Computational Modeling Studies of the Beta-Amyloid Protein Binding to Develop Drugs for the Treatment of Alzheimer’s Disease
Sanidhya D Tripathi1,2, Patrycja A Krakowiak1,2 and Jerry A Darsey2,3*
1Arkansas School for Mathematics, Sciences, and the Arts, Hot Springs, AR, USA
2Center for Molecular Design and Development, Little Rock, AR USA
3Department of Chemistry, Little Rock, AR USA
Submission: December 17, 2017; Published: December 21, 2017
*Corresponding author: Jerry A Darsey, Department of Chemistry & Center for Molecular Design and Development at the University of Arkansas at Little Rock, Little Rock, AR 72204, USA, Tel: 501 569-8828; Email: jadarsey@ualr.edu
How to cite this article: Sanidhya D T, Patrycja A K, Jerry A D. Computational Modeling Studies of the Beta-Amyloid Protein Binding to Develop Drugs for the Treatment of Alzheimer’s Disease. Curr Trends Biomedical Eng & Biosci. 2017; 10(5): 555799. DOI: 10.19080/CTBEB.2017.10.555799.
Abstract
Alzheimer’s Disease (AD) is a progressive neurodegenerative disease that affects memory and other mental functions due to the accumulation of Amyloid-Beta (Aβ) plaques. The functions carried out by the Aβ protein leads to abnormalities in the suffering patient. If a molecule binds to the Aβ protein, it could lead to potential treatment. To propose new treatments for AD, novel molecules were identified that bind to the active site of Aβ. First, a correlation between reported IC50 values of fourteen chemicals and their energy gap, dipole moment, and total energy were analyzed. The best correlation was discovered between 1/IC50 values and total energy (R2=0.9279). Then, each identified molecule was virtually modified and its new IC50 value was determined utilizing Gaussian 09, a molecular modeling program, which was used to predict total energies. Some of the most useful modifications included the addition of oxygen (-O), chlorine (-Cl), and fluorine (-F) atoms. The results of this study can be used as a tool to finding novel, medically relevant new molecules for other neurodegenerative diseases and assist researchers in developing new drugs to treat AD
Keywords: Alzheimer’s disease; Beta-Amyloid protein; Drug design; Protein misfolding; amyloid-β aggregation; Oxidative stress; Neuroinflammatory processes; Parkinson’s disease; Huntington’s disease; Amyotrophic lateral sclerosis (ALS), IC50; Computational molecular modelling; Density functional theory
Introduction
Nearly six million people suffer from Alzheimer’s disease (AD) in the United States, and the number of patients with this multifaceted neurodegenerative disorder could rise to include as many as 16 million people by 2050 [1]. Moreover, in 2017, this disease is estimated to cost $259 billion in global health care expenditures [1]. Alzheimer’s disease is an irreversible neurological disorder that is characterized by memory loss, personality changes, and decline in thinking abilities. Though memory loss is the most widely recognized symptom of Alzheimer’s disease, many other complications, such as a range of behavioral and cognitive deteriorations, are also common. The cognitive symptoms include memory loss, confusion, and disorientation while the behavioral symptoms include depression, insomnia, and hallucinations [2]. Due to the often-intense nature of the symptoms, AD patients can suffer in a multitude of ways and eventually become bedridden and dependent on custodial care. On average, death occurs nine years after diagnosis [3].
AD is characterized at a molecular level by factors such as protein misfolding and aggregation, oxidative stress, and neuroinflammatory processes. The complex pathogenesis has provided for only a limited development of effective therapeutic solutions to combat disease progression [3,4]. Currently, there are several symptomatic medications and treatments for AD. Some of these registered treatments include cholinesterase inhibitors, donepezil, galantamine, and rivastigmine [2,4,5]. Although these drugs are beneficial to some degree, they only provide temporary improvements in cognition and function of the diagnosed patient. Because there is still no efficient medication to prevent the deterioration of the neurodegenerative processes in AD patients, there remains an urgent need for novel drugs that may slow the neurodegenerative disease and potentially reverse its effects. Overall, no treatment with a strong disease modifying effect is currently available.
The primary suspects that are thought to cause Alzheimer’s disease are amyloid-beta (Aβ) and hyper-phosphorylated tau proteins. Agents inhibiting one or both forms of aggregation are promising drug candidates for AD [6]. Amyloid-β peptide chains consist of 40 to 42 amino acids and are suspected to contribute to the degenerative condition of AD. The amyloidbeta precursor protein (APP) is found in many types of cell membranes. The actions of the β-secretase and γ-secretase release the Aβ proteins at an elevated rate into the cytoplasm. These proteins accumulate at high levels in neural cells; they are internalized and folded into a β-pleated shape which results in stacking that forms long aggregates known as senile plaques [2]. Acting as monomers, dimers, or multimers in cell membranes, the Aβ proteins interfere with neurotransmission and memory formation. As the protein aggregates continue to increase in size, they become insoluble in the bloodstream. Scientists have begun to study this problem and suggest treatment strategies, which include inhibitors of the β-secretase and γ-secretase activities, as drugs and physiological compounds to prevent amyloid-β aggregation in the brain [7].
Much research in pursuit of novel compounds that have efficient inhibitory properties of AD still needs to be completed. For example, innovative techniques are needed to offset the financial expenditures related to the traditional tissue culture and model organism experiments. Though several vaccines are in the process of being developed, a breakthrough discoveries are needed to speed up this slow process. A successful drug discovery process would not only revolutionize the treatment of AD and improve the lives of countless people all over the world, but it could also act as an example of how beneficial drugs for other diseases could be discovered in the future. In conjunction with previous research, drug candidates that bind to Aβ proteins could be designed based on specific properties known to enhance this binding. For example, chemical compounds that have a low IC50 value (a measurement of the effectiveness of a drug or inhibitorof binding to a biological molecule by IC50 % and expressed in micromolar concentration, M) could be identified and evaluated to better treat AD. As part of the described study, it was proposed that there exists a correlation between the IC50value and other notable parameters of a chemical compound and its interaction with Aβ proteins. Using computational molecular modeling and virtual simulations, molecules best suited to bind and therefore inhibit Aβ proteins were identified and tested.
As shown in previous studies, determining a molecule to bind to Aβ would result in a major step towards creating a drug to help treat Alzheimer’s disease [8,9]. By determining specific molecules that bind to the Aβ active site, a correlation can be found between its binding affinities and energies. To achieve this goal, computational molecular modeling was used in this research. The goal of this study was to create a molecule that would bind efficiently to the Aβ protein by finding correlations between chemical properties when creating the modified molecule. The plan of this research was to alter molecules that have been previously identified as inhibitors of Aβ and then determine by how much these modified molecules change the IC50 values of the “parent” molecule. By collecting data such as the dipole moment (the product of the separation of two ends of a dipole molecule and the magnitude of the charges), highest occupied molecular orbital (HOMO), lowest unoccupied molecular orbital (LUMO), energy gap (value found by subtracting the HOMO value from the LUMO value), and plotted vs. the IC50 value, a correlation was determined and analyzed after the modifications were incorporated. New, IC50 values were then predicted using the line of best fit. The newly designed compounds with a lower predicted IC50 values would be a potential new drug to treat Alzheimer’s disease.
Materials and Methods
The research was initiated by locating and analyzing the native structure of the Beta-Amyloid (Aβ) protein. After a detailed study of their properties and relevance to Alzheimer’s disease, 21 compounds were identified in publications and patents that had been tested as potential inhibitors of the Aβ protein [10]. However, the molecules mentioned in various sources were not always described using the IUPAC naming system. Thus, the correct name of these molecules had to be determined.
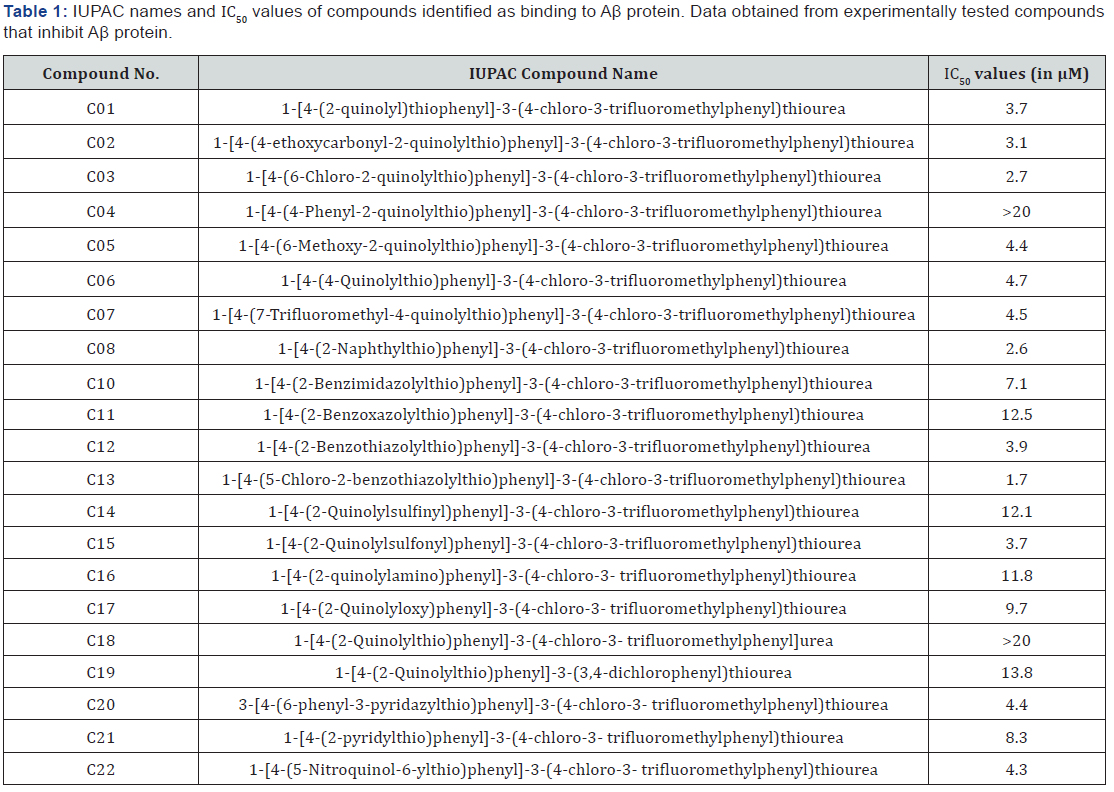
To accomplish this task, PubChem (https://pubchem.ncbi. nlm.nih.gov/) was utilized as the database to find important information about the molecules and their activities with respect to Aβ protein. PubChem, a free public tool is maintained by the National Center for Biotechnology Information (NCBI) Center at the National Institutes of Health (NIH) and provides the IUPAC (International Union of Pure and Applied Chemistry) name, InChIKey (International Chemical Identifier), and chemical structures of more than 92.1 million compounds. To enable the extraction of this information from the PubChem database, the OPSIN tool (Open Parser for Systematic IUPAC nomenclature), freely available from the University of Cambridge, was used. The InChIKey was identified for each chemical and was then used to search for chemical properties on the PubChem website. In ChIKey is a textual chemical identifier for compounds that encodes chemical information and can be used to search online databases. To extract all InChIKeys for the 22 identified compounds, a Google Sheet was created with the first column showing the original name of each chemical as cited in each publication. Table 1 shows the compounds identified by thoroughly searching the literature and their respective IC50 values [10].
Compound Alterations
Using the computational molecular modeling program (Gaussian 09), each chemical was altered by adding specific functional groups and thus resulted in the generation of a new group of compounds. Elements, bond types/sizes, and chemical groups were altered using tools accessible to the user. Each molecule was analyzed using the optimization method, which was applied to clean the molecular bonds and structures utilizing the DFT (Density Functional Theory) 6-31G* basis set (defined for atoms H to Zn and is a double-zeta polarized basis set). The total energy (a.u.), dipole moment (Debye), and HOMOLUMO (Highest Occupied Molecular Orbital-Lowest Unoccupied Molecular Orbital) energy gap (Hartrees), were established and recorded for each molecule.
Statistical correlations
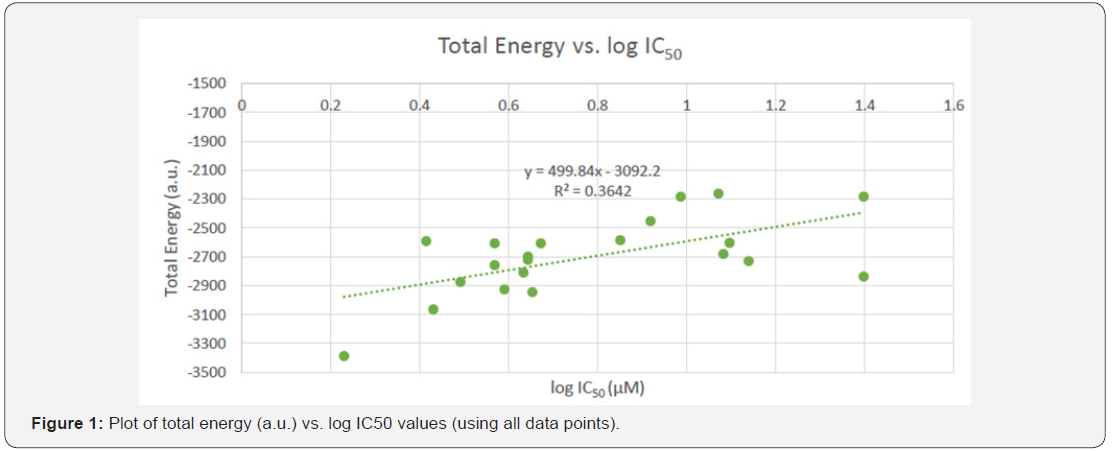
Once the data from all the trials was recorded, binding affinities, measured by the experimentally determined IC50 values of each of the 22 molecules. These values would serve as a tool to later produce modified compounds and their IC50 values to be compared to the original unmodified molecule. The data was entered into an Excel 2016 spread sheet and graphed using the scatter plot tool. Plots included IC50 , log IC50 , and 1/ IC50 values vs. the various calculated parameters (the total energy, dipole moment (Debye), and HOMO-LUMO energy gap). From these plots, the strength of correlations were determined. It should be noted that some data points were outliers and were therefore removed from the data set since they were more than two standard deviations away from the linear trendline. Plots were included with and without the outliers (Figures 1-4).

Results
Data points from the plots of the various Amyloid-Beta (Aβ) inhibitors were used to identify characteristics that would show the best correlations with IC50 values. The values of the total energy, dipole moment, and energy gap of the promising compounds were calculated using the Gaussian 09 software. Numerous correlations were looked at including total energy vs. log IC50 , total energy vs. 1/ IC50 , total energy vs. IC50 , dipole moment vs. log IC50, dipole moment vs. 1/IC50, dipole moment vs. IC50 , energy gap vs. log IC50 , energy gap vs. 1/ IC50 , and energy gap vs. IC50 values. The analysis revealed that the most highly correlated properties were total energy vs. 1/ IC50 values and total energy with log IC50 values (Figures 1-4). This information was then used to find novel, modified molecules and more efficient Aβ compounds that could become new drugs to potentially reduce the negative physiological effects of Alzheimer’s Disease.
The log IC50 values of all the literature derived compounds were used to determine a relationship between total energy and IC50 values. The plot reveals that the log IC50 values for all. The log IC50 values were used to determine a relationship between total energy and IC50 values. The plot reveals that the log IC50 values are proportional to the total energy; this statement is supported by the linear equation and R2 =0.8157 value of the trendline.
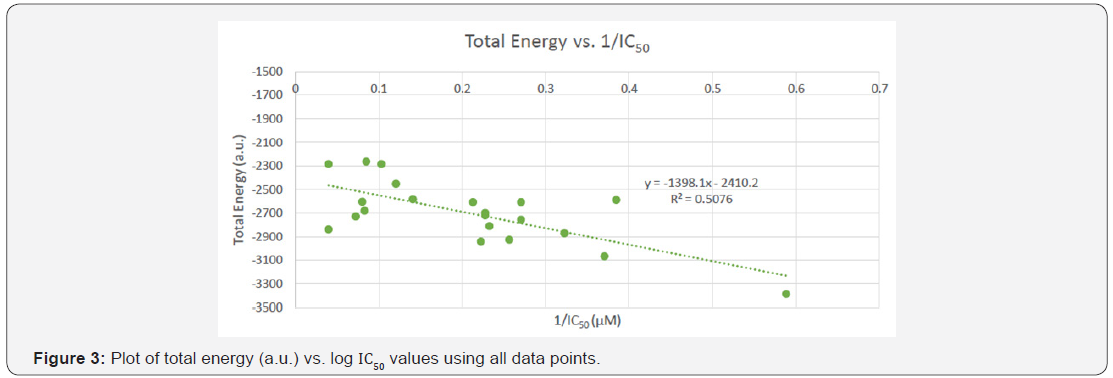
The 1/ IC50 values of all the literature derived compounds were used to determine a relationship between total energy and IC50 values and are plotted with respect to total energy that was determined using the Gaussian 09 program (see Figure 3). The plot depicts a correlation between the 1/ IC50 values and total energy, with an R2=0.5076 value.
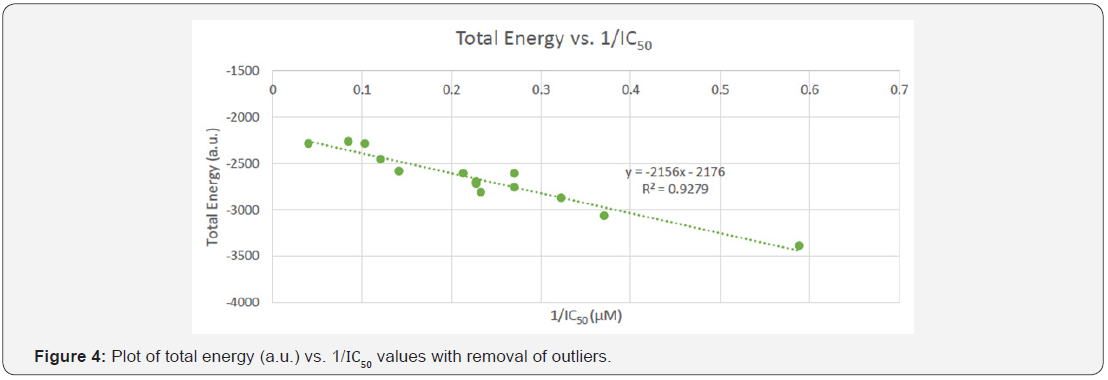
The data points in Figure 4, show the I IC50 values that were obtained from previous experimental research after the removal of the outliers; while the total energy associated with each was obtained using the Gaussian 09 software. The 1/I C50 values were used to determine a relationship between the two parameters. The graph depicts how the 1/ IC50 values are inversely proportional to the total energy, according to the equation and R =0.9279 value of the trendline.
Modifications
Another set of trials was conducted after the highest correlation was determined i.e. total energy vs. 1/IC50 values). To find novel molecules with lower IC50 values than those mentioned in the literature, simple modifications had to be made to accomplish this task. Therefore, the fourteen compounds were modified by an addition of chemical groups/elements to the existing structure and a new IC50 value was calculated after the total energy value was outputted from the Gaussian 09 program. For this round of testing, each molecule was run first under the optimization method, and then using the DFT (Density Functional Theory) with the 6-31G* basis set. Table 2 shows the results of the new IC50 values after modifications had been completed.
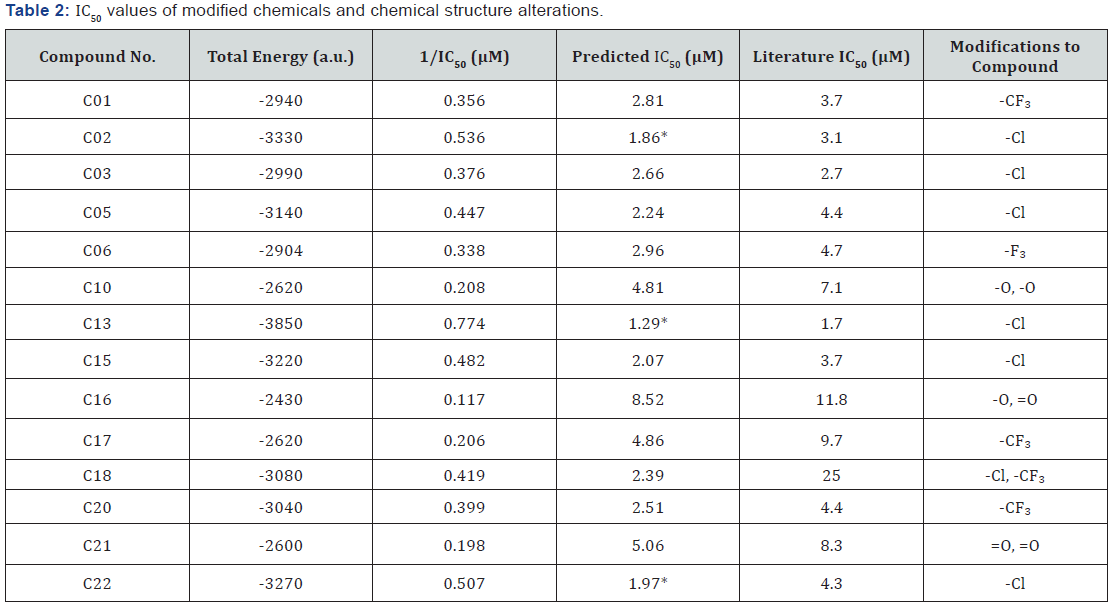
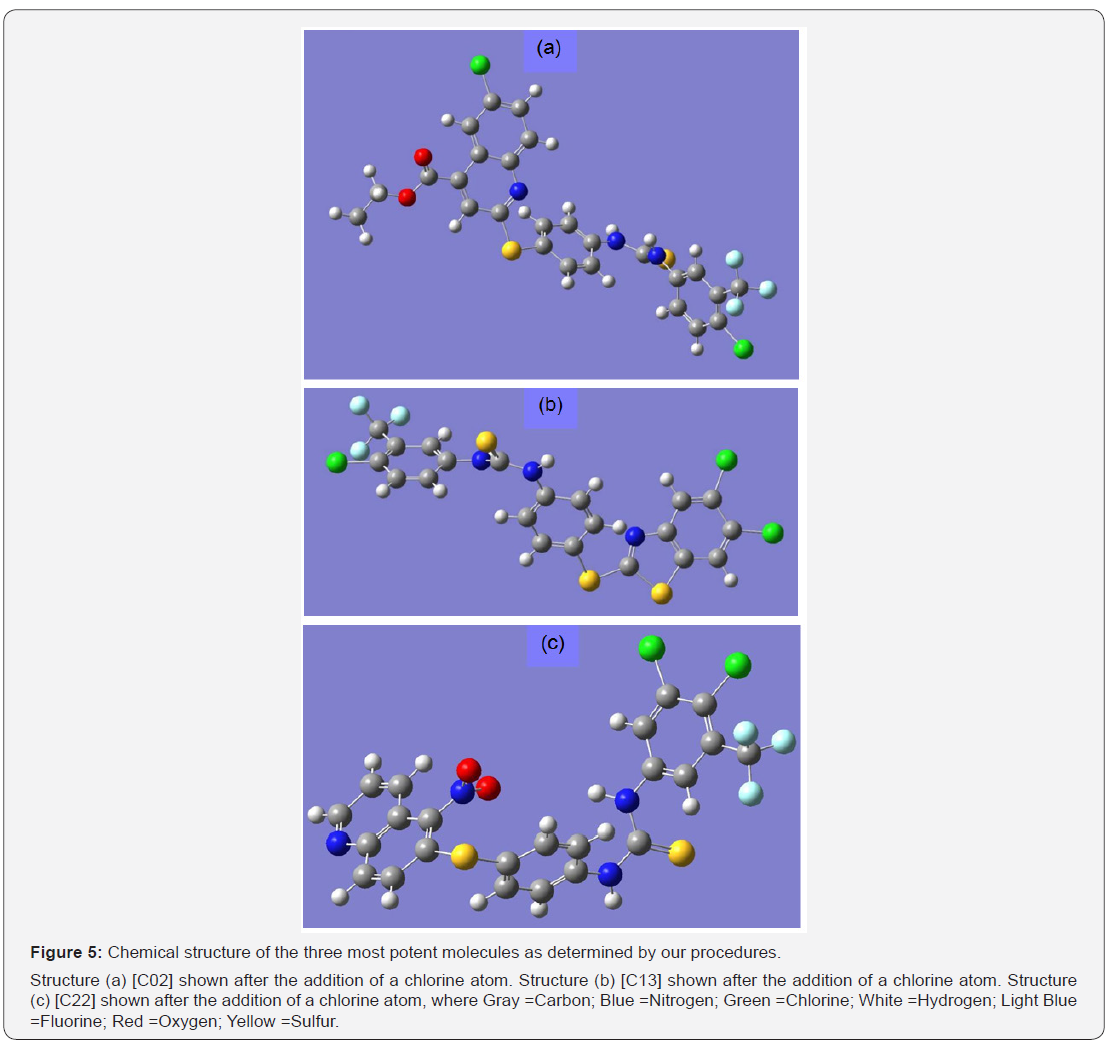
After running the DFT with the 6-31G* basis set on all the compounds, 3 compounds identified as C02, C13, C22 yield the lowest IC50 (largest binding affinities) values in the sets tested. Their chemical structures are show in order in Figure 5a-5c.

Discussion
The accumulation of the Beta-Amyloid (Aβ) proteins, which are derived from the Amyloid Precursor Protein (APP), can form plaques within the human brain [6,11-15]. This ability to form plaque can cause numerous health problems to the patient, which culminate in Alzheimer’s disease. It was therefore beneficial to identify molecules that could bind to these plaque molecules marking them for excretion from the body. The purpose of the current study was to investigate not only whether such potential drug molecules exist, but also to design modifications that would improve their binding capacity. To accomplish this goal, first, it was important to establish if there is a correlation between the literature-derived IC50 values of various chemicals and notable properties including total energy, energy gap, and/or the dipole moment. The property that correlated best with was the 1/IC50 values vs. total energy with an R2=0.9279. Therefore, total energy was calculated for the proposed modifications to the original compounds and the IC50 values for these were determined using total energies and the linear regression equation in Figure 4. The chemical modifications with lower determined IC50 values were selected for further consideration. Any chemical that exhibited lower IC50 values than its original parent molecule should be tested experimentally for efficacy of binding Aβ plaques and its ability to treat Alzheimer’s disease.
In the computational molecular modeling field of research, having a reference chemical is a vital aspect of future research. Based on the procedures of this current study, novel compounds could be identified for various other diseases such as Parkinson’s, Huntington’s, and Amyotrophic lateral sclerosis (ALS). In addition to the distinguishing aspect of this research that allows for modeling treatment for various notable diseases, it also allows for far less time consuming laboratory work since so much of the process is done computationally. Because synthesizing novel chemicals and measuring their IC50 values is both laborious and expensive, the computational research reported here is invaluable to the future expansion of pharmacology. Because scientists do not have to spend large amounts of time synthesizing chemicals and testing them, the computer-aided approach offers a much faster and efficient way to obtain data critical to improving treatment of many deadly diseases. Using this procedure, hundreds of molecules can be analyzed and a far greater number of novel compounds can be identified as promising medications. Ultimately, identifying compounds with high binding affinities for the Aβ protein (with lower IC50 values) that can act as inhibitors, can be applied to Alzheimer’s disease research to develop new drugs to treat patients.
Conclusion
The set of experiments described here reveal an opportunity to design new molecular Amyloid Beta (Aβ) inhibitors using computational molecular modeling techniques. The hypothesis that new IC50 values of virtually altered chemicals could be predicted by first finding a linear correlation between reported IC50 values and a specific, calculable property of these molecules was supported when a strong correlation identified between Total Energy and 1/ IC50 values led to the discovery of lower IC50 values of the modified molecules. More specifically, the linear regression equation generated after graphing 1/ IC50 values vs. Total Energy was used to predict how modifying the previously reported chemicals could lower their predicted IC50 values and so increase their binding capacities to Aβ. The data that allowed for these predictions was generated using the quantum mechanical Gaussian 09 software program. The significance of this novel work is that it suggests new drug candidates to treat Alzheimer’s disease (AD). One of the main causes of the mental symptoms of AD are a buildup of Aβ plaques in the brain. By proposing modifications to various chemicals that improve binding and therefore inhibit Aβ, the experiments described here allow for a set of novel chemicals to be tested in tissue culture and animal models with Aβ buildup. The benefit of doing the preliminary experiments virtually using a computer-aided approach is that it is cheaper, faster and more efficient than biological synthesis and testing. Finding leading drug candidates for the treatment of AD, is critical for future researchers to be able to develop effective medications to combat this debilitating and deadly disease.
The presented research could also lead to investigations of other protein misfolding diseases including Huntington’s disease, Parkinson’s disease, and various Prion diseases. The computational molecular modeling techniques described in this manuscript could be used to identify molecules that bind proteins associated with specific diseases, and then promote the discovery of novel drugs by proposing modifications that could make these molecules more efficient inhibitors, thus producing better treatments for these neurodegenerative diseases. These computationally driven findings could allow for quicker discoveries of treatments that could improve and even save millions of lives.
Competing Interests
The authors have declared that there exist no known competing interests.
Author Contributions
Sanidhya Tripathi (ST) carried out all the procedures. ST and Jerry Darsey (JD) conceived of the study and performed the statistical analysis. ST drafted the manuscript with the help of PK and JD. JD and PK read and approved the final manuscript.
Acknowledgement
The authors would also like to thank the University of Arkansas at Little Rock Center for Molecular Design and Development and the Arkansas School for Mathematics, Sciences, and the Arts for facilitating my research and providing the venue to share these research findings.
Furthermore, the authors would like to thank Ms. Sumreen Gul, a Ph.D. student, for help with running the Gaussian 09 software and giving one of the authors (ST) consistent feedback and tips throughout the process. Finally, the author (ST) would like to thank the parents Sanjay and Arati Tripathi for all the encouragement and support.
References
- Alzheimer’s Association (2016) Alzheimer’s disease facts and figures Alzheimer’s Dementia 12(4): 459-509.
- Godyń J, Jończyk J, Panek D, Malawska B (2016) Therapeutic strategies for Alzheimer’s disease in clinical trials. Pharmacological Reports 68(1): 127-138.
- Citron M (2002) Alzheimer’s disease: treatments in discovery and development. Nature neuroscience 5: 1055-1057.
- Lahiri DK, Farlow MR, Greig NH, Sambamurti K (2002) Current drug targets for Alzheimer’s disease treatment. Drug Development Research 56(3): 267-281.
- Jia Q, Deng Y, Qin, H (2014) Potential therapeutic strategies for Alzheimer’s disease targeting or beyond β-amyloid: insights from clinical trials. BioMed research international 2014: 837157.
- Arbor SC, LaFontaine M, Cumbay M (2016) Amyloid-beta Alzheimer targets-protein processing, lipid rafts, and amyloid-beta pores. The Yale Journal of Biology and Medicine 89(1): 5-21.
- Seeman P, Seeman N (2011) Alzheimer’s disease: β‐ amyloid plaque formation in human brain. Synapse 65(12): 1289-1297.
- Dovey HF, John V, Anderson JP, Chen LZ, de Saint Andrieu P, et al. (2001) Functional gamma‐ secretase inhibitors reduce beta‐ amyloid peptide levels in brain Journal of neurochemistry 76(1): 173-181.
- Fuse S, Matsumura K, Fujita Y, Sugimoto H, Takahashi T (2014) Development of dual targeting inhibitors against aggregations of amyloid-beta and tau protein. Eur J Med Chem 85: 228-234.
- Reel JK, Simon RL, Whitesitt CA (1997) US Patent No. 5,624,937. DC: U.S. Patent and Trademark Office, Washington, USA.
- Friedrich RP, Tepper K, Rönicke R, Soom M, Westermann M, et al. (2010) Mechanism of amyloid plaque formation suggests an intracellular basis of Aβ pathogenicity. Proceedings of the National Academy of Sciences 107(5): 1942-1947.
- Irvine GB, El-Agnaf OM, Shankar GM, Walsh DM (2008) Protein aggregation in the brain: the molecular basis for alzheimer’s and parkinson’s diseases. Molecular Medicine 14(7-8): 451-464.
- Estrada LD, Soto C (2007) Disrupting β-amyloid aggregation for Alzheimer disease treatment. Current topics in medicinal chemistry 7(1): 115-126.
- Murphy MP, LeVine H (2010) Alzheimer’s Disease and the β-Amyloid Peptide. Journal of Alzheimer’s Disease : JAD, 19(1): 311.
- Selkoe DJ, Schenk D (2003) Alzheimer’s disease: molecular understanding predicts amyloid-based therapeutics. Annual review of pharmacology andtoxicology 43(1): 545-584.






























