asterisk, and period separation symbols that interacted between keywords, and this will affect the results and inference making of the study. Second, there are many algorithms used for SNA. We merely applied the separation components shown in Figure 3. Any changes made along with algorithm used will present different pattern and judgment. Third, we applied Ferguson’s delta as a uniform distribution index that cannot represent any better or worse performance to the journal when the cohesion measure is an indicator used in the study. The major cohesion measure (i.e., the mean of the minor cohesion measures in papers) is suggested to be used to determine the focus of journal aims and scope attained. A cutting point is needed to determine in the future for any specific journal. Fourth, social network analysis is not subject to the Pajek software we used in this study. Others, such as Ucinet [42] & Gephi [43], are recommended to readers for use in future studies considering the topic of journal keyword analysis.
Conclusion
It is necessary to apply the compute module in inspecting whether keywords are within abstracts. The cohesion measure provides journal editors with a way to examine keywords accurately within an abstract before reviewing the paper.
Acknowledgment
We thank Frank Bill who provided medical writing services to the manuscript and Chi-Mei Medical Center for offering grand fund to the cost spent of the study.
References
- International Committee of Medical Journal Editors (1997) Uniform requirements for manuscripts submitted to biomedical journals. N Engl J Med 336: 309-316.
- http://www.inderscience.com/www/id31_keywords.pdf
- Nadim A (2005) How to write a scientific paper? ain shams. Journal of Obstetrics and Gynecology 2: 256-258.
- Rhodes W (2010) Guest editorial: the abstract as a marketing tool. Optical Engineering 49: 7.
- Day R, Gastel B (2006) How to write and publish a scientific paper, (6th edn), Westport, Greenwood Press, Connecticut, USA.
- Luo L, Li L, Hu J, Wang X, Hou B, et al. (2016) A hybrid solution for extracting structured medical information from unstructured data in medical records via a double-reading/entry system.BMC Med Inform Decis Mak 16: 114.
- Knoblock CA, Lerman K, Minton S, Muslea I (2003) Accurately and reliably extracting data from the web: a machine learning approach. Intelligent exploration of the web, pp. 275-87.
- Feldman R, Sanger J (2007) The text mining handbook: advanced approaches in analyzing unstructured data. Cambridge University Press, Cambridge, UK.
- Buneman P, Davidson S, Fernandez M, Suciu D (1997) Adding structure to unstructured data. In: Database Theory-ICDT’97, pp. 336-350.
- Buneman P, Davidson S, Hillebrand G, Suciu D (1996) A query language and optimization techniques for unstructured data. ACM, pp. 505-516.
- Buneman P, Davidson SB, Suciu D (1995) Programming constructs for unstructured data. IRCS Technical Reports Series, p. 121.
- Blumberg R, Atre S (2003) The problem with unstructured data. DM Rev 13(42-49): 62.
- Sebastiani F (2002) Machine learning in automated text categorization. ACM Comput Surv (CSUR) 34(1): 1-47.
- Holzinger A, Stocker C, Ofner B, Prohaska G, Brabenetz A, et al. (2013) Combining HCI, natural language processing, and knowledge discovery-potential of IBM content analytics as an assistive technology in the biomedical field. Human-Computer Interaction and Knowledge Discovery in Complex, Unstructured, Big Data, pp. 13-24.
- Hripcsak G, Friedman C, Alderson PO, Dumouchel W, Johnson SB, et al. (1995) Unlocking clinical-data from narrative reports - a study of natural-language processing. Ann Intern Med 122(9): 681-688.
- Tran T, Luo W, Phung D, Harvey R, Berk M, et al. (2014) Risk stratification using data from electronic medical records better predicts suicide risks than clinician assessments. BMC Psychiatry 14: 76.
- Domingos P (2012) A few useful things to know about machine learning. Communications of the ACM 55(10): 78-87.
- Verhoef PC, Kooge E, Walk N (2016) Creating value with big data analytics: making smarter marketing decisions. Routledge, London, UK.
- Power DJ (2002) What is the “true story” about data mining, beer and diapers? DSS News 3(23).
- Takahashi Y, Ishizaki T, Nakayama T, Kawachi I (2016) Social network analysis of duplicative prescriptions: One-month analysis of medical facilities in Japan. Health Policy 120(3): 334-341.
- Stewart SA, Abidi SS (2012) Applying social network analysis to understand the knowledge sharing behaviour of practitioners in a clinical online discussion forum. J Med Internet Res 14(6): e170.
- DeNooy W, Mrvar A, Batagelj V (2011) Exploratory Social Network Analysis with Pajek: Revised and Expanded (2nd edn), NY: Cambridge University Press, New York, USA.
- Zare Farashbandi F, Geraei E, Siamaki S (2014) Study of co-authorship network of papers in the Journal of Research in Medical Sciences using social network analysis. J Res Med Sci 19(1): 41-46.
- Sadoughi F, Valinejadi A, Shirazi MS, Khademi R (2016) Social network analysis of Iranian researchers on medical parasitology: a 41 year coauthorship survey. Iran J Parasitol 11(2): 204-212.
- Osareh F, SeratiShirazi M, Khademi R (2014) A Survey on coauthorship network of Iranian researchers in the field of pharmacy and pharmacology in web of science during 2000–2012. J Health Admin17 (56): 33-45.
- Landon BE, Keating NL, Barnett ML, Onnela JP, Paul S, et al. (2012) Variation in patient-sharing networks of physicians across the United States. JAMA 308(3): 265-273.
- Barnett ML, Christakis NA, O’Malley J, Onnela JP, Keating NL, et al. (2012) Physician patient-sharing networks and the cost and intensity of care in US hospitals. Med Care 50(2): 152-160.
- Landon BE, Onnela JP, Keating NL, Barnett ML, Paul S, et al. (2013) Using administrative data to identify naturally occurring networks of physicians. Med Care 51(8): 715-721.
- Osareh F, Khademi R, Rostami MK, Shirazi MS (2014) Co-authorship Network Structure Analysis of Iranian Researchers’ scientific outputs from 1991 to 2013 based on the Social Science Citation Index (SSCI). Collnet J Scientometr Info Manag 8(2): 263-71.
- Liu X, Bollen J, Nelson ML, Van de Sompel H (2005) Co-authorship networks in the digital library research community. Info Processing & Managment 41(6): 1462-1480.
- Ferguson GA (1949) On the theory of test discrimination. Psychometrika 14(1): 61-68.
- Hankins M (2008) How discriminating are discriminative instruments? Health Qual Life Outcomes 6: 36.
- Hankins M (2007) Questionnaire discrimination: (re)-introducing coefficient Delta. BMC Medical Research Methodology 7:19.
- Chien TW, Djaja N (2014) Using Rasch simulation data to verify whether Ferguson Delta coefficient can report students’s abilities are equal in a class. Rasch Meas Trans 28(3): 1484-1485.
- Chien TW, Lin WS (2016) Simulation study of activities of daily living functions using online computerized adaptive testing. BMC Med Inform Decis Mak 16(1): 130.
- Badar K, Hite JM, Badir YF (2013) Examining the relationship of coauthorship network centrality and gender on academic research performance: the case of chemistry researchers in Pakistan. Scientometrics 94(2): 755-775.
- Andrienko N, Andrienko G (2005) Exploratory analysis of spatial and temporal data. A systematic approach. Springer
- Rouleau G, Gagnon MP, Côté J, Payne Gagnon J, Hudson E, et al. (2016) How Do Information and Communication Technologies Influence Nursing Care? Stud Health Technol Inform 225: 934-935.
- Golan R, Bernstein AN, McClure TD, Sedrakyan A, Patel NA, et al. (2017) Partial gland treatment of prostate cancer utilizing high-intensity focused ultrasound in the primary and salvage setting: a systematic review. J Urol 5347(17): 54786-54792.
- Takahashi Y, Ishizak T, Nakayama T, Kawachi I (2016) Social network analysis of duplicative prescriptions: One-month analysis of medical facilities in Japan. Health Policy 120(3): 334-341.
- Zhao K, Wang X, Cha S, Cohn AM, Papandonatos GD, et al. (2016) A multirelational social network analysis of an online health community for smoking cessation. J Med Internet Res 18(8): e233.
- Borgatti SP, Everett MG, Freeman LC (2002) Ucinet for windows: software for social network analysis. MA: Analytic Technologies, Harvard, USA.
- Bastian M, Heymann S, Jacomy M (2009) Gephi: an open source software for exploring and manipulating networks. International AAAI Conference on Weblogs and Social Media.






























