Virulence of Pasteurella Multocida Isolated During Pasteurellosis Outbreak among Saigas in Kazakhstan
Kamshat A Shorayeva*, Nurkuisa M Rametov, Mukhit B Orynbayev, Kainar . Barakbayev, Nurlan T Sandybayev, Abylay R Sansyzbay and Kulyaisan T Sultankulova
The research Institute for Biological Safety Problems, Ministry of Education and Science of Republic of Kazakhstan, Kazakhstan
Submission: August 04, 2017; Published: August 29, 2017
*Corresponding author: Kamshat A Shorayeva, The research Institute for Biological Safety Problems, Ministry of Education and Science of Republic of Kazakhstan-Science Committee, Gvardeiskiy, Kordaiskiy rayon, Zhambylskaya oblast, 080409, Republic of Kazakhstan, Email: K.A.shorayeva@mail.ru
How to cite this article: Kamshat A S, Nurkuisa M R, Mukhit B O, Kainar . B, Nurlan T S, et al. Virulence of Pasteurella Multocida Isolated During Pasteurellosis Outbreak among Saigas in Kazakhstan. Curr Trends Biomedical Eng & Biosci. 2017; 8(2): 555735. DOI: 10.19080/CTBEB.2017.08.555735
Abstract
For the past few years mass deaths of saiga antelopes was recorded in the Republic of Kazakhstan. The outbreaks of Pasteurella infection caused drastic reduction of saiga population. The isolates of P.multocida belong to multocida subspecies. Dedicated from the dead saigas in 20102011 gg. Pasteurella multocida isolates contained virulent genes. The study was performed with use of 25 (2010) and 8 (2011) P.multocida field isolates from different organs of dead saigas.
Fifteen virulence-associated genes (ptfA, ompA, ompH, oma87, plpB, fimA, hsf-2, pfhA, exbB, tonB, hgbA, fur, nanB, nanH and pmHAS) were detected by multiplex PCR. In isolates of P.multocida from saigas (year 2010) the following genes were detected: tonB (100%), ompA (92%), nanH (92%), fimA (68%), ptfA (68%), exbB (68%), oma87 (48%), plpB (44%), ompH (16%) and pfhA (12%). The genomes of P.multocida isolated from saigas in 2011 contained the following genes: hgbA (100%), ompA (100%), tonB (100%) and nanH (100%); fimA (87.5%), ptfA (87.5%) and exbB (87.5%); pfhA (62.5%), hsf2 (62.5%), fur (62.5%) and none of plpB, oma87, ompH, nanB and pmHAS genes was found in them. All assayed isolates showed high content of virulent hgbA, ompA, tonB and nanH genes. Field isolates of P.multocida (years 2010 and 2011) from saiga antelopes differ in virulence gene profiles.
Keywords: Pasteurella multocida; virulence factors; PCR; saiga; Kazakhstan
Introduction
Saiga antelope (Saiga tatarica) is entered in the International Red Book because of drastic decline of its population. Mass death of saigas was observed on the territory of the Republic of Kazakhstan in 2010-2011. About 12000 in 2010 and over 500 animals in 2011 died [1] Taking into account the current low saiga numbers emergence of epizooty can be a real threat to the life story of this animal species. Assay of samples from dead saigas (Borsy village, West Kazakhstan oblast, Republic of Kazakhstan) at the RIBSP in 2010-2011 resulted in isolation of pathogenic organisms that were differentiated as Pasteurella [2]. is a Gram negative bacterium belonging to Pasteurellaceae family. Subject to the specific type of capsular antigens P.multocida has serologic differences and subdivides into capsular serogroups (A, B, D, E, F) [3,4]. The currently known virulence factor genes include genes encoding the outer membrane proteins (ompA, ompH, oma87, plpB), adhesins (hsf1, hsf2), iron transport genes (fimA, ptfA, exbB, tadD, fur), enzymes (nanH, nanB, pmHAS). It should be also noted that there are genes responsible for high virulence of P.multocida; they are pfhA gene encoding filamentous hemagglutinin, hgbB gene that is responsible for hemoglobin control in a body [5-7].
The work was conducted towards determining subspecies and serotyping of P.multocida bacteria isolated from saiga antelopes in 2010 and 2011 in the Republic of Kazakhstan. Pathogenicity of P.multocida strains and isolates is assessed by presence (%) of virulent genes [8,9]. It is clear that high mortality rate among saiga antelopes in the Republic of Kazakhstan is directly related to the virulence factors of P.multocida. On the basis of the above stated the objective of the study was to detect the virulent genes in the genomes of P.multocida isolated from saiga antelopes in 2010 and 2011 in the Republic of Kazakhstan.
Materials and Methods
All ethics, field and laboratory studies were reviewed and approved by the appropriate committees of the Research Institute for Biological Safety Problems (RIBSP), Ministry of Education & Science in Gvardeiskiy, Zhambylskaya oblast, Republic of Kazakhstan.
Clinical examination, sampling and specimen delivery
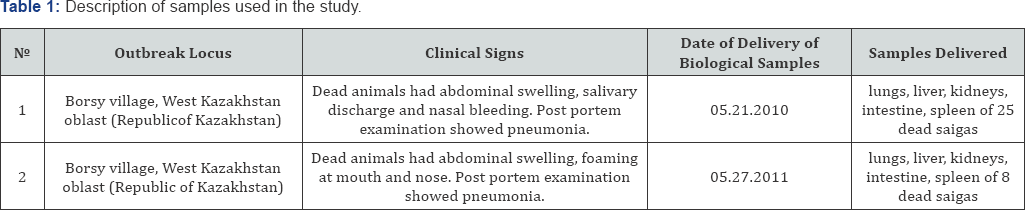
Twenty five samples taken from dead saigas in 2010 and 8 samples of year 2011 were delivered to the RIBSP by veterinary specialists from West Kazakhstan oblast within the network of epidemiological surveillance to determine the cause of saigas illness and death. Suspected pasteurellosis clinical cases were reported, and biological samples were taken from dead saigas for diagnosis. The samples taken from each animal were placed separately in indelibly marked plastic bags with the species name, animal i.d. and tissue name. These were packed in iced refrigerator bags and delivered frozen to the RIBSP. Detailed information on clinical signs and samples taken is shown in Table 1.
DNA Extraction
DNA was extracted with use of "PrepMan Ultra Sample Preparation Reagent" kit (Applied Biosystems) following the manufacturer's guidelines.
Pasteurella multocida species specific PCR
The species specific primers designed by Townsend et al. [10] KMTIT7 and KMTISP6 were used to amplify the gene sequences in P.multocida. The PCR reaction mixture and the thermal cycle protocol were as follows: initial denaturation-94 oC for 5 min, followed by 30 cycles, each cycle consisting of 3 steps; denaturation at 95oC for 1 min; annealing at 55 oC for 1 min; extension at 72 oC for 1 min; final extension at 72 oC for 9 min
Detection of virulence associated genes by multiplex PCR
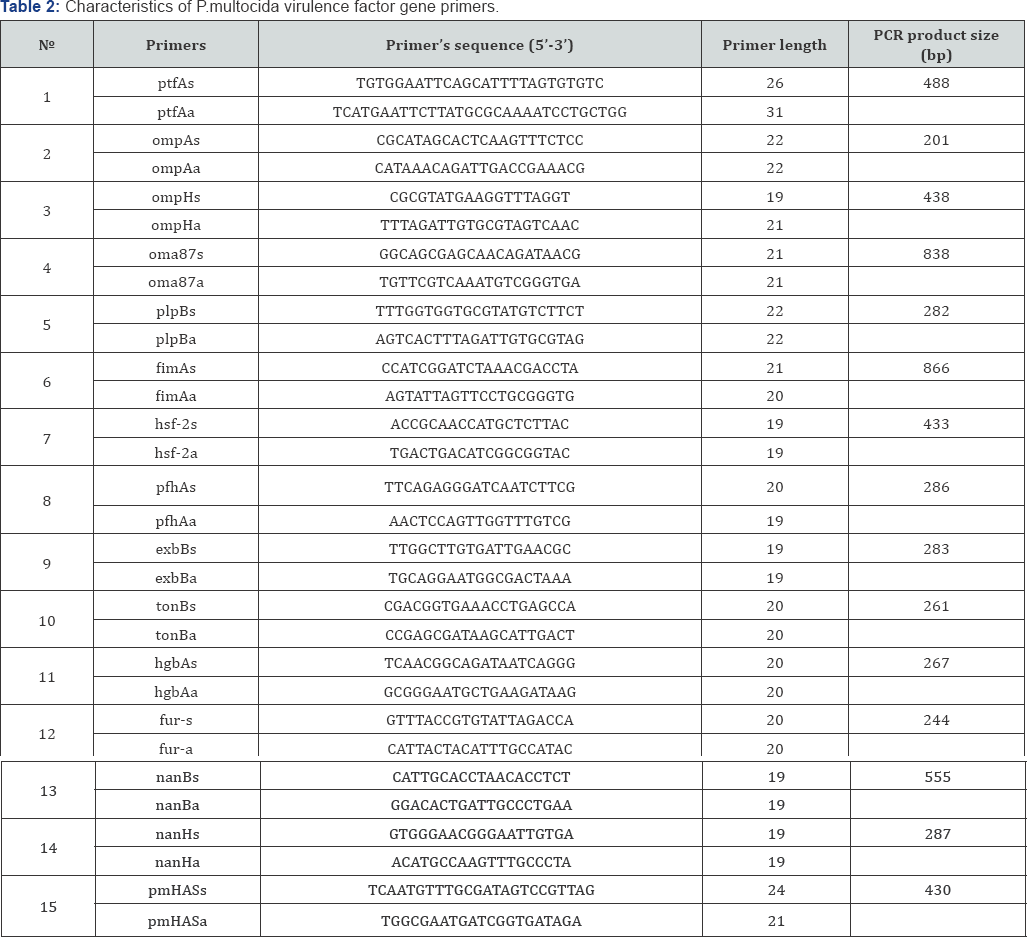
AccuPrime Taq DNA Polymerase High Fidelity kit manufactured by Invitrogen (USA) was used to produce PCR products of P.multocida virulent genes. Multiplex polymerase chain reaction (PCR) with 15 primer pairs was used for amplification (Table 2) [4,5,11,12].
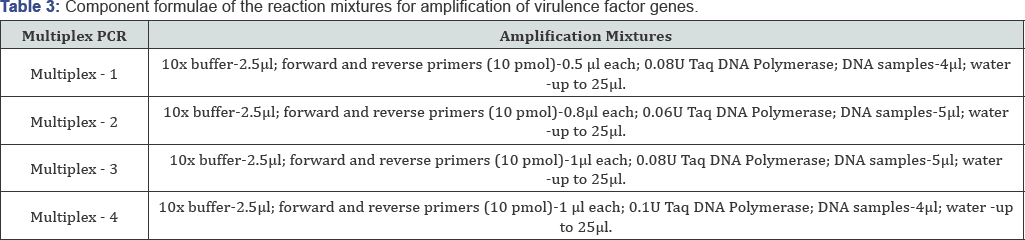
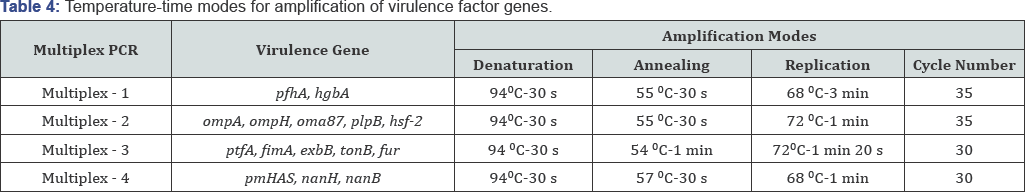
The parameters of the multiplex PCR that enable to detect mixtures and modes for amplification of virulence factor genes several virulence gene profiles at a time have been chosen. While were selected. The parameters of the multiplex PCR are shown performing multiplex PCR optimal component structures of PCR in Table 3 & 4.
DNA electrophoresis in 2% agarose gel was performed with TBE buffer containing 1 μl/ml ethydium bromide in the unit for electrophoresis of nucleic acids GNA-100, Pharmacia (Sweden).
Statistical Analysis
The resulted statistical data were processed with use of SPSS software for Windows 12.0. The diagrams were drawn with the help of Microsoft Excel 2010. Statistical analysis was performed using Fisher's exact test (two tailed) in the GraphPadPrism 6.0 software.
Results
All 25 samples taken from dead saigas in 2010 and 8 samples of year 2011 were identified as P.multocida. It has been shown that KMTI gene is specific for reference strains and all P.multocida isolates [10]. Size of the specific product obtained with use of primers KMTI77 and KMTISP6 is 460 bp. The results of the assay are shown on Figure 1.
As Figure 1 shows PCR fragments of KMTI gene sized 460 bp have been found in DNA of P.multocida isolates under study. So, the assayed isolates are of Pasteurella genus, multocida species. The mechanisms that take place in virulence factor genes of P.multocida provide a molecular basis for studying the degree of its pathogenicity. Therefore our task was to detect the presence of virulent genes in the genomes of P.multocida isolated from saiga antelopes. The results of the multiplex PCR for detection of virulent genes in P.multocida isolated from saiga antelopes in Kazakhstan are shown on Figure 2 & 3.
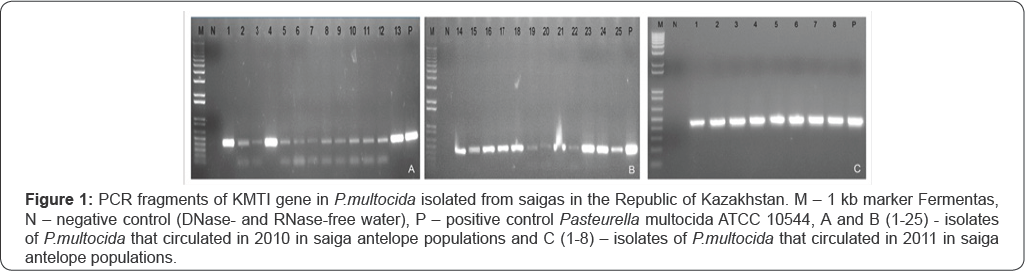
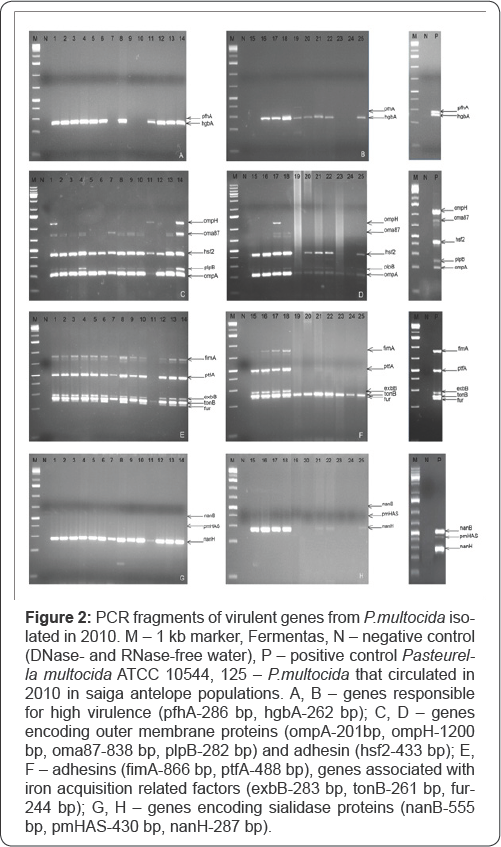
Figure 2 shows the PCR fragments of virulent genes encoding the outer membrane proteins, adhesins, iron acquisition related factors and sialidase proteins. HgbA-267 bp gene responsible for hemoglobin regulation in a body was not found in genomes of isolates Nos. 7, 9, 10, 15, 23, 24. Region of pfhA-286 bp gene encoding filamentous hemagglutinin was detected only in genomes of isolates Nos. 8, 11, 18, 21.
Figures 2C & 2D show PCR fragments of virulent genes encoding outer membrane proteins and adhesins of Pasteurella bacteria isolated from saiga antelopes in 2010. Region of OmpH gene encoding outer membrane protein H 1200 bp long was found only in genomes of 4 assayed isolates: 1, 11, 14 and 17. Fragment of the virulent Oma87 gene was detected in genomes of 13 isolates under study (1, 2, 3, 4, 7, 8, 9, 10, 13, 14, 16, 17, 18). Fragment of hsf2 gene sized 433 bp was present in genomes of all isolates except Nos. 19, 23 and 24. PlpB gene was detected in genomes of 11 isolates (1, 2, 3, 4, 7, 8, 9, 10, 13, 14, 18) and it was absent in 14 isolates (5, 6, 11, 12, 15, 16, 17, 19, 20, 21, 22, 23, 24 and 25). Fragment of OmpA gene encoding outer membrane protein A 201 bp long was detected in all isolates except Nos. 19 and 24.
Figures 2E & 2F show PCR fragments of virulent genes associated with iron acquisition related factors: exbB-283 bp, tonB-261 bp, fur-244 bp and adhesins: fimA-866 bp, ptfA-488 bp. As one can see the PCR fragments of virulent fimA, ptfA and exbB genes are present in genomes of 17 isolates except Nos. 11, 19, 20, 21, 22, 23, 24 and 25. PCR fragment of tonB gene sized 261 bp was detected in genomes of all isolates. PCR fragment of fur gene 244 bp long was found in genomes of 18 isolates and not detected in genomes of isolates 7, 11, 19, 20, 21, 23 and 24.
On Figures 2G & 2H PCR fragments of virulent genes encoding sialidase proteins (nanB-555 bp; pmHAS-430 bp; nanH-287 bp) are shown. We failed to find virulent nanB and pmHAS genes in genomes of all assayed isolates. Site of virulent nanH gene 287 bp long were present in all isolates except Nos. 11, 19, 20, 23 and 24.
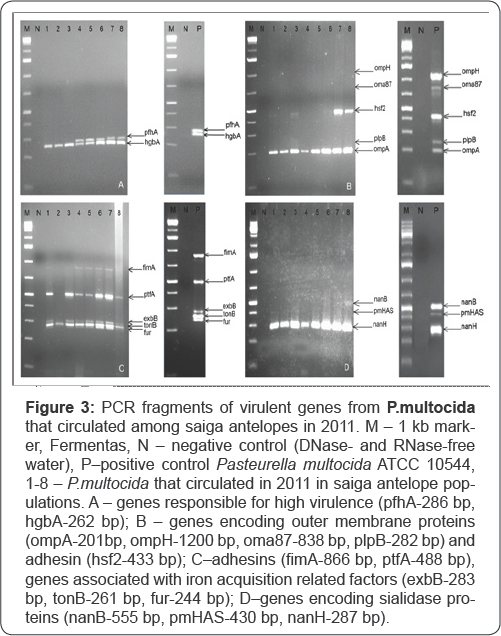
Figure 3A shows that PCR fragment of hgbA gene sized 267 bp has been found in the genomes of the assayed isolates. Product of pfhA gene 286 bp long was detected in genomes of isolates Nos. 4, 5, 6, 7, 8. The results of PCR for presence of ompH, oma87, plpB, hsf2, plpB and ompA genes (Figure 3B) show that ompH, oma87 and plpB genes are absent in P.multocida isolates. The region of hsf2 gene sized 433 bp was detected in genomes of isolates Nos. 1, 3, 6, 7, 8. OmpA gene encoding the outer membrane protein A was present in genomes of all isolates.
Figure 3C shows PCR products of fimA, ptfA, exbB, tonB, fur genes. TonB and exbB genes associated with iron acquisition related factors were present in genomes of all isolates whereas the fimA, ptfA and fur genes were not detected in the genome of isolate No. 2.
Only one gene (nanH) sized 287bp, of all genes encoding sialidase proteins was found in the assayed isolates (Figure 3D). In genomes of P.multocida isolated from dead saigas in 2011 nanB and pmHAS genes were absent. Table 5 shows quantitative content (%) of virulent genes in genomes of P.multocida isolated from saiga antelopes in years 2010 and 2011.
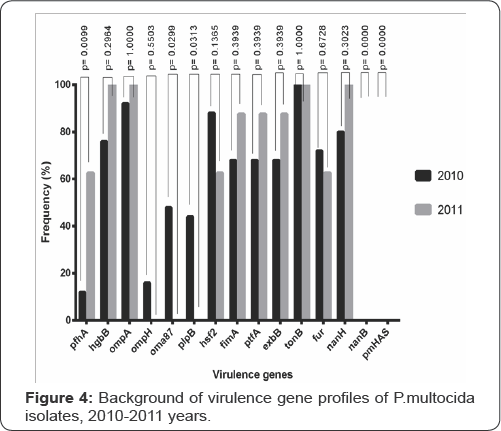
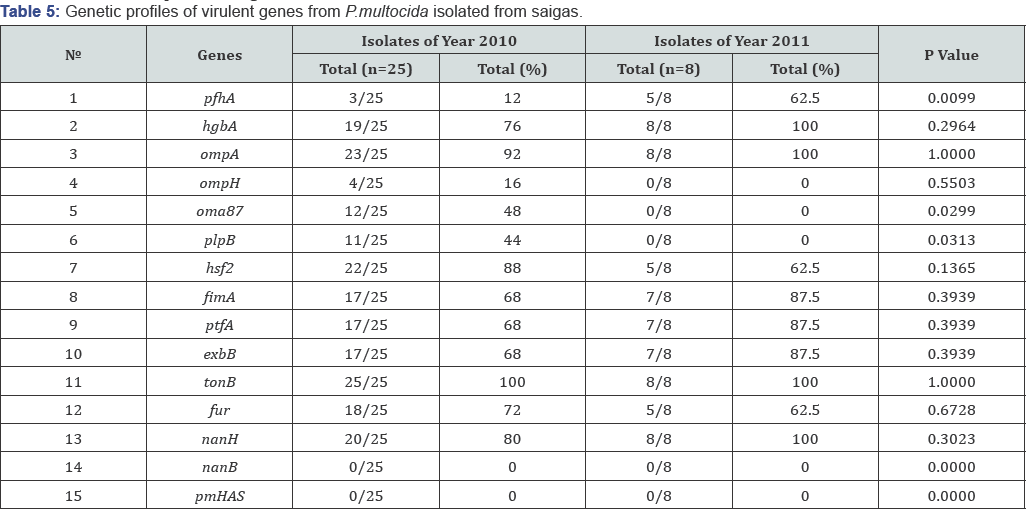
The study towards determination of virulent gene profiles of P.multocida isolated from dead saiga antelopes in 2G1G Table 5 has shown that hgbA, pfhA, ompA, ompH, oma87, plpB, hsf2, fimA, ptfA, exbB, tonB, fur, nanH genes are present in the genomes of the isolates while nanB and pmHAS genes are absent. On the basis of the obtained data (Figures 2,3 &5) the diagram has been plotted (Figure 4) to reflect the background of virulence gene profiles of P.multocida isolated from dead saigas in the Republic of Kazakhstan during years 2G1G-2G11.
Judging on the results of the comparative analysis of data plotted on diagram (Figure 4) the background of virulence gene profiles of P.multocida isolated from saiga antelopes in 2G1G include 13 genes while in P.multocida isolated from saigas in 2G11 only 1G genes were detected. In 2G11 sharp drop in mortality rate among saigas (only about 500 animals) versus year 2010 («12000 animals) was recorded maybe owing to reduction of the virulence background due to the influence of some biological, physical and chemical factors on the causative agent.
Percentage of virulent genes in the genomes of isolates under study appeared to be different, e.g., content of tonB gene was 100%, of fur gene-72%, of hgbB gene-76%, pfhA-12%, ompA- 92%, oma87-48%, plpB-44%, ompH-16%, nanH-80 %, and percentage of fimA, ptfA and exbB genes was 68%. Statistical analysis of data has been performed using Fisher's exact test and the resulted p values are shown in Table 4 and Figure 4. Differences in content of virulent pfhA, oma87 and plpB genes in P.multocida isolated from saiga antelopes in years 2G1G and 2011 were considerable (p<.05). As it concerns virulent hgbA, ompA, ompH, hsf2, fimA, ptfA, exbB, tonbB, fur and nanH genes the differences were insignificant (p>0.1).
Determination of the virulent gene profiles of P.multocida isolated from saigas in 2G11 showed 100% content of hgbA, ompA, tonB and nanH genes. Percentage of fimA, ptfA and exbB genes in genomes of P.multocida isolates was 87.5%, of genes pfhA, hsf2 and fur-62.5%. PlpB, oma87, ompH, nanB and pmHAS genes were absent.
Discussion
So far the principles of pathogenicity mechanisms and immune processes in P multocida are not known. The putative determinants of virulence such as genes encoding the outer membrane proteins, genes participating in iron transport, enzymes, toxins and adhesins play the certain role in these processes and their presence in P.multocida genomes is associated with virulence.
Some authors recommend to use hgbA gene as an epidemiological marker for characterization of bacterial strains and isolates of P multocida [4,13]. Content of the virulent hgbB gene in genomes of Kazakhstanean isolates from saigas in 2010 was 76% whereas in isolates of year 2011 it was 100%. A number of foreign scientists have shown 100% presence of the virulent hgbA gene in genomes of P multocida isolated from different animal species [7].
Filamentous hemagglutinin pfhA gene is used as an epidemiological marker in P multocida isolated from cattle. In our study only 12% of the assayed isolates of year 2010 contained the virulent pfhA gene whereas its content in the isolates of year 2011 was 62.5% (p<0.05). Adler et al. [14] paid special attention to detection of surface antigens participating in pathogenesis of P multocida. They extracted the protective protein of outer membrane oma87 that participates in export of other virulence factors from a bacterial cell. Our studies show 48% content of oma87 gene in genomes of Kazakhstanean isolates of P multocida from saiga antelopes in 2010 and its absence in isolates of year 2011 (p<0.05).
Also the assayed Kazakhstanean isolates of P multocida showed low content of ompH gene, e.g. only 16% of P multocida isolated in 2010 contained this gene, and in isolates of year 2011 this gene was not detected. One more gene encoding the outer membrane protein is ompA gene. As Table 3 shows content of this gene in the genomes of Kazakhstanean isolates of P multocida is high. In isolates of year 2010 the percentage of this gene is 92%, and in isolates of year 2011 it is 100%.
According to Lu YS et al. [15] the protein encoded by ompA gene as well as the protein encoded by ompH gene is protective, that is why the results of molecular and genetic analysis are significant in studying pathogenesis of P multocida.
Content of plpB gene encoding outer membrane lipoprotein in the bacteria isolated in 2010 was 44%, whereas in P multocida isolates of year 2011 it was absent (p<0.05). Adhesins are essential to colonization and invasion of the host by P multocida and as a rule their presence on bacterial surface is associated with virulence [3]. They include fimA (fimbrial), hsf-1 and hsf-2 (autotransporter adhesins), pfhA (filamentous hemagglutinins), ptfA (subunits of type IV pili) genes. Out of all isolates of year 2010 68% contained virulent fimA gene and it was detected in 87.S% of isolates of year 2011. Occurrence of virulent ptfA gene in genomes of P muItocida isolated in Kazakhstan in years 2010 and 2011 was respectively 68% and 87.5%. Content of the virulent hsf-2 gene in genomes of Kazakhstanean isolates of years 2010 and 2011 was respectively 88% and 62.5%. These data are close to the results of other authors [6].
The virulent tonB gene was detected in all assayed Kazakhstanean isolates of P muItocida. Content of the virulent exbB gene in isolates of year 2010 was 68%, and in isolates of year 2011 it was 87.S%. Virulent fur gene was present respectively in 72% and 62.5% of P muItocida isolated in years 2010 and 2011.
Sialidase proteins encoded by nanH and nanB genes can localize on the surface of host cells and destroy the receptors of the immune system [16,17]. In our study virulent nanH gene was detected respectively in 80% and 100% of bacterial genomes of P.multocida isolated in years 2010 and 2011. Virulent nanB gene was absent in genomes of Kazakhstanean isolates. Virulent pmHAS and nanB genes were not found in genomes of P multocida isolated from saiga antelopes in Kazakhstan in years 2010 and 2011.
References
- Grachev Y, Bekenov A (2010) Mass mortality among saigas in Kazakhstan: 12,000 dead. Saiga News 11: 2-3.
- Orynbayev MB, Rystayeva RA, Kerimbayev AA, Kospanova MN, Kopeev SK, et al. (2G13) Cases of Mass Mortality Among Saiga Antelopes of the Ural Population in Kazakhstan. Topical issues of Veterinary Biology 1(17): 20-26.
- Chung JY, Wilkie I, Boyce JD, Townsend KM, Frost AJ, et al. (2001) Role of Capsule in the Pathogenesis of Fowl Cholera Caused by Pasteurella multocida Serogroup A. Infect Immun 69(4): 2487-2492.
- Ewers C, Lübke A, Bethe A, Kiebling S, Filter M, et al. (2006) Virulence genotype of Pasteurella multocida strains isolated from different hosts with various disease status. Vet Microbiol 114(3-4): 3G4-317.
- Sayegh J, Atashpaz S, Hajazi MS (2008) Virulence genes profile and typing of ovine Pasteurella multocida. Asian J Anim Vet Adv 3(4): 206213.
- Ferreira TS, Felizardo MR, Sena de Gobbi DD, Gomes CR, et al. (2012) Virulence genes and antimicrobial resistance profiles of Pasteurella multocida strains isolated from rabbits in Brazil. Scientific World J 38(1): 58-67.
- Furian T, Borges K, Rocha S, Rodrigues E, Nascimento V, et al. (2013) Detection of virulence-associated genes of Pasteurella multocida isolated from cases of fowl cholera by multiplex-PCR. Pesq Vet Braz 33(2): 177-182.
- Fuller TE, Kennedy MJ, Lowery DE (2000) Identification of Pasteurella multocida virulence genes in a septicemic mouse model using signature-tagged mutagenesis. Microb Pathog 29(1): 2S-38.
- Harper M, Boyce JD, Adler B (2006) Pasteurella multocida pathogenesis: 125 years after Pasteur. FEMS Microbiol 265(1): 1-10.
- Townsend KM, Frost AJ, Lee CW, Papadimitriou JM, Dawkins HJ (1998) Development of PCR assays for species and type - specific identification of Pasteurella multocida isolates. Clin Microbiol 36(4): 1096-1100.?
- Doughty SW, Ruffolo CG, Adler B (2000) The type 4 fimbrial subunit gene of Pasteurella multocida. Vet Microbiol 72(1-2): 79-90.
- Tang X, Zhao Z, Hu J, Wu B, Cai X, et al. (2009) Isolation, antimicrobial resistance, and virulence genes of Pasteurella multocida strains from swine in China. J Clin Microbiol 47(4): 951-958.
- Roehrig SC, Tran HQ, Spehr V, Gunkel N, Selzer PM, et al. (2007) The response of Mannheimia haemolytica to iron limitation: implications for the acquisition of iron in the bovine lung. Vet Microbiol 121(3-4): 316-329.
- Adler B, Chancellor R, Homchampa P, Hunt M, Ruffolo C,et al. (2006) Immunity and vaccine development in Pasteurella multocida infections. J Biotechnol 44(1-3): 139-144.
- Lu YS, Lai WC, Pakes SP, Nie LC (1991) A monoclonal antibodies against a Pasteurella multocida outer membrane protein protects rabbits and mice against Pasteurellosis. Infect Immun 59(1): 172-180.
- Dabo SM, Taylor JD, Confer AW (2007) Pasteurella multocida and bovine respiratory disease. Anim Health Res Rev 8(2): 129-150.
- Tang X, Zhao Z, Hu J, Wu B, Cai X, et al. (2009) Isolation, antimicrobial resistance, and virulence genes of Pasteurella multocida strains from swine in China. J Clin Microbiol 47(4): 951-958.






























