Allosteric regulation in drug design
Ashfaq Ur Rehman1,2*, Shah Saud3, Nasir Ahmad4, Abdul Wadood2and R Hamid5
sup>1State Key Laboratory of Microbial Metabolism, Department of Bioinformatics and Biostatistics, China
2Department of Biochemistry, Abdul Wali Khan University Mardan, Pakistan
3Laboratory of Analytical Biochemistry and Bio separation, Shanghai Jiao Tong University, China
4Department of Chemistry, Islama College University Peshawar, Pakistan
5Department of Bioinformatics, Muhammad Ali Jinnah University Islamabad, Pakistan
Submission: May 02, 2017; Published: May 23, 2017
*Corresponding author: Ashfaq Ur Rehman, State Key Laboratory of Microbial Metabolism, Department of Bioinformatics and Biostatistics, Shanghai Jiao Tong University, 800 Dongchuan Road, Shanghai 200240, China, Tel: 86-13162094886; Fax: 86-21-34204348; Email: raysjtu@sjtu.edu.cn
How to cite this article: Ashfaq U R, Shah S, Nasir A, Abdul W, R Hamid. Allosteric regulation in drug design. Curr Trends Biomedical Eng & Biosci. 2017; 4(1): 555630. DOI:10.19080/CTBEB.2017.04.555630.
Abstract
Protein and enzymes play significant roles in biological processes of all living organisms; their functions are regulated through allosteric mechanism, which are initiated through attachment of ligand or inhibitors with the protein or enzymes other than active (orthosteric) sites. This mini review involved mechanism, types and importance of allosteric regulations in drug design process.
Keywords: Allosteric, Activator: Drug design
Introduction
For the survival of all organisms the significance of protein function is pivotal. As all the cell processes are under carful control and if not properly controls this leads to the abnormality and ultimately cause disease. While various biological processes expressed the control at different points in life time of protein included regulation of gene expression, translation into protein through control of activity and at last degradation of protein [1].
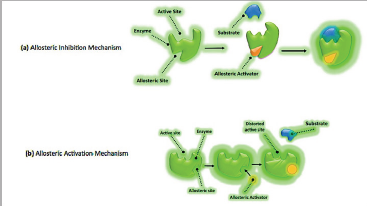
This is a mechanism of protein function, which occurs at one site of protein structure while its impact is on other site of protein. For example, the attachment of the ligand or mutation of amino acid in one allosteric site changes the catalytic activity of at far orthosteric site or a second binding site. The ligand is called effector; it may be another protein while its binding site is other than orthosteric site; here we called it allosteric site. Two possibilities came out from ligand binding either these will enhance the function of protein called allosteric activators or do inhibition called allosteric inhibitors, as shown in Figure 1. Mechanism of allostery and its Flsexibility key for allostery.
Allostery is the process by which biological macromolecules; commonly proteins, relay the effect of binding at one to another site, often distal, functional site, allowing for regulation of activity. The allosteric trepidation related to non-ligand sources includes mutation of amino acid and covalent modification at an allosteric site. By this definition allostery is an intrinsic property of all
proteins. All the surfaces of proteins are susceptible to ligand binding and mutation to cause allostery. All protein surfaces are then potential allosteric sites subject to ligand binding or to single or multiple mutations that might introduce the property of allostery. Whereas the lack flexibility in fibrous, structural proteins leads to the uniform and stable conformations which stopped the allostery [2,3].
The flexibility of protein is the only characteristic which leads to the production of allostery and catalysis that in turn produced large number of conformations which are interconvert able with different time levels. The allosteric ways regulate both enzymes and the non-catalyticprotein There are various types of allosteric regulation which includes binding of small molecules to the allosteric site through opening and closing of orthosteric sites, changes involved in conformations of orthosteric site, changes to electrostatic properties of orthosteric site, allosteric control of complex formation, allosteric regulation by phosphorylation, binding of protein, regulation of making and breaking of disulfide bond and regulation through changes in protein flexibility [4].
Allosteric regulation rule in drug discovery
The pharmacological drug design can be developed through deep understanding of mechanism of allostercity. The main reason is the when the ligand bind with the allosteric site it completely change function of protein and this site can be used as drug target for novel compounds. The conservation of the allosteric site is common because the evolution has little impact on it and that is the main reason for selective inhibitors across the species [5,6]. In the field of chemical biology and designing of new class of antibiotics the species-specific inhibitors plays important role. The allosteric inhibition has been deeply studied for kinases, G protein-coupled receptors and ligand-gated ion channels [7-10].
Conclusion
The need for drugs with rare side effects cannot be overstressed. Today, most drugs modify the actions of enzymes and other molecules by directly binding to their orthosteric sites. Nevertheless, orthosteric site configuration is similar in several proteins performing related functions and this leads to a lower specificity of a drug for the desired protein. Consequently, such drugs may have adverse side effects. A new basis of drug discovery is emerging based on the binding of the drug molecules to sites away (allosteric) from the orthosteric sites. It is possible to find allosteric sites, which are unique and hence more specific as targets for drug discovery. It is predicted that the drug discovery exploiting allosteric sites will lead to more effective therapeutic agents with rare side effects.
References
- AB Pardee (2006) Regulatory molecular biology. Cell Cycle 5(2006): 846-852.
- Nina MG, Stephen JB (2008) Nature Chemical Biology 4: 474-482.
- Monod J, Wyman, J Changeux JP (1965) On the nature of allosteric transitions: a plausible model. J. Mol. Biol. 12, 88-118.
- Kuriyan J, Eisenberg D (2007) The origin of protein interactions and allostery in colocalization. Nature 450: 983-990.
- Salvesen GS, Riedl SJ (2007) Caspase inhibition, specifically. Structure 15: 513-514.
- Schweizer A, Roschitzki-Voser H, Amstutz P, Briand C, Gulotti-Georgieva M, et al. (2007) Inhibition of caspase-2 by a designed ankyrin repeats protein: specificity, structure, and inhibition mechanism. Structure 15(5): 625-636.
- Bertrand D, Gopalakrishnan M (2007) Allosteric modulation of nicotinic acetylcholine receptors. Biochem Pharmacol 74: 1155-1163.
- Raddatz R, Schaffhauser H, Marino MJ (2007) Allosteric approaches to the targeting of G-protein-coupled receptors for novel drug discovery: a critical assessment. Biochem Pharmacol 74(3): 383-391.
- Ross RA (2007) Allosterism and cannabinoid CB1 receptors: the shape of things to come. Trends Pharmacol Sci 28: 567-572.
- Shi Z, Resing KA, Ahn NG (2006) Networks for the allosteric control of protein kinases. Curr Opin Struct Biol16: 686-692x.






























