Current and Future Applications for DNA Sequence for Forensics, Biosurveillance, Clinical Health and Detection
John P Jakupciak*
Cipher Systems, USA
Submission: January 24, 2017; Published: February 06, 2017
*Corresponding author: John P Jakupciak, Institute of Analytical Sciences, 2661 Riva Rd Ste 1000 Fl 5, Annapolis, MD 21401, USA, Email: Johnjakupciak@cipher-sys.com
How to cite this article: John P J. Current and Future Applications for DNA Sequence for Forensics, Biosurveillance, Clinical Health and Detection. Curr Trends Biomedical Eng & Biosci. 2017; 1(3): 555561. DOI: 10.19080/CTBEB.2017.01.555561
Abstract
Many DNA/RNA/protein analytical methods are currently available, ranging from the standard culturing methods, which are tedious, slow, and dependent on achieving culture of the agent, to simplistic single biomarker genomic based tests, to high resolution DNA sequencing, which for a few unique techniques, provide identification of metagenomics samples at the strain level. The latter are faster than culture but perform in a near real-time response window. Real-time PCR limited detection methods are restricted to multiplex identification of few bioagents and test results contain the potential of error due to innate amplification errors, coupled with the requirement to know in advance what the agent is in the sample prior to design primers (probes) for detection. Rapid direct sequencing, coupled with data base matching, offers the most reliable, effective, reproducible, and cost effective approach to biological detection at strain level with details to measure major and minor population content. The evidence is now convincing, although a steep education curve is daunting to decision-maker acceptance, that strain to strain variation in genomic sequence renders probabilistic identification from direct sequence the method of choice for forensics, biosurveillance, clinical health and detection. Herein, the application of NGSbioinformatics tools for forensic analyses of bacterial samples was examined against specially prepared samples. These results were used to elucidate benefits, caveats, and potential pitfalls of direct-sequence analysis; revealed subtle errors in sequence information that are overlooked by the community and demonstrated utility of sequencing to match evolved populations back to its source.
Keywords: Biothreats, Infectious disease, Sequencing, Forensic science
Introduction
Specialists from the FBI, Army National Guard, US NORTHCOM, Department of State, experts at hospitals and universities, and key opinion leaders from the private sector all agree that the most effective approach for comprehensive genetic variation discovery is by high-throughput sequencing [1-3]. Genome identification via DNA sequencing is a concept advocated by DHS leaders, e.g., Tom Ridge [4] and other leaders [5].
Real time DNA sequencing is required for rapid genome (organism) identification. Sequencing methods are generally costly and have evolved from diagnostic applications. Critical to the success of biothreat surveillance is the ability to screen for and detect multiple agents rapidly in a single reaction [6]. Currently, there is intense research for new molecular detection technologies that could be used for rapid and very accurate detection to share and implement strategies for improved local research issues in detection of infectious diseases, identification, studying and then enhancing global disease surveillance for pathogen detection and response. Real-time sequencing can serve as the foundation to monitor health and detection capacity for environmental and clinical measurements of microbials, viruses, and biological signals. Such indicators provide incident information to local, state, and national decision makers, as a few examples [7].
The most effective criteria for the challenge to produce fast and reliable tools for the identification of traditional and novel (including genetically modified) pathogenic organisms in complex samples needs to be sequencing instrument specific and focus only on performance time and false positive and false negative outcome of the analysis results.
Prior studies examined implementation of whole genome sequencing (WGS) for genetic applications [8-10]. WGS has potential to greatly advance the precision of source attribution of microbial populations [11-13]. Before realizing any such advance for microbial forensics, it is critical to fully characterize the benefits, limitations, and proper protocols for application of WGS. Presented are several genomic characterization methods, along with critiques for improving forensic value of WGS analysis. Multiple strategies for genomic analysis and variant validation were conducted. To determine bacterial sample attribution to a source, twelve separate colonies of Bacillus anthracis strain Ames were cultured under stressful media conditions to establish samples possessing differing sets of mutations (subpopulations), bearing known lineages of descent. Analytical pipeline protocols were tested to assess their abilities to discern microbial sample relationships.
Findings
Sequencing of genomes from progenitor and descendent B. anthracis colony lineages revealed more sequence variants than expected. Phylogenetic relationships derived from SNP data frequently deviated from the known relationships between clones within the same lineage and with theprogenitor clones of the lineage. WGS bears great potential in microbial forensics. The major strengths of this forensic method are the non-arbitrary determinations of data validation and relatedness metrics, as well as the ability to compare microbialgenomes with or without a reference database of related genomes [13].
The ability to develop unique genomic signatures requires comparisonof pathogens of interest to all possible background signatures. The application of real-time sequencing is further challengeby the need to continually re-check uniqueness of all DNA or amino acid signatures as new genomes are added to private/public databases. The pace of genome sequencing continues to increase, making it essential that efficient algorithms, combined with powerful computing facilities, be applied to this problem in order to incorporate the latest genome data into the signatures. In this study, whole genome sequence analysis was investigated to characterize associations between closely related populations of known ancestry under controlled conditions. Study goals were to verify whether whole genome sequencing (WGS) genomic analysis was a reliable microbial forensic method for attributing relatedness, characterizing the extent of evolutionary mutations in Bacillus anthracis populations over time, and understanding the strengths and limitations of whole genome sequence analysis in a forensics context. We can compare the results to prior studies on infectious agents [11].
Although nucleotide and amino acid sequence based approaches have been used to infer microbial evolutionary relationships, over the last 19 years these methods have been increasingly used for typing and characterizing their populations [7,11-18]. Sequencing methods provide standardized and unambiguous data that are portable through web based databases with direct access to the information needed to identify and monitor emerging pathogenic agents [19-22]. More importantly, sequence data, unlike many other forms of molecular typing data, provide direct genealogical information that can be used efficiently to estimate phylogenetic relationships and parameters associated with population dynamics [23].
Use of biological agents, such as anthrax, presents unique challenges to the forensic investigator, since the genetic signature of the evidence is changeable rather than static. Understanding pathogen diversity in nature and under lab conditions is critical to improve forensics science and counter bioterrorism and distinguish outbreaks from intentional attacks. In these experiments, B. anthracis clones from a single source were cultured in parallel to evaluate the direction of mutation and test novel bioinformatics tools to link the end state, passaged materials back to the source. This study has a significant impact on bioforensics and the ability to use direct sequence analysis to provide a probabilistic description of major/minor population members in samples.
Bacillus anthracis, the etiologic agent of anthrax, [24,25] is a Gram positive spore forming bacterium. The organism is very closely related to low and non-virulent bacteria of the B. cereus sensu lato species. The primary virulence factors for mammalian pathogenesis are carried on horizontally transferrable plasmids pXO1 and pXO2 (lethal factor toxin and a poly-D-glutamic acid based capsule, respectively). Rare instances of close relatives carrying virulence determinants on plasmids have been reported. The B. anthracis species shows low sequence diversity, indicative of relatively recent global spread of a clone strain. There are a very limited number of unique chromosomal DNA targets, the most important being a group of defective prophages.
Whole genome sequencing revealed more than 3,500 SNPs among different strains of B. anthracis [26]. Evolutionary analysis revealed that many of the SNPs were evolutionarily stable. Due to the predominant sequencestability of SNP variants in B. anthracis, [27] developed a series of “canonical” SNPs to classify evolutionary relationships among 88 global isolates. Each canonical SNP was used to distinguish a separate node of a phylogenetic tree with which showed similar geographical distributions among related strains.
Another variant class sufficiently present in B. anthracis genomes is single nucleotide repeats (SNRs) [28] where selected SNR loci are used for multilocus variable-number tandem repeat analysis (MLVA). Since SNRs have the highest mutation rates for B. anthracis, SNR analysis has the greatest resolving power between closely related strains, but is not well suited for identifying phylogenetic relationships between distantly related strains [29].
Bioforensics analysis of B. anthracis may be improved in many ways. Current research priorities include increasing the number of genomes of B. anthracis to increase the reference sequence space for characterization, building of SNP trees establishing evolutionary relationships among variants, and determining the extent of population substructure and geographic associates to that substructure. New genomes should have extensive metadata on isolation date, geographic location, virulence and phenotypic traits. Very important to the further study of biothreat agent fauna is to move away from the paradigm of studying single microbial isolates. Diversity and evolution of microbial populations can be understood better through direct sequence analysis of entire populations.
A challenge to bacterial forensics is the ability to identify and differentiate between samples in a timely fashion without need of intensive resources. Possibly the greatest challenge is the need to have the proper background biological data collected to adequately analyze metagenomic sequence. Accurate characterization of a sample is dependent on accurate measurement of the genetic variation between samples with resolution down to the strain level: This challenge and validated genome identification was addressed as an approach for sample attribution.
The main attribution question: are the genetic contents of samples A and B relatively identical? With any two populations, there will be slight differences between minor constituents and potentially acquired sequence variations. Relatively identical signifies that the minor differences between the genomic content of the two samples are within an expected amount of variation for populations deriving from the same source (Figure 1).
Figure 1 Short sequence matching is used to calculate the probability of similarity (identity) based on genome clustering, based on the core/pan-genomes and the distance or relatedness of the unknown sample genetic content
There are three main components to variation between samples of same source:
- sequencing run variation
- recently acquired mutations/lateral gene transfer
- differences in relative proportions of microbial constituents
The art of genomic analysis in microbial forensics is thus the ability to distinguish between acceptable levels of differences caused by factors 1-3 above vs. distinction between two different source populations.
The purpose of this experiment was threefold:
- Observe DNA sequence mutations arising from an originally clonal isolate of Bacillus anthracis strain Ames when cultured under stressful conditions in the laboratory
- Determine the strengths and limitations of whole genome sequence analysis for characterizing variation between similar substrains
- Advance methods for determining “relatedness” between microbial samples.
Herein is the report on the genetic variation across 12 independent, but identical starting B. anthracis colonies after eight passages each. Our resulting data allow us to characterize levels and patterns of genetic variation within the context of a repeated passage experiment. The results demonstrate the feasibility of direct sequence analysis on samples for genome identification and addressed challenges centered on the bioinformatics software.
Materials and Methods
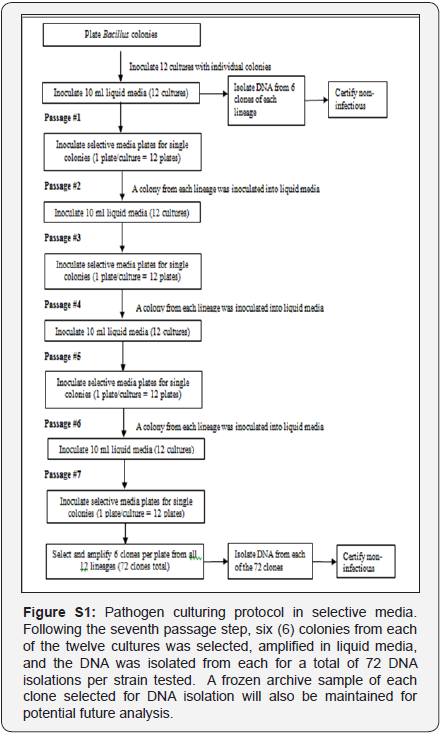

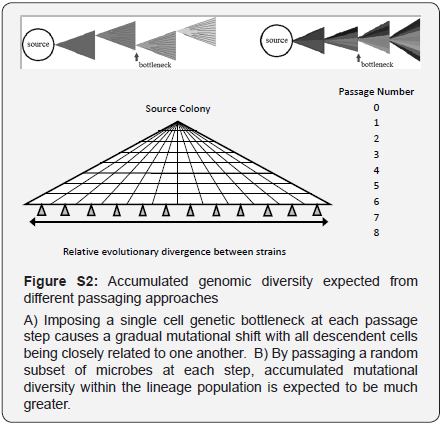
A single colony of B. anthracis strain Ames (BEI#NR-411) was passaged into 12 different plates. These twelve bacterial cultures were maintained separately over the course of seven more passages (Figure S1). Each culture passage was started with a single clonal colony streaked out on a petri dish. This created a single genome bottleneck at each passage step. Mutational variations differentiating each lineage were thus a result of initial variation in the source clonally derived culture plus mutations accumulated during the course of the eight growth and passage steps (Figure S2).Colonies were alternately cultured in tryptic soy broth, followed by culturing on selective Cereus identification agar plates for a total of seven more passages. At the end of eight passages, six clones from each of the twelve lineages were collected. DNA was isolated from each clone and sequenced using an Illumina Genome Analyzer IIx platform.
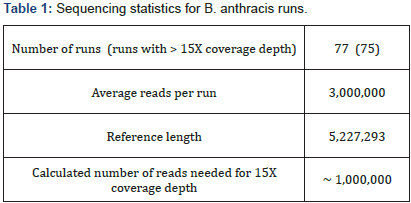
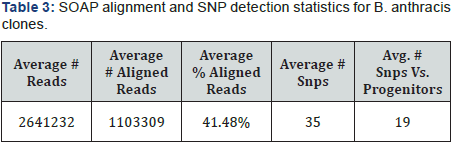
Single ended DNA sequencing was performed using a Genome Analyzer IIx (GA IIx) (Illumina, San Diego, CA). Library preparation was performed using a genomic DNA sample preparation Kit. DNA clusters were generated according to the manufacturer’s instructions using an Illumina cluster generation kit (Multiplexing Sequencing Primers and PhiX Control Kit v2) on an Illumina cluster station. All sequencing runs were performed with the GA IIx using the Illumina TruSeq SBS kit v5. Fluorescent images were analyzed with the Illumina CASAVA 1.8 software to obtain FASTQ-formatted sequence data of the short reads. For further details of the experimental methods [30]. Average characteristics of the sequencing runs is given in Table 1.
First, the performance of different assembly software were tested. Four sequence alignment software tools were selected, namely GNUMap [31], AMOScmp [32], SOAP2 [33] and BWA [34]. These alignment tools were selected based on popularity and third party performance comparisons. All of these methods can perform gapped and ungapped aligments.
Results and Discussion
Results of this study are contained in the descriptions of each step of our genome analysis pipeline. Develop phylogenetic distance mapping algorithms.
DNA signatures, barcodes, and other unique sequences can be used to detect the presence of an organism and to distinguish that organism from all other species. Real-time DNA sequencing files e.g. sff files are used for database matching, phylogenetic classification, reshuffled and randomly matched to calculate the degree of novelty. This approach leverages genetic information harbored across entire genomes and through matrix analysis enables comparison of specific targets of defined genomic value or weight to identify targets even ones with various degrees of relatedness.
First step – Selection of assembly software
To test different assembly algorithms, four software tools were selected [31-34]. Ade novo assembler (MIRA3) was also tested to explore whether reference genome influences the ability to assemble a genome in length and number of used reads. Although all reference based assemblers performed similarly, we selected two implementations, GNUMAP and SOAP, for the final analysis. AMOScmp, BWA and SOAP all implement the same algorithm (Burrows-Wheeler transform) whereas GNUMAP uses a probabilistic Needleman-Wunsch algorithm, which takes advantage of Illumina probability files to improve the mapping accuracy for lower quality reads and increase the amount of usable data produced in a given experiment. Therefore, the combined use of both SOAP and GNUmap allows an opportunity to take advantage of both approaches. MIRA3 de novo assembler yielded a shorter assembled genome and threw away a larger amount of reads, which depending on the dataset, ranged between 30% to 45%.
Second step - pipeline construction, assembly and SNP calling
A pipeline was constructed to manage, edit, and analyze the raw genomic WGS data. This pipeline was fully written in Python and uses elements of the BioPython package. The basic outline of this pipeline goes as follows:
- QSEQ -> FASTQ
- 1B. Optionally recalibrate quality scores for FASTQ data
- SOAPalign -> SOAPsnp -> FASTA genome and SNP file / GNUMAP -> SAM file -> BAM file -> FASTA genome and SNP file
- Pull “genes” (CDS, rRNA and tRNA) from the reference, BLAST against new genome
- Annotate new genome according to coordinates found by BLAST (including E-values)
- Annotate SNPs in new genome (including posterior probabilities)
- Cross reference SNPs with genes
- SNP annotations list the names of parent loci
- Gene annotations list SNPs within gene.
One advantage of this pipeline is that is fully adaptable to assemble new genome data. Some of the obstacles to overcome when developing the pipeline included normalizing data quality according to Sanger formatting prior to assembly and developing an approach for dealing with ORF frameshifts when annotating the genomes.
SNP calling was done under a Bayesian Inference framework, as implemented in SOAPsnp, by comparing an assembled genome against the reference genome used. The output files of the pipeline are: a nucleotide fasta file per chromosome, a GenBank annotated file per chromosome, and a SNP file with information on all chromosomes.
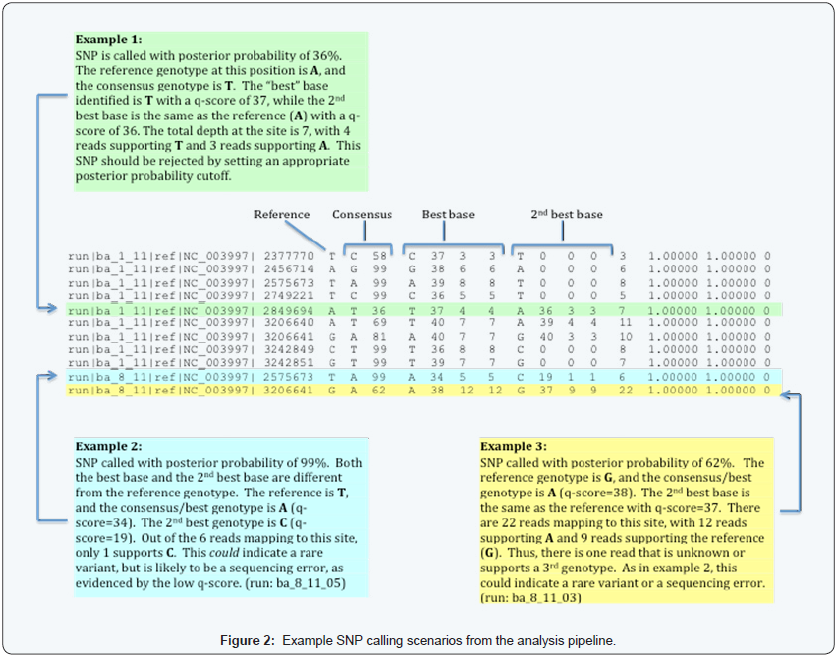
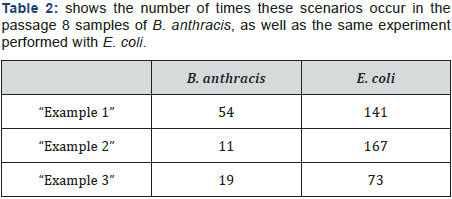
Figure 2 shows examples of the *.snp file and three SNP calling scenarios. The example shows a possible SNP where the 2nd best base is the same as the reference and has “good” support. Example 2 shows a possible SNP where the reference genotype, best base, and 2nd best base are all different from each other. Example 3 shows a possible SNP with reads that do not support either the best base or the 2nd best base. In example 1, “good” support means that 1) the average 2nd base quality is greater than 35; and 2) 40% or more of mapped bases (reads) are the reference/2nd genotype.
Population variation adds complexity to forensics analysis in addition to variation caused by foreign DNA present in biological assay reagents, machine errors from sequencing platforms, and errors introduced through shortcut assumptions used in assembly and alignment software. To further examine the variation, the differences were plotted for all of the clones across all of the lineages from the single point source. In Figure 3, we compared the depth of coverage in terms of the number of reads covering a particular SNP to the resulting posterior probability (PP) of the SNP read. The reference sequence for each passage 8 sample was its ancestral “progenitor” after passage 1. The resulting analysis shows that there is a general trend of increasing PP with increasing depth. Due to the single genome bottelnecking design of the experiment, the majority of SNPs in each lineage were identical for the same passage number.
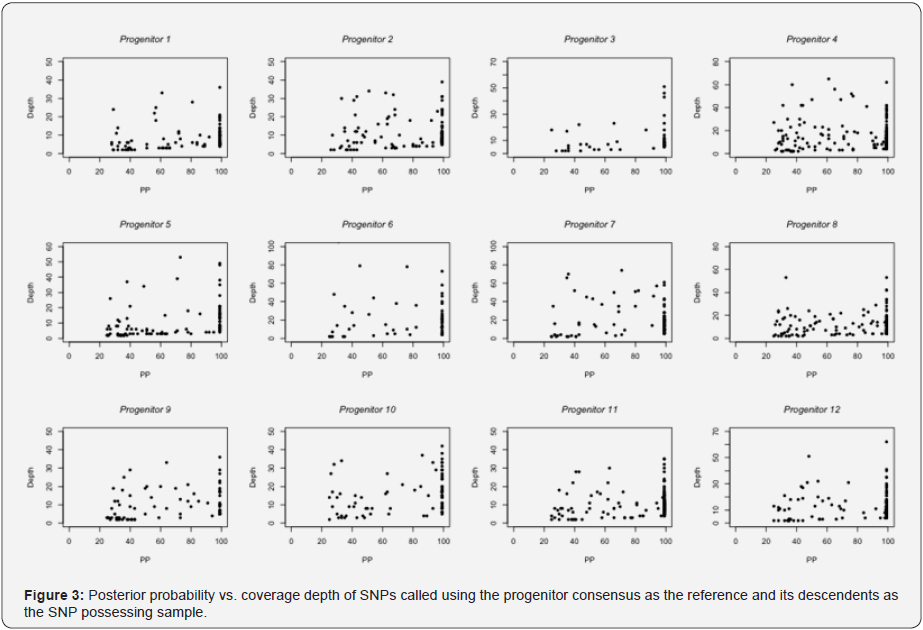
*SNP accuracy and depth of coverage3
SNPs were detected and validated by the SOAPsnp software tool. Statistics for average SNPs per clone detected are shown in Table 2. SOAPsnp does an acceptable job of detecting many SNPs, but fails to detect SNPs that are closely spaced together, due to an arbitrary rule in the SOAPsnp algorithm. SOAPsnp also does not require SNPs to be detected in both the forward and reverse directions – a validation requirement worth consideration (Table 3).
Third step – genome comparison and visualization
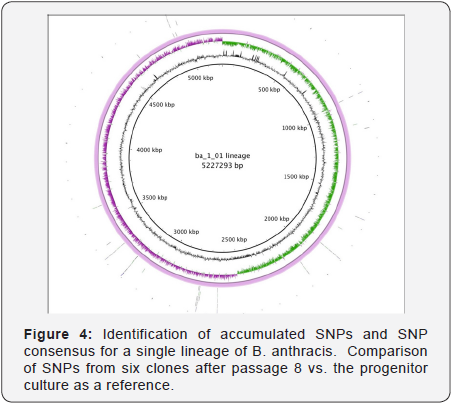
This graphical representation of the genomes show that the passaged genome is one component of the population of genomes from the isolate. This comparison (Figure 4) is one of 80 possible variants that represent major and minor components of the population. All the positions and types of variants were catalogued and used to build SNP phylogenies.
Assessment of inter-population genetic variation from a single point source. The total number of SNPs were shown relative to the confidence criteria described. This illustrates the different genotypes making up the population represented by the potions and with respect to which genome of that specific lineage (7 concentric circles of data points). This Figure 5 shows a composite view of the SNPs for one of the twelve lineages (01 lineage).
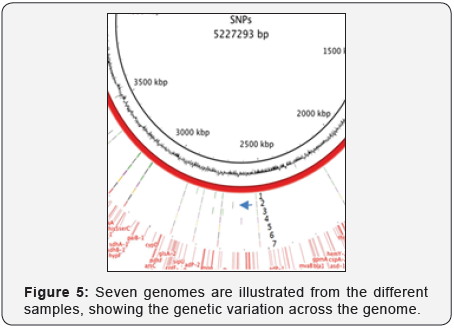
Figure 5 indicates the unique mutations serving as “DNA fingerprints.” In addition, we mapped the direction of mutation and compared taxonomic relationships and assembled genomes even when there were minor differences between related genomes. Using biothreat agents, e.g., B.anthracis, Y.pestis, B.mallei, Brucella, etc., cultured isolates and environment mixtures, mutations were tracked and assigned during passage, per lineage, from time-points along the collection schema across representative members of each population.Herein results for Bacillus anthracis are reported. Their individual genomes were built and phylogenetical analyses of their relationships revealed the unique DNA fingerprints associated with each lineage. These populations were measured and their diversity mapped with passage, which in turn enables traceability and attribution to a single source.
A substitution rate was calculated from this data using only the number of SNPs with posterior probabilities of occurrence above a trusted level of 95%. Nearly all SNPs scoring above 95% posterior probability scores also scored above 99%. The time period of seven culture passages was sufficient where SNPs were accumulated with complete consensus within each lineage. The culturing step after passage eight allowed for new SNPs to be acquired, yet no subsequent bottlenecking step was produced that would have forced 100% consensus for each SNP. Thus virtually all SNPs receiving posterior probability scores above 99% would derive from acquired mutations occurring after passage 1 and before passage eight-a period of seven passages.
While the number of generations from passage 1 until passage 8 are unknown, a relative substitution rate may still be calculated to be 1.4 SNPs per culture passage step along a single cell lineage. This rate is higher than what may be expected, its reputation as a slowly evolving species. Other factors involved in the slow environmental mutation rate of B. anthracis are the capability of a long dormant spore state and the stabilization of certain preferred SNP sequences and elimination of unfavorable mutations.
Fourth step - genome annotation and ortholog alignment
Genome annotation was approached using local alignments between a given target genome and an annotated reference genome. To this end, an in-house python script was developed in which the target genome was aligned against each CDS, rRNA and tRNA from the reference to then record the coordinates found by BLAST (including E-values). The output of this analysis is consolidated in a file formatted as a Gen Bank record that can be submitted to NCBI.
Since most population genomics and phylogenetic methodolgies depend on orthology assignment through multiple sequence alignment, we have explored two pieces of software able to perform whole genome alignments, namely Mauve and MAFFT.
Annotation of mutated genes
Validated SNPs from among the 84 clones were annotated along the reference Ames strain genome as shown in Figure 6. Overall, the validated SNPs were dispersed throughout the genome.
Fifth step - Phylogenetic and diversity estimates
Phylogenetic relationships were estimated via both Maximum Likelihood (RAxML) and a network approach (SplitsTree) to estimate relationships among strains. As expected from the experimental design (effectively a star-phylogeny evolutionary history), the strains showed little phylogenetic structure among lineages (Figure 7).
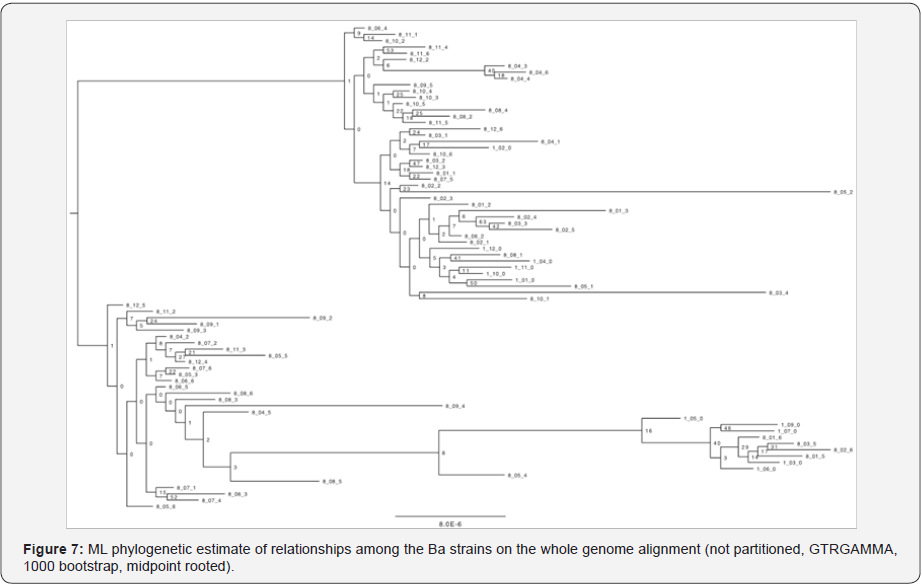
Reliability of different phylogeny approaches for determine relatedness within the same lineages and between different lineages were compared, as well as comparisons back to the reference isolate for each respective passage (t=1) lineage progenitor. From the experimental design, one would expect a split network to appear among the clones after 8 passages. Theoretically, the rays of the star phylogeny should converge in the middle where belongs the original isolate strain that the initial cultures were derived from. Logically, closer to the center of the star should be the progenitor strains, while the descendent colonies (8th passage) are expected to occur on the periphery of the star. An understanding of a pathogen’s genomic diversity will aid in attribution strategies (Figure 8).
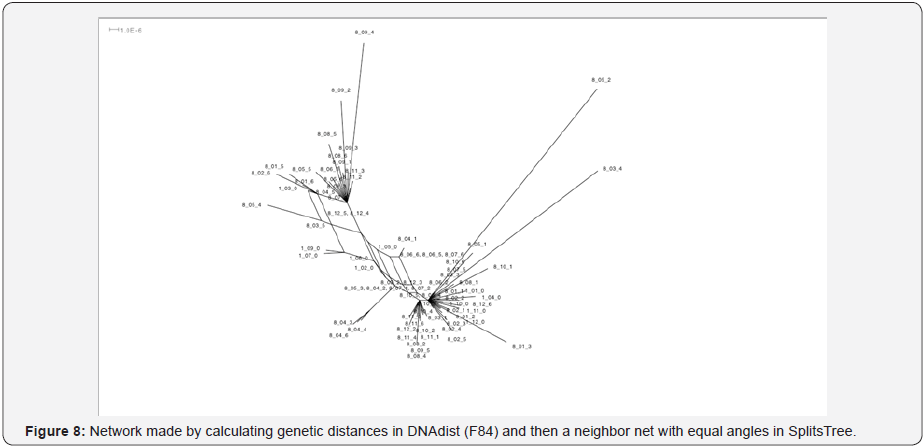

Discussion
A major goal of this study was to understand the limitations of whole genome analysis in a forensics context. The greatest surprise in our results was the recurrence of apparent convergent evolution at the level of new SNPs shared among different lineages. Many of the SNPs that arose after the stressed culturing conditions were shared among the lineages at passage 8, but not among the progenitors of each lineage. This strongly suggests that the biochemical changes imparted by these random mutations were potential imparted by the particular laboratory manipulation in the stressful media environment.
Phylogenic relationships derived from SNP data frequently deviated from the known relationships between clones within the same lineage and their progenitor of the lineage. It is known from other studies [27] that a number of SNPs can provide reliable determinant markers for distinguishing relationships between B. anthracis strains. SNPs that are evolutionarily favorable are, perhaps, less reliable markers of phylogenetic relationships than SNPs with no biological significance, since environmental stress will not dictate which biologically insignificant SNP variants are preserved or eliminated. This hypothesis could be extended to suggest that ideal genomic SNP markers for phylogeny would include patterns of multiple SNPs, with each SNP bearing little biological significance. Such low biological impact SNPs would include synonymous mutations, sequences of uncoded DNA, or conversion between similar amino acids in noncritical protein regions, such as conversion between leucine and isoleucine away from active sites of the protein.
Another consideration when developing phylogenies of closely related strains, these particular analysis SNP based phylogeny methods had difficulty distinguishing relationships at so close a level. At greater evolutionary distance or without the high level of evolutionary strain, the SNP differences between lineages ought to be more distinctive, thus allowing more accurate association of clones within lineages and discrimination between lineages. Longer reads with higher coverage or targeted amplicons should provide the resolution and strengthened data reliability.
In an uncontrolled microbial forensics investigation, the actual lineages of samples would not be known a priori, as they were in this study. On the other hand, the very close relationships between samples in this study probably made discrimination between these closely related genomes more difficult. The introduction of variously stressful and nonstressful environments further complicated the relationships by probably introduction of convergent evolution in validated SNP markers, although real life infections do involve varying host environments and correlated changes in evolutionary pressures.
WGS analysis of microbial samples is ofuse in forensics, however community acceptance and reliability of current analytical methods requires considerable refinement before the genomic analysis results are prepared to stand up in court on their own.
The global consequences of innovation and increased reliance on information technology by all nations, groups and individuals is changing both the speed at which threats at threats can develop and the tools our adversaries have at their disposal. The strategic outlook indicates the community must adjust to be able to field collection faster against an increasingly wide range of threat actors.
DNA sequencing and Population-sequencing [12] leverages a thorough understanding of evolutionary relationships within a species, combined with geographic distribution. We are better able to identify abnormal patterns that may be indicative of outbreaks/nefarious events and initiate abatement procedures. Phylogeographic and population genetic knowledge also forms the foundation for source attribution. While this work has been reactive, in the future, the methods used and concepts explored, will ultimately contribute to predictive modeling for disease prevention and abatement that will be applicable throughout the world.
Acknowledgement
This study was funded by Department of Homeland Security contract Whole Genome Approach to Microbial Forensics (WGAMF) HSHQDC-10-C-00140.Samples were prepared and handled at the CUBRC BSL facility and cultured under standard procedures. DNA from serial passages of biothreat agents was extracted and sequenced. Sequencing was conducted at ECBC as per Illuminarecommended protocols.
References
- Budowle B (2005) Crit Rev Microbiology 31: 233-254.
- 3rd Annual CBRN conference (2008) Tysons Corner, VA Jan.
- Science Needs for Microbial Forensics (2014) Washington DC, The National Academies Press. Washington DC, USA.
- Homeland security.
- Nations Medical Countermeasure stockpile (2016) Washington DC, The National Academies Press, Washington DC, USA.
- Cirino NM, Musser KA, Egan C (2004) Expert Review of Molecular Diagnostics 4: 841-857.
- Jakupciak JP, Colwell RR (2009) Biological agent detection technologies. Mol Ecol Resour 9 (Suppl s1): 51-57.
- Vallone PM, Jakupciak JP, Coble MD (2007) Forensic application of the Affymetrix human mitochondrial resequencing array. Forensic Sci Int Genet 1(2): 196-198.
- Jakupciak JP, Maggrah A, Maragh S, Maki J, Reguly B, et al. (2008) Facile whole mitochondrial genome resequencing from nipple aspirate fluid using MitoChip v2.0. BMC Cancer 8: 95.
- Maragh S, Jakupciak JP, Wagner PD, Rom WN, Sidransky D, et al. (2008) Multiple strand displacement amplification of mitochondrial DNA from clinical samples. BMC Med Genet 9: 7.
- Jakupciak JP, Wells JM, Karalus RJ, Pawlowski DR, Lin JS, et al. (2013) Population-Sequencing as a Biomarker of Burkholderia mallei and Burkholderia pseudomallei Evolution through Microbial Forensic Analysis. J Nucleic Acids 2013: 801505.
- Jakupciak JP (2013) Population-Sequencing as a Biomarker for Sample Characterization. J Biomark 2013: 861823.
- Jakupciak JP, Wells JM, Lin JS, Feldman AB (2013) Population Analysis of Bacterial Samples for Individual Identification in Forensics Application. J Data Mining Genomics Proteomics 4: 138.
- Maiden MC (2006) Multilocus sequence typing of bacteria. Annu Rev Microbiol 60: 561-588.
- Urwin R, Maiden MC (2003) Multi-locus sequence typing: a tool for global epidemiology. Trends Microbiol 11(10): 479-487.
- Sachse K (2004) Specificity and performance of PCR detection assays for microbial pathogens. Mol Biotechnol 26(1): 61-80.
- Pettersson E, Lundeberg J, Ahmadia A (2009) Generations of sequencing technologies. Genomics 93(2): 105-111.
- Clarke J (2009) Continuous base identification for single-molecule nanopore DNA sequencing. Nat Nanotechnol 4: 265-270.
- Treangen, TJ.,Salzberg SL (2011) Repetitive DNA and next generation sequencing: computational challenges and solutions. Nat Rev Genet 13(1): 36-46.
- Reddy RM, Mohammed MH, Mande SS. (2012) TWARIT: An extremely rapid and efficient approach for phylogenetic classification of metagenomic sequences. Gene 505(2): 259-265.
- Liu B, Gibbons T, Ghodsi M, Treangen T, Pop M (2011) Accurate and fast estimation of taxonomic profiles from metagenomic shotgun sequences. BMC Genomics 12(Suppl 2): S4.
- Sundberg K, Clement M, Snell Q, Ventura D, Whiting M, et al. (2012) Phylogenetic search through partial tree mixing. BMC Bioinformatics13 Suppl 13: S8.
- Pérez-Losada M, Jobes DV, Sinangil F, Crandall KA, Arenas M, et al. (2011) Phylodynamics of HIV-1 from a phase III AIDS vaccine trial in Bangkok, Thailand.PLoS One 6(3): e16902.
- Read TD, Peterson SN, Tourasse N, Baillie LW, Paulsen IT, et al. (2003) The genome sequence of Bacillus anthracis Ames and comparison to closely related bacteria. Nature 423: 81-86.
- Read TD, Salzberg SL, Pop M, Shumway M, Umayam L, et al. (2002) Comparative genome sequencing for discovery of novel polymorphisms in Bacillus anthracis. Science 296(5575): 2028-2033.
- Rasko DA, Worsham PL, Abshire TG, Stanley ST, Bannan JD, et al. (2011) Bacillus anthracis comparative genome analysis in support of the Amerithrax investigation. Proc Natl Acad Sci U S A 108(12): 5027- 5032.
- Van Ert MN, Easterday WR, Huynh LY, Okinaka RT, Hugh-Jones Me, et al. (2007) Global genetic population structure of Bacillus anthracis. PLoS One 2(5): e461.
- Stratilo CW, Lewis CT, Bryden L, Mulvey MR, Bader D (2006) Single- Nucleotide Repeat Analysis for Subtyping Bacillus anthracis Isolates. J Clin Microbiol 44(3): 777-782 .
- Keim P, Van Ert MN, Pearson T, Vogler AJ, Huynh LY, et al. (2004) Anthrax molecular epidemiology and forensics: using the appropriate marker for different evolutionary scales. Infection, Genetics and Evolution 4(3): 205-213.
- Eppinger M, Worsham PL, Nikolich MP, Yinong Sebastian , Sherry Mou, et al. (2010) Genome sequence of the deep-rooted Yersinia pestis strain Angola reveals new insights into the evolution and pangenome of the plague bacterium. Journal of Bacteriology 192(6): 1685-1699.
- Clement NL, Snell Q, Clement MJ, Hollenhorst PC, Purwar J, et al. (2010) The GNUMAP algorithm: Unbiased probabilistic mapping of oligonucleotides from next-generation sequencing. Bioinformatics 26(1): 38-45.
- Pop M, Phillippy A, Delcher AL, Salzberg SL (2004) Comparative Genome Assembly. Briefings Bioinformatics 5(3): 237-248.
- Li R, Yu C, Li Y, Lam TW, Yiu SM, et al. (2009) SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics 25(15): 1966- 1967.
- Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25(14): 1754-1760.






























