Protein Sequence Textuary of Biomedical Images Based on Protein Modelization of MRI
Mohammad Reza Dawoudi1* and Attila Gyenesei2
1Salo Nursing Campus, Turku University of Applied Sciences, Finland
2The Finnish Microarray and Sequencing Centre, Finland
Submission: August 17, 2017; Published: August 29, 2017
*Corresponding author: Rafiul Amin Laskar, Mutation Breeding Lab., Department of Botany, Aligarh Muslim University, Aligarh, 202002, UP, India, Email: rafihkd@gmail.com
How to cite this article: Mohammad Reza D, Attila G. Protein Sequence Textuary of Biomedical Images Based on Protein Modelization of MRI. Curr Trends Biomedical Eng & Biosci. 2017; 8(2): 555731. DOI: 10.19080/CTBEB.2017.08.555731.
Abstract
Serving an image as a text is one of the items on the data-base. A variety of methods could be employed. In this paper we have proposed a novel method for converting medical images into textuary formats. Through this method it is possi-ble to create protein sequences of the current biomedical medical images. This is achieved using the protein modeliza-tion of digital medical images. Protein modelization of a digital image is an arrangement for allowing numbers of pix-els values to be presented as sequences of characters. In this system every pixel values of color scale is procedure into four protein characters. One advantage of protein sequence textuary compared to other image serving methods is the browsing, retrieving and sequence blasting from database.
Introduction
This author kit is designed to assist you in preparing your submission. It is an exact representation of the format expected by the editor. A template in Microsoft Word is availa-ble on the Medinfo 2013 World Wide Web server at http://www. medinfo2013.dk. This example provides all the information you require to comply with the Medinfo 2013 submission standards for Scientific demonstrations. The official language of the Medinfo 2013 Congress in Co-penhagen, Denmark is English. Values should be in Interna-tional System units.
To ensure that the greatest opportunity for participation in Medinfo 2013 is made available to researchers and institu-tions, authors may appear in multiple submissions but may have only ONE submission per category considered where they are the primary or first author [1-6].
Materials and Methods
Protein Modelization of a digital image is an arrangement for allowing numbers of pixels values and pixels maps to be pre-sented as sequence of protein characters. The possible pixel value is arranged from 0 to 255 and the permutations with repetitions of four letters are 256, therefore every pixel values could be procedure into four letters. The first step is to carry out the direct coding of the digital image, with the four letters A, G, C, and T. Alphabet {A, C, G, T} is chosen, because it is the same as in real DNA sequences. The second stage is translation DNA sequence to protein sequence. This is per-formed with translated tools which allow the translation of DNA sequences to protein sequences. ExPASy [7] and TTans-late Nucleic Acid Sequence Tool [8] are examples of free online translation tools. Last section will execute with multi-ple sequence alignment. The results will be obtained by the statistical significance of match's ratio. The similar image could be finding by corresponding degree of similarity (match's ratio) with image list.
Experiment Results

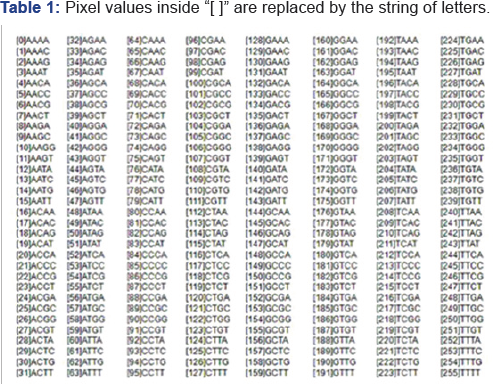
In this section we present the experiment result of protein modelization using Translate tool and Multiple Sequence Alignment by CLUSTALW. The experiment performs protein modelization of the eight real lateral ventricles (brain) images (Image). In the first stage the pixels values of each images converted into the virtual DNA sequences. These conversions should be done based on Table 1. The second stage is performed with DNA translation in to protein sequences using translated tools. The statistical signif-icance of matches is calculated by sequence alignments. There are various algorithms and tools could be implemented. The DNA and protein sequences of images are shown in Ap-pendix I and II.
Multiple sequence alignment by CLUSTALW
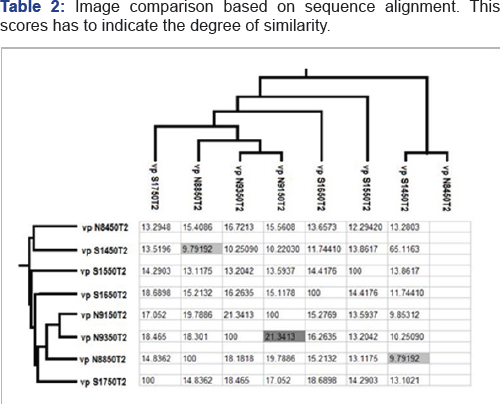
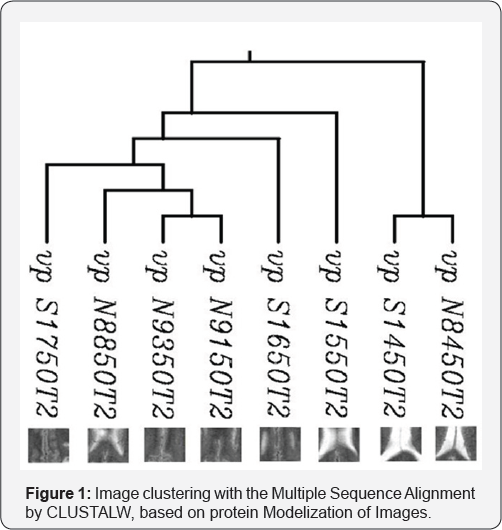
Multiple sequence alignment tools determine the ratio of sim-ilarity between protein sequences. The ratio of similarity be-tween eight protein sequences has computed using CLUSTALW. The ratios of similarity between sequences (sta-tistical significance of matches) are shown in Table 2. The ratio of distance between eight protein sequences has computed using CLUSTALW, and the below graph has been obtained (Figure 1).
Results
The sequence alignments are executed to detect percentage of similarity ratio in virtual protein sequences. Based the degree similarity between virtual protein sequences we can categories the images. In every category a collection of objects are similar and according to a given score we can identify the group of objects which are "close" to this score.
Acknowledgement
This article was carried out at The Finnish Microarray and Se-quencing Centre (FMSC) in Turku, Finland at the year 2009 and 2010. Special thanks go to head of the centre Dr. Attila Gyenesei for providing me the possibility to do my research on this significant subject.
References
- Barnard K, Shirahatti N. A method for comparing content based image retrieval methods.
- Marqes O, Mayron L. System and methods of image retrieval. Patent application. USPC Class: 382305.
- http://www.searchenginejournal.com/a-look-into-reverse-image- search-tools/14666/trackback
- http://www.searchenginejournal.com/a-look-into-reverse-image- search-tools/14666/trackback/
- http://au.expasy.org/tools/dna.html
- http://align.genome.jp/
- http://au.expasy.org/tools/dna.html
- http://biotools.umassmed.edu/cgi-bin/biobin/transeq






























