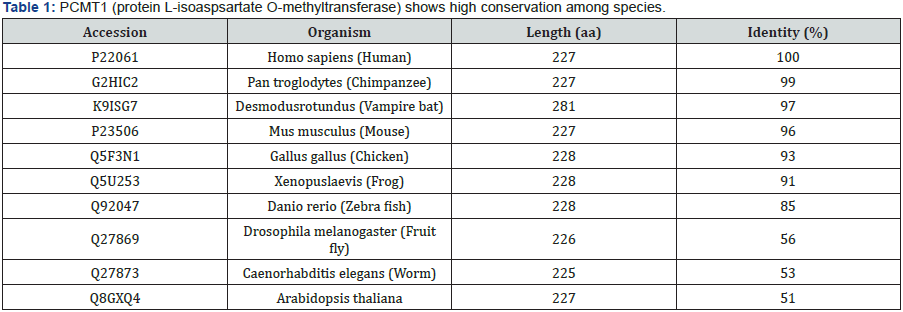
Protein L-Isoaspartate O-Methyltransferase (PCMT1): A Key Player of Spontaneously Arisen Protein Damage Repair
Burcu Biterge*
EpiGen Biotechnology Consultancy Inc, Turkey
Submission: July 15, 2017; Published: July 21, 2017
*Corresponding author: Burcu Biterge, EpiGen Biotechnology Consultancy Inc, Büklüm Sk Kavaklhdere 5/15, Çankaya, Turkey, Tel: (0312) 419 2215; Fax: (0312) 419 2216; Email: b.biterge@epigen.com.tr
How to cite this article: Burcu B. Protein L-Isoaspartate O-Methyltransferase (PCMT1): A Key Player of Spontaneously Arisen Protein Damage Repair. Anatomy Physiol Biochem Int J. 2017; 3(1): 555605. DOI:10.19080/APBIJ.2017.03.555605.
Abstract
Proteins undergo spontaneous non-enzymatic chemical modifications due to the exposure to e.g. oxidative reagents. The cell copes with the accumulation of such damaged proteins by proteosomal degradation. However, aged or damaged proteins can also be enzymatically repaired. Protein L-isoaspartate O-methyltransferase (PCMT1) catalyzes the methylation of isoAspartate (isoAsp) residues that spontaneously arise as a result of protein aging and facilitates their restoration to the normal state.PCMT1 plays a significant role in maintenance of protein homeostasis as well as in cellular function and integrity by acting on a wide variety of substrates.
Keywords: Spontaneous Protein Damage, Aging, Aspartate Methylation, Isoaspartate, PCMT1, Protein L-Isoaspartate O-Methyltransferase
Introduction
Methylation of aspartic acid residues was first described in the literature in erythrocytes as a possible step of repairing aged membrane proteins [1]. During the process of aging, L-aspartyl residues are spontaneously converted to L-isoaspartyl via the unstable intermediate L-succinimide which undergoes a spontaneous hydrolysis, generating a mixture of normal L-aspartate (15-30%) and abnormal L-isoaspartate (70-85%), pointed out as steps 2 and 3 in Figure 1 respectively [2]. Accumulation of this abnormal form of aspartate is recognized as damage in the cell and therefore needs to be repaired [3].
It has been previously shown that Protein L-isoaspartate O-methyltransferase (PCMT1, or alternatively called PIMT) can rapidly convert L-isoaspartyl sites to -carboxyl-O-methyl esters in vivo (step 4), which at physiological pH and temperature undergo spontaneous demethylation and give rise to the L-succinimide intermediate (step 5) [4]. Although multiple cycles of the repair mechanism is required to prevent L-isoaspartate sites from accumulating, it is the only repair pathway known so far (Figure 1). The methylation of isoAsp residues by PCMT1 is also suggested to act in targeting aged proteins for proteosomal degradation [5].

Protein L-isoaspartate O-methyltransferase (PCMT1) is encoded by a single gene located on human chromosome 6q22.3-24 [6]. Alternative splicing of the RNA transcripts gives rise to two isoforms; differing only in their last two or three aminoacids at the C-terminus. The more basic isoform carries a –RWK sequence whereas the more acidic isoforms bears a -RDEL sequence, which is recognized as ”endoplasmic reticulum retention signal” and is most likely to be cytoplasmic [7].Catalytic properties of these isozymes appear to be identical [8]. PCMT1 is a ubiquitously expressed enzyme. It was previously shown as a cytoplasmic protein in all mammalian tissues tested [9]; however, a recent study revealed that PCMT1 localizes both to the nucleus and to the cytoplasm [10]. It is well conserved among species; from human to mouse, chicken, fish, worms, insects and plants, with more than 95 percent similarity in amino acid sequence within mammals (Table 1).

PCMT1 has a wide range of substrates includingBcl-xL, p53, LaminA, Akt, MAP-2, - and -Tubulin, - and -Synuclein, Calmodulin, Calreticulin [3,11,12]. Hence, PCMT1 has been implicated in regulation of many cellular processes and signaling pathways (Figure 2). PCMT1 mediated methylation of p53 has been reported to directly down regulate p53 protein levels and thereby suppress the transcription of p53-target genes [11]. It has also been shown that the methylation of Bcl-XL deamidated at Asp52 and Asp66 is required for maintaining its anti-apoptotic functions and that the over expression of PCMT1 prevents apoptosis in endothelial cells after H2O2 induction [13]. In line with this, the miRNA 15a/16-1 cluster mediated silencing of PCMT1 was shown to interfere with the repair of the deamidatedBcl-XL, making cells more susceptible to cisplatin-induced apoptosis [14]. Another important function has been attributed to PCMT1 in PI3K/Akt pathway due to the activation of PI3K/Akt pathway and increased levels of insulin receptor in PCMT1 deficient mice. There are several reports that PCMT1 can also act on histone proteins to repair isoaspartate residues [2,5,15]. Furthermore, we recently showed that PCMT1 methylates histone H4 at aspartate 24 (H4D24me), implicating a possible role for PCMT in marking aged histones for replacement or degradation; which for the first time linked histone modifications with histone aging and histone protein homeostasis.
Conclusion
PCMT1 is a crucial component of the cellular protein repair machinery of the cell, which is evident by its wide variety of substrates that are important regulators of cellular function. In line with this, it has significant physiological and therapeutic implications. For instance, mice lacking PCMT1 exhibit accumulation of soaps in several tissues including brain, which may result in neuronal dysfunction, and they ultimately die of epileptic seizures, suggesting PCMT1 as a potential therapeutic target in certain brain diseases including epilepsy [16]. Therefore, future studies on PCMT1 will not only provide new insights into the cellular protein repair mechanisms, but also possible new targets for therapy.
References
- Janson CA, Clarke S (1980) Identification of Aspartic Acid as a Site of Methylation in Human Erythrocyte Membrane Proteins. J Biol Chem 255(24): 11640-11643.
- Carter WG, Aswad DW (2008) Formation, Localization and Repair of L-isoaspatyl Sites in Histones H2A and H2B in Nucleosomes from Rat Liver and Chicken Erythrocytes. Biochemistry 47(40): 10757-10764
- Vigneswara V, Lowenson JD, Powell CD, Thakur M (2006) Proteomic Identification of Novel Substrates of a Protein Isoaspartyl Methyltransferase Repair Enzyme. J Biol Chem 281(43): 32619-32629.
- Mc Fadden PN, Clarke S (1981) Methylation at D-aspartyl Residues in Erythrocytes: Possible Step in the Repair of Aged Membrane Proteins. Proc Natl Acad Sci U S A 79(8): 2460-2464.
- Young GW, Hoofring SA, Mamula MJ, Doyle HA (2005) protein L-isoaspartyl Methyltransferase Catalyzes in vivo Racemization of Aspartate- 25 in Mammalian Histone H2B. J Biol Chem 280: 26094- 26098.
- MacLaren DC, O’Connor CM, Xia YR, Mehrabian M, Klisak I, et al. (1992) The L-isoaspartyl/D-aspartyl Protein Methyltransferase Gene (PCMT1) Maps to Human Chromosome 6q22.3-24 and the Synthenic Region of Mouse Chromosome 10. Genomics 14(4): 852-856.
- MacLaren DC, Kagan RM, Clarke S (1992) Alternative Splicing of the Human Isoaspartyl Protein Carboxyl Methyltransferase RNA leads to the Generation of a C-terminal -RDEL sequence in Isozyme II. Biochem Biophys Res Commun 185(1) 277-283.
- Ota IM, Gilbert JM, Clarke S (1988) Two Major Isozymes of the Protein D-aspartyl/L-isoaspartyl Methyltransferase from Human Erythrocytes. Biochem Biophys Res Commun 151: 1136-1143.
- Clarke S (1985) Protein Carboxyl Methyltransferases: Two Distinct Classes of Enzymes. Annu Rev Biochem 54: 479-506.
- Biterge B, Richter F, Mittler G, Schneider R (2014) Methylation of histone H4 at aspartate 24 by protein L-isoaspartate O-methyltransferase (PCMT1) links histone modifications with protein homeostasis. Sci Rep 4: 6674.
- Lee MJ, Villa R, Trojer P, Norman J (2007) Demethylation of H3K27 regulates polycomb recruitment and H2A ubiquitination. Science 318(5849): 447-450.
- Farrar C, Houser CR, Clarke S (2005) Activation of the PI3K/Akt signal transduction pathway and increased levels of insulin receptor in protein repair-deficient mice. Aging Cell 4(1): 1-12.
- Cimmino A, Capasso R, Muller F, Sambri I (2008) PLoS One 3(9): e3258
- Sambri I, Capasso R, Pucci P, Perna AF (2011) The microRNA 15a/16- 1 cluster down-regulates protein repair isoaspartyl methyltransferase in hepatoma cells: implications for apoptosis regulation. J Biol Chem 286(51): 43690-700.
- Young AL, Carter WG, Doyle HA, Mamula MJ (2001) Structural Integrity of Histone H2B in vivo requires the Activity of Protein L-isoaspartate O methyltransferase, a Putative Protein Repair Enzyme. J Biol Chem 276: 37161-37165.
- Ogawara M, Takahashi M, Shimizu T, Nakajima M (2002) Adenoviral Expression of Protein L-isoaspartyl Methyltransferase (PIMT) Partially Attenuates the Biochemical Changes in PIMT-deficient Mice. J Neurosci Res 69(3): 353-361.






























