α22-μ-Globulin Fragment: A Unique Protein in the Kidneys
Nadeem Kizilbash* and Farah Siddiqui
Department of Medical Laboratory Technology, Northern Border University, Saudi Arabia
Submission: April 18, 2018; Published: June 11, 2018
*Corresponding author:Nadeem Kizilbash, Department of Medical Laboratory Technology, Faculty of Applied Medical Sciences, Northern Border University, Saudi Arabia, Email: fsd707@gmail.com
How to cite this article: Nadeem K, Farah S. 2-μ-Globulin Fragment: A Unique Protein in the Kidneys. Curr Trends Biomedical Eng & Biosci. 2018; 15(2): 555910. DOI: 10.19080/CTBEB.2018.15.555910.
Abstract
α2-μ-Globulin (A2), an 18.6kDa protein produced in the liver, accumulates in the proximal tubule as a 15.5kDa cleavage product called “α2-μ-globulin fragment” (A2-f). A2-f possesses several unusual properties: (i) it escapes lysosomal degradation in the proximal tubules which is the common degradative pathway for all other proteins processed at this site; (ii) it accumulates selectively in the cytosol of proximal tubule, where it may represent the most abundant protein; and (iii) it crosses a biological membrane to enter the cytosol. The structural features that mediate this unusual behavior have never been identified.
Background and Significance
α2-μ-Globulin fragment (A2-f) represents about 5% of the total soluble protein content of the kidney [1,2]. It was initially thought to be a unique kidney fatty acid binding protein for several reasons: (a) it has a similar molecular size as all known FABPs, is found in the soluble protein fraction; and (b) it is abundant in cells with high fatty acid flux and binds long chain fatty acids in vitro [2,3]. Localization of A2-f to the proximal tubule, the site with the highest fatty acid flux in the kidney, further supported the idea that the 15.5kDa protein was a unique kidney fatty acid binding protein (kFABP) [2]. The complete sequence analysis of the protein revealed it is derived from the proteolytic conversion of the precursor protein, A2.
It is believed that unusual structural features permit A2-f to be targeted to the proximal tubule cell, to escape lysosomal degradation and to enter the cytosol [4,5]. A study showed that A2 is synthesized in the liver but not in the kidney. RT-PCR confirmed that mRNA for A2-f is abundant in the liver but cannot be detected in the kidney [6]. Removal of 9 amino acid residues from the C-terminus and 2-3 residues from the N-terminus of A2 (18.6kDa) produces A2-f, a protein with an apparent molecular weight of 15.5kDa [6]. Although the proteolytic conversion of A2 to A2-f is believed to be mediated by lysosomal cysteine proteases [7], the specific enzyme(s) responsible for converting A2 to A2-f have never been identified. To date, successful conversion of A2 to A2-f in vitro has not been reported.
A2-f binds long chain fatty acids in vitro with a 1:1 stoichiometry [2,3]. It is known that A2-f displays a very high binding affinity for long chain fatty acids with a Kd that ranges from 0.1-2mM, depending upon the structure of the specific fatty acid ligand [2]. Although the structure of A2-f has not been studied, the tertiary structure of A2 has only been determined by X-ray crystallography [8,9]. According to the published structure, each protein molecule envelopes a molecule of long chain fatty acid in its β-barrel cavity and transports it to mitochondria and peroxisomes where the fatty acids undergo β-oxidation to produce acetyl CoA and ultimately, generates ATP.
Structural Information
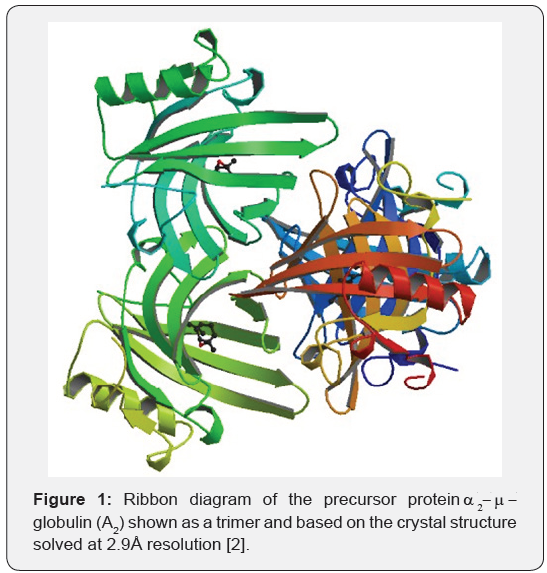
The crystal structure of the precursor protein, A2 (18.6kDa) has been reported at 2.8Å resolution (Figure 1). It identifies Tyr-124 as a key residue involved in hydrogen bonding of the carboxyl group of the ligand fatty acid [8]. According to the authors, the hydroxyl group of Try-124 interacts with the ionized form of the carbonyl group of the fatty acid in the ligand binding site. Tyr-124 points towards the electron density of the bound ligand (whose electron density appears to resemble that of a pheromone). Even though some structural features of A2 and A2-f may differ, crystallographic information from A2 is likely be very important for beginning the sequential assignment of the NMR resonances of A2-f in our study.
Conclusion
It has become increasingly clear over a period of time that A2-f plays many key physiological roles in the cell, including retinol, fatty acid and pheromone transport. It may also fulfill important functions in maintenance of innate immunity, cell regulation, morphogenesis and angiogenesis. It is believed that its precursor protein, A2, may represent an unusual example of a physiologic protein capable of accumulating as a fragment (A2-f) in a distant cell type.
References
- Hai A, Kizilbash NA, Alruwaili J (2014) International Journal of Bioscience, Biochemistry and Bioinformatics 4(2): 121-124.
- Lam KT, Borkan S, Claffey KP, Schwartz JH, Chobanian AV, et al. (1988) Properties and differential regulation of two fatty acid binding proteins in the rat kidney. J Biol Chem 263(30): 15762-15768.
- Hamilton JA (1998) Fatty acid transport: difficult or easy? J Lipid Res 39(3): 467-481.
- Hai A, Kizilbash NA (2013) α2-μ-Globulin fragment from kidneys of male rats. Bioinformation 9(3): 145-149.
- Wang Y, Shia MA, Christensen TG, Borkan SC (2000) Hepatic α2μ- globulin localizes to the cytosol of rat proximal tubule cells. Kidney International 57(3): 1015-1026.
- Kimura H, Odani S, Suzuki J, Arakawa M, Ono T (1989) Kidney fatty acid binding protein: identification as A2m-globulin. FEBS Letters 246: 101-104.
- Saito K, Kaneko H, Isobe N, Nakatsuka I, Yoshitake A, et al. (1992) Differences in A2m-globulins in male rat kidneys following treatment with A2m-globulin accumulating agents: cysteine protease(s) play(s) an important role in production of kidney-type A2m-globulin. Toxicology 76: 77-186.
- Böcskei Z, Groom CR, Flower DR, Wright CE, Phillips SE, et al. (1992) Pheromone binding to two urinary proteins revealed by X-ray crystallography. Nature 360(6400): 186-188.
- Chaudhuri BN, Kleywegt GJ, Bjorkman J, Lehman McK, Oliver JD, et al. (1999) The structures of alpha 2u-globulin and its complex with a hyaline droplet inducer. Acta Crystallogr D Biol Crystallogr 55(Pt 4): 753-762.






























