Sorting Out Non-Synonymous Single Nucleotide Polymorphism Leads to Novel Biomarker Discovery for Disease Prognosis
Mohammad Uzzal Hossain1, Keshob Chandra Das2, U.S. Mahzabin Amin2, Chaman Ara Keya3 and Md. Salimullah2*
1Bioinformatics Division, National Institute of Biotechnology, Ganakbari, Ashulia, Savar, Dhaka-1349, Bangladesh
2Molecular Biotechnology Division, National Institute of Biotechnology, Ganakbari, Ashulia, Savar, Dhaka-1349, Bangladesh
3Department of Biochemistry and Microbiology, North South University, Bashundhara, Dhaka-1229, Bangladesh
Submission: February 28, Published: April 24, 2017
*Corresponding author: Md. Salimullah, Director General (Additional Charge) & Chief Scientific Officer, National Institute of Biotechnology, Ganakbari, Ashulia, Savar, Dhaka-1349, Bangladesh, Tel: 88027788443; Fax: 880-2-7789636; Email; salim2969@gmail.com
How to cite this article: Hossain MU, Das KC, Amin USM, Keya CA, Salimullah M. Sorting Out Non-Synonymous Single Nucleotide Polymorphism Leads to Novel Biomarker Discovery for Disease Prognosis. Curr Trends Biomedical Eng & Biosci. 2017; 3(2): 555608. DOI: 10.19080/CTBEB.2017.03.555608
Abstract
Hereditary genetic variation which is considered to be primarily caused by single nucleotide polymorphism (SNP), is a significant drawback for developing universal therapy against diseases. Among others, non-synonymous SNP (nsSNP) could be fatal due to its effect on structure and function of the ultimate gene product. Therefore, study of functional nsSNP's would provide an insight into the exact cause underlying the onset of genetic variation and possible methodologies for the cure or early management of the disease. Various in silico tools could be employed to screen and map the deleterious nsSNP's to the protein structure for predicting the structure-function effects. Further, these nsSNPs upon experimental verification would be ideal candidate for the disease risk assessment. Positive linkage study would enforce novel biomarker discovery for specific disease prognosis.
Rationale
Extensive effort was given over the past years on revealing how genetic changes give upsurge to the molecular effects that cause diseases and phenotypes [1],2]]. These efforts enforced the growth to number of databases, web resources, and tools for prioritizing possible single nucleotide polymorphisms (SNPs). These resources and online tools are designated on the basis of their genomic context as well as annotations. Till now, most of the focus is on human genome annotations, although some resources provide insight into SNP data from model organisms such as mouse, fruit fly, or chimpanzee [3]]. Typically, SNP data is used as a marker in the context of a linkage or population-based association study. However, there are a number of challenges such as the position, expression of functional products (RNA, Protein), and experimental validation of target SNP etc. to identify these so-called functional variants. Therefore, systematic in silico approach is needed to reduce the burden of scrutinizing a large number of SNPs available for a target disease. The approach might concentrate in three important areas: identification of candidate genes that may have causal variants, selection of candidate causal SNPs and focus on nsSNP's (Figure 1). We suggest targeting nsSNPS as these polymorphisms are most likely to affect the functionality of the target gene product. In this regard, bioinformatics tools can play a pivotal role to identify specific disease related nsSNP through structural and functional assessment.
Bioinformatics Tools and Resources for nsSNP�x2019;s Discovery and Analysis
Generally, the discovery and prioritization of SNPs are carried out by sequencing. SNPs discovery based on the various sites isolating from the sequence, assesses frequency of the error in total numbers of the selected sequences, isolates parologous and then determines genotype. In silico approaches play an important role in SNP discovery and scrutiny. These methods mark genes that encompass SNPs, let researchers to retrieve data about SNPs based on gene of interest, genetic or physical map location, or expression pattern.
The polymorphism data is available from several databases such as NCBI dbSNP database (http://www.ncbi.nlm.nih.gov/SNP/), the Ensemble genome browser (www.ensembl.org/), and the UniProt database (www.uniprot.org). The NCBI dbSNP database is the most extensive SNP database among the others, but it contains both validated and non validated polymorphisms. There are now many databases that provide access to SNP or disease mutation data. Many genotype-phenotype databases are available as well including the Human Gene Mutation Database (HGMD, http://www.hgmd.cf.ac.uk/ac/index.php) [4], Online Mendelian Inheritance in Man (OMIM, http://www.ncbi.nlm.nih.gov/sites/entrez?db=omim) [5], the Pharmacogenetics Knowledge Base (PharmGKB, http://www.pharmgkb.org/) [6], database of Genotype and Phenotype (dbGAP, http://www.ncbi.nlm.nih.gov/sites/entrez?db=gap) [7] etc. There are also a growing number of databases of resequencing polymorphism data including the SeattleSNPs project (http://pga.mbt.washington.edu/) and sequencing of somatic mutations in cancer [8,9]. This has led to a wealth of genetic variation data.
Besides, there are some tools and softwares which could provide meaningful insights of target polymorphisms (Table 1). These tools could be utilized for prioritization of SNPs based on their functionality and stability. SNPs with the potential to cause structural modifications due to the amino acid substitution as well as their functional abnormality could also be predicted with these tools (Figure 1, Table 1). In this scenario, nsSNP would be the best choice to study as it could show the most deleterious effect of polymorphism [5].
infection received peg-interferon and ribavirin treatment for 48 weeks, out of nine patients showed Resistance to the treatment. Blood sampling were made on at start and end of the treatment. Based on the therapeutic response to antiviral treatment, those 18 patients could divide into two groups: Treated (Responder, R) 9 patients, and Resistant (Non-responder, NR) 9 patients.

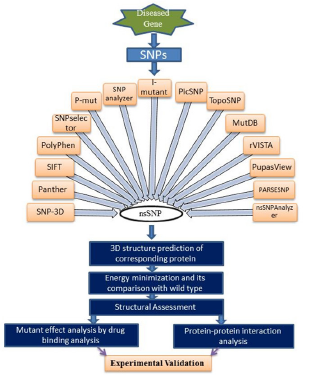
Experimental validation of target nsSNP

After the identification of possible non synonymous polymorphism researcher can employ these nsSNP's into experimental validation for the novel biomarker discovery Various methods may be considered for the experimental validation of the nsSNP in specific disease (Figure 2) [4]. If the verification and statistical output of selected nsSNP gives the frequent occurrence in the specific region of the target gene, it could be evaluated as a biomarker for disease risk assessment. Furthermore, linkage or population based association study should also be performed to declare a nsSNP as biomarker.
nsSNPs: Future Biomarkers?
The relationship between SNPs and various diseases, has long been established with a wide range of human diseases resulting from different nsSNP's. Particular population or individual's disease susceptibility, severity of illness etc depend on those nsSNP�x2019;s. Also, nsSNP's contributed to individual�x2019;s response to drug, drug resistance etc. However, the establishment of association of nsSNP's with diseases is going with slow pace due to the lack of their proper identification. Therefore, the overwhelming task of characterization of 8.2 million SNPs in Human genome [28] for disease association, bioinformatics tools together with wet laboratory research could be the best option for the obvious future biomarker ‘nsSNP’ development.
References
- Mooney S (2005) Bioinformatics approaches and resources for single nucleotide polymorphism functional analysis. Brief Bioinform 6(1): 44-56.
- Ng PC, Henikoff S (2006) Predicting the effects of amino Acid substitutions on protein function. Annu Rev Genomics Hum Genet 7: 61-80.
- Steward RE, MacArthur MW, Laskowski RA, Thornton JM (2003) Molecular basis of inherited diseases: a structural perspective. Trends Genet 19(9): 505-513.
- Cooper DN, Stenson PD, Chuzhanova NA (2006) The Human Gene Mutation Database (HGMD) and its exploitation in the study of mutational mechanisms. Curr Protoc Bioinformatics doi: 10.1002/0471250953.bi0113s12.
- Hamosh A, Scott AF, Amberger J, Valle D, McKusick VA (2000) Online Mendelian Inheritance in Man (OMIM). Hum Mutat 15(1): 57-61.
- Altman RB (2007) Pharm GKB: a logical home for knowledge relating genotype to drug response phenotype. Nat Genet 39(4): 426.
- Mailman MD, Feolo M, Jin Y, Kimura M, Tryka K, Bagoutdinov R, et al. (2007) The NCBI dbGaP database of genotypes and phenotypes. Nat Genet 39(10): 1181-1186.
- Sjoblom T, Jones S, Wood LD, Parsons DW, Lin J, et al. (2006) The consensus coding sequences of human breast and colorectal cancers. Science 314(5979): 268-274.
- Greenman C, Stephens P, Smith R, Dalgliesh GL, Hunter C, et al. (2007) Patterns of somatic mutation in human cancer genomes. Nature 446(7132): 153-158.
- Sunyaev S, Ramensky V, Koch I, Lathe W, Kondrashov AS, et al. (2001) Prediction of deleterious human alleles. Hum Mol Genet 10(6): 591¬597.
- Ng PC, Henikoff S (2003) SIFT: Predicting amino acid changes that affect protein function. Nucleic Acids Res 31(13): 3812-3814.
- Ferrer-Costa C, Gelpi JL, Zamakola L, Parraga I, de la Cruz X, et al. (2005) PMUT: a web-based tool for the annotation of pathological mutations on proteins. Bioinformatics 21(14): 3176-3178.
- Mooney SD, Altman RB (2003) MutDB: annotating human variation with functionally relevant data. Bioinformatics 19(14): 1858-1860.
- Xu H, Gregory SG, Hauser ER, Stenger JE, Pericak-Vance MA, et al. (2005) SNPselector: a web tool for selecting SNPs for genetic association studies. Bioinformatics 21(22): 4181-4186.
- Yue P, Melamud E, Moult J (2006) SNPs3D: candidate gene and SNP selection for association studies. BMC Bioinformatics 7: 166.
- Stitziel NO, Binkowski TA, Tseng YY, Kasif S, Liang J (2004) topoSNP: a topographic database of non-synonymous single nucleotide polymorphisms with and without known disease association. Nucleic Acids Res 32(Database issue): D520-D522.
- Chang H, Fujita T (2001) PicSNP: a browsable catalog ofnonsynonymous single nucleotide polymorphisms in the human genome. Biochem Biophys Res Commun 287(1): 288-291.
- Karchin R, Diekhans M, Kelly L, Thomas DJ, Pieper U, et al. (2005) LS- SNP: large-scale annotation of coding non-synonymous SNPs based on multiple information sources. Bioinformatics 21(12): 2814-2820.
- Ryan M, Diekhans M, Lien S, Liu Y, Karchin R (2009) LS-SNP/PDB: annotated non-synonymous SNPs mapped to Protein Data Bank structures. Bioinformatics 25(11): 1431-1432.
- Zhao T, Chang LW, McLeod HL, Stormo GD (2004) PromoLign: a database for upstreamregion analysis and SNPs. Hum Mutat 23(6):534-539.
- Marinescu VD, Kohane IS, Riva A (2005) MAPPER: a search engine for the computational identification of putative transcription factor binding sites in multiple genomes. BMC Bioinformatics 6: 79.
- Loots GG, Ovcharenko I (2004) rVISTA 2.0: evolutionary analysis of transcription factor binding sites. Nucleic Acids Res 32(Web Server issue): W217-W221.
- Capriotti E, Fariselli P, Casadio R (2005) I-Mutant2.0: predicting stability changes upon mutation from the protein sequence or structure. Nucleic Acids Res 33(Web Server issue): W306-W310.
- Conde L, Vaquerizas JM, Ferrer-Costa C, de la Cruz X, Orozco M, et al. (2005) PupasView: a visual tool for selecting suitable SNPs, with putative pathological effect ingenes, for genotyping purposes. Nucleic Acids Res 33(Web Server issue): W501-W505.
- Taylor NE, Greene EA (2003) PARSESNP: A tool for the analysis of nucleotide polymorphisms. Nucleic Acids Res 31(13): 3808-3811.
- Bao L, Zhou M, Cui Y (2005) nsSNPAnalyzer: identifying disease- associated nonsynonymous single nucleotide polymorphisms. Nucleic Acids Res 33(Web Server issue): W480-W482.
- Mi H, Huang X, Muruganujan A, Tang H, Mills C, et al. (2017) PANTHER version 11: expanded annotation data from Gene Ontology and Reactome pathways, and data analysis tool enhancements. Nucleic Acids Res 45(D1): D183-D189.
- Zhao Z, Zhang F (2006) Sequence context analysis of 8.2 million single nucleotide polymorphisms in the human genome. Gene 366(2): 316¬324.






























