Late Onset Neonatal Sepsis in Sudan: Incidence, Bacteriological Profiles, Patterns of Antimicrobial resistance and Fatality
Abeer Abd Elmoniem Elhussein Salih1, Abd Elgadir Ali Bashir2, Mona Elfaki eltahir Elfaki3, Abu Bakr Mohamed Ali Nasr4 and Eltahir Awad Gasim Khalil5*
1 Sudan Medical Specialization Board, Sudan
2 Federal Ministry of Health, Sudan
3 Department of Microbiology and Immunology, College of Medicine King Khalid University, Kingdom of Saudi Arabia
4 Department of Obstetrics and Gynecology, Faculty of Medicine, University of Khartoum, Sudan
5 Institute of Endemic Diseases, University of Khartoum, Sudan
Submission: August 13, 2019; Published: September 10, 2019
*Corresponding author: Professor. Eltahir Awad Gasim Khalil, Department of Clinical Pathology & Immunology, Institute of Endemic Diseases, University of Khartoum, Khartoum, Sudan
How to cite this article: Abeer Abd Elmoniem Elhussein Salih, Abd Elgadir Ali Bashir, Mona Elfaki eltahir Elfaki, Abu Bakr Mohamed Ali Nasr, Eltahir Awad Gasim Khalil. Late Onset Neonatal Sepsis in Sudan: Incidence, Bacteriological Profiles, Patterns of Antimicrobial resistance and Fatality. Acad J Ped Neonatol. 2019; 8(1): 555784. DOI: 10.19080/AJPN.2019.08.555784
Abstract
Objective: Neonatal sepsis is a systemic bacterial infection of newborns with high mortality rates in developing countries. This study aimed to determine the incidence, bacteriological profiles, antibacterial susceptibility patterns and mortality of neonates with septicemia in ICUs of a referral maternal hospital in Sudan.
Materials and Methods: Following parents’ consent, neonates with clinical suspicion of septicemia were enrolled [n=317]. Blood cultures were collected. Identification of organisms and their antibacterial susceptibility patterns were done.
Results and Discussion: The incidence of septicemia was calculated at 14.6% (146/1000 neonates/year). Around half (149/317; 47.0%) of neonates had positive blood cultures, more than half (55.7 %; 83/149) grew one organism, while 44.3% (66/149) had poly-microbial infections. A fifth (29/149, 19.5 %) of culture-proven septicemic neonates perished the majority (62%, 18/29) due to K. pneumoniae. Hospitalization for more than 7 days (OR: 5.7, 95% CI: 3.3-9.7), low birth weight [1500-2499 grams] (OR: 4.5, 95% CI2.7-7.6) and prematurity (OR: 3.8, 95% CI: 2.05 -5.56) were incriminated in culture positive neonatal septicemia. Pseudomonas spp. were seen in about a third (34.2%, 76/222), Staphylococcus aureus in 22% (49/222) and Klebsiella spp. in 15.8% (35/222). Staphylococci were highly sensitive to Vancomycin and Meropenem, high levels of resistance to the penicillin G, cephalosporins and aminoglycosides were detected among gram negative bacteria.
Conclusion: Neonatal septicemia incidence and mortality rate were 146/1000 neonates/day and 195 deaths/ 1000 live births respectively. Pseudomonas spp, S. aureus and Klebsiella spp. were the commonest organisms. Hospitalization for >7 days, low birth and pre-maturity were important risk factors.
Keywords: Neonatal septicemia; Bacteriological profile; Antibacterial susceptibility; Sudan
Introduction
Neonatal sepsis refers to systemic and generalized bacterial infection of newborns, documented by a positive blood culture in the first 4 weeks of life with high mortality rates in developing countries. It is characterized by fever, hypothermia, malaise, tachycardia, hyperventilation, toxicity and/or prostration which results from circulating multiplying bacteria. Neonatal sepsis is an important cause of mortality and morbidity and is a life-threat ening emergency where prompt antibiotics treatment is essential for favorable outcomes. It accounts for 30-50% of total neonatal deaths each year. Neonatal sepsis is classified into early (EOS) and late onset (LOS) on the basis of presentation within 72 hours or after 72 hours to 30 days of life respectively. The timing of the transition from EOS to LOS is not clear-cut and depends on the causative pathogen. The causative organisms of neonatal sepsis vary from nursery to nursery and depend on prenatal factors, quality and quantity of the health care personnel, flora of the delivery rooms and availability of antibiotics. EOS is usually due to microorganisms that are acquired from the mother during the ante-partum or intra-partum periods, while pathogens causing LOS are generally acquired in the post-natal environment. Infections with group B Streptococci presenting within the first 7 days of life are usually regarded as early-onset and therefore of maternal origin. Whereas, infections with coagulase negative Staphylococci presenting at any age are likely to be hospital acquired. The usual mode of transmission of microbes in the nursery is by direct physical contact or indirectly from another infant through hands of facility personnel. Maternal factors like fever, pre-mature rupture of membranes, premature onset of labor, chorio-amnionitis, urinary tract infections and group B Streptococcus colonization play important roles in development of neonatal sepsis. Laboratory-confirmed bloodstream infection (LCBI) in neonates must meet the following criteria: patient ≤ 1 year of age and has at least 1 of the following: fever or hypothermia (rectal temperature >38 ℃ or <37 ℃) respectively), apnea or bradycardia and positive laboratory results that are not related to an infection at another site. The gold standard for diagnosis of neonatal septicemia is the isolation of bacterial agents from blood culture from one or two blood cultures with other laboratory evidence of sepsis, such as an elevated serum C-reactive protein level (CRP). Gram-negative bacteria [E. coli, P. aeruginosa, K. pneumonae, H. influenzae], Staphylcoccus spp and β hemoliticus Streptococcus remain the major cause of infection with increasing resistance to commonly used antibiotics in developing countries. As many as 60% of blood cultures would be falsely negative for common neonatal pathogens due to a number of causes. In cases of clinically suspect culture-negative neonates, adjunct laboratory tests like serum pro-calcitonin, C-reactive protein, IL-6/IL-8 levels can be helpful. The polymerase Chain Reaction (PCR) is emerging as an important technique for accurate and rapid diagnosis of neonatal septicemia [1-15].
Management of Neonatal Sepsis is challenging due to several physiological peculiarities, different neonatal antibiotic pharmacokinetics compared to older children. The knowledge of anti-microbial susceptibility patterns of common causative pathogens of neonatal sepsis helps to inform antibiotics choice provided periodic active surveillances are carried out. Antibiotics are used prophylactically to manage asymptomatic newborns when the mother has risk factors for infection or has a confirmed infection. WHO recommendation, of Ampicillin and Gentamicin combination for treatment of neonatal sepsis may no longer be effective in treating many neonates with sepsis. This is due to the fact that the majority [71%] of Klebsiella and 50% of E. coli are reportedly resistant to Gentamicin, leading to increase mortality due to neonatal sepsis in developing countries [16-21].
This study aimed to determine the bacteriological profiles, antibiotics susceptibility patterns of isolates, risk factors and mortality rates of late neonatal septicemia in intensive care units in a central maternal referral hospital in Sudan.
Materials and Methods
Ethical Considerations
The research proposal was reviewed and approved by the Scientific and Ethics Committees of the Federal Ministry of Health, Sudan. Parents were enlightened and asked to sign a consent form if they agree to enroll their neonates in the study.
Study Design and site
This was a prospective, cross-sectional and facility-based study that was conducted at the Intensive Care Unit of Omdurman Maternity Hospital Sudan, during the period of January to April 2014. Three hundred and seventeen neonates of 0-28 days of age were enrolled. Inclusion criteria included: neonates suspected of having septicemia on clinical grounds [fever/hypothermia, rectal temperature >38 ℃, <37 ℃ respectively); apnea; bradycardia; tachycardia; hyperventilation; toxicity and/or prostration]. Exclusion criteria were out-patient born babies, birth weight <1500 grams and presence of apparent congenital anomalies.
Sampling: Sample frame: Sample size:
The study sample size was calculated as follows:
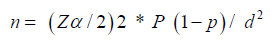
Z = Standard normal deviation of 1.96 corresponding to 95% confidence interval
P= frequency of occurrence of event = prevalence rate = 0.25
1-p = frequency of non-occurrence =0.75
d = is a degree of accuracy of the results (marginal error), set at 0.5
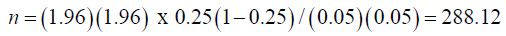
To adjust for non-responses, 10% of the calculated sample size will be added to n as follows:
288 x 0.1 = 28.8
The Total sample size = 288 + 28.8 = 316.8
Therefore, n was set as = 317 neonates
Sampling technique
Systematic random sampling was chosen as the sampling technique, where the first one was selected randomly and another one will be after k interval which was set as equal to 3.
Study variables
Dependent variables: septicemia.
Independent variables: Age, sex, birth weight, mood of delivery, Apgar score, duration of mechanical ventilation, intra-vascular catheterization, antibiotic usage, duration of hospitalization, profile of infecting bacteria and its antibiotic susceptibility patterns.
Microbiological material and data
Blood sampling and processing: Blood samples were drawn aseptically, 2mls of venous blood were collected and immediately inoculated directly into a tube containing Thioglycollate broth. Samples were incubated aerobically at 37 °C for18-24 hours and were examined daily for 7 days for visible signs of bacterial growth as manifested by turbidity in the media above the red cell level, gas production, heamolysis or/and coagulation of the broth.
Identification of organisms
Culture and gram staining: Blood cultures that showed signs of microbial growth were sub-cultured onto blood, chocolate and MacConkey agar. The blood, MacConkey and chocolate agar plates were incubated in aerobic in micro-aerophilic atmospheres using a candle jar at 37 °C for 24-48 hours. All positive blood cultures were identified by their characteristic appearance, gram-staining and confirmation by the pattern of biochemical reactions using standard methods. Cultures that showed no microbial growth within 7 days were reported as negative.
Biochemical tests
Members of the Enterobacteriaceae family were identified by indole/H2S production, citrate utilization, motility test, urease test, oxidase, carbohydrate utilization (Biomerieux, France). Gram-positive bacteria were identified using coagulase, DNase, catalase, bacitracin and optochin susceptibility tests [22].
Antimicrobial susceptibility testing
Small filter paper disks (6mm) impregnated with a standard amount of antibiotic (commercially available) were placed onto a Muller Hinton agar plates to which test bacteria had been swabbed. The plates were incubated overnight at 37 ℃ and the zone of inhibition of bacterial growth was used as a measure of susceptibility. Interpretation of results was done as suggested by Clinical Laboratory Standards Institute (CLSI) [23].
Data management and analysis
Data was tabulated, entered into Microsoft excel sheets and uploaded to SPSS (Statistical Package for Social Science) software. Frequencies with 95% confidence interval (CIs) and Odds Ratio were calculated.
Results
Maternal and Neonatal Characteristics
The majority of the study neonates (65.3%; 207/317) were preterm, have low birth weights (64.0%; 203/217)) and were predominantly males (males 197/317; 62.1%) with a male to female ratio of 5: 3. About half (48.9%; 155/317) were delivered by caesarian section/instrumentation. In the first minute after birth, the vast majority (83%; 263/317) had an Apgar score of ≤ 7, while at 5 minutes the per cent age dropped to 60.2% (191/317). The majority of the neonates (68.2%, 216/317) were hospitalized for more than 3 days about a third (36.3%; 115/317) received mechanical ventilation for more than 7days (Table 1).
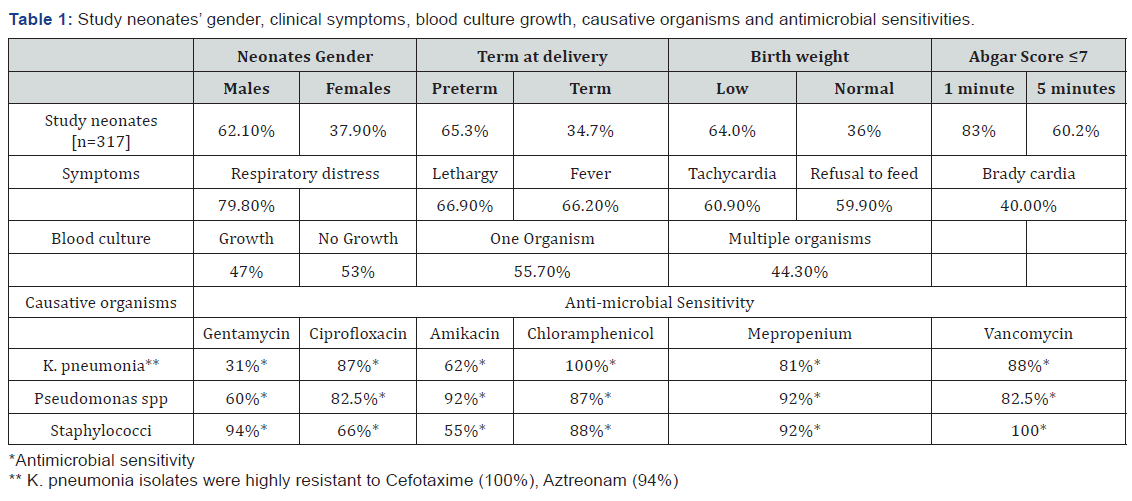
Respiratory distress was the commonest clinical feature of LOS and was seen in the majority (79.8%; 253/317) followed by lethargy (66.9%; 212/317), fever (66.2%; 210/317), tachycardia (60.6%; 192/317), refusal to feed (59.9 %, 190/317), bradycardia (40%; 127/317), central cyanosis (35.0%; 111/317) and hypothermia (21.1%; 67/317). Other symptoms like un-explained jaundice (14.2%; 45/317), seizures (10.1%; 32/317) and bulging anterior fontanelle (6.0%; 19/317) were seen with lower frequencies.
Blood culture results
Around half of the study neonates (149/317; 47.0%) had positive blood cultures, while 53% (168/317) did not grow any organism. More than half (55.7 %; 83/149) grew one organism, while 44.3% (66/149) had poly-microbial infections. The incidence rate of neonatal septicemia was calculated at 14.6% (32/1000 patients/ day).
Risk factors of Late Onset neonatal culture-proven septicemia
Hospitalization for more than 7 days has been strongly correlated to developing culture-proven neonatal sepsis (OR: 5.7, 95% CI: 3.3-9.7), followed by low birth weight of 1500-2499grams (OR: 4.5, 95% CI2.7-7.6) and preterm deliveries (OR: 3.8, 95% CI: 2.05 -5.56).
Mortality and causative bacterial pathogens
About a fifth (19.5%; 29/149) of culture-proven septicemic neonates perished. The main causative organisms were K. pneumonia (62%), Pseudomonas spp (18%), Coagulase-negative Staphylococci (9%), S. aureus (9%), while other bacteria (E. coli, S. pneumoniae, S. pyogenes) was seen in a minority (2%).
Antimicrobial susceptibility results
All coagulase-negative Staphylococci (n=15; 100%) were resistant to Penicilin G, three quarters (76%) were resistant to Trimethoprim/ sulphamethoxazole and 69% were resistant to Fusidic acid. All these isolates were sensitive to Meropenium and Vancomycin. While their sensitivities to Ciprofloxacin, Amikacin, Chloramphenicol and Gentamicin were 94%, 89%, 87% and 78% respectively. Staphylococci aureus isolates (n=32) showed complete resistance to Penicilin G (100%) followed by Fusidic acid (98%) and Trimethoprim/sulphamethoxazole (75%). The isolates were highly sensitive to Vancomycin (100%) followed by Gentamicin (94%), Meropenium (92%), Chloramphenicol (88%), Ciprofloxacin (66%) and Amikacin (55%). Most of Pseudomonas spp. isolates (n=40) were sensitive to Meropenum (92%) and Amikacin (92%). Sensitivities to Chloramphenicol, Ceftazidime, Vancomycin, Ciprofloxacin and Gentamicin were 87%, 85%, 82.5%, 82.5% and 60% respectively. Resistance to Trimethoprim/sulphamethoxazole, Cefotaxime, Aztreonam and Tetracycline was considerably high at 100%, 100%, 98%, 78% respectively. Klebsiella pneumonae isolates (n=16) were highly resistant to Cefotaxime (100%), Aztreonam (94%), Gentamicin (69%) and Tetracycline (69%), while they were highly sensitive to Chloramphenicol (100%), Vancomycin (88%), Ciprofloxacin (87%), Ceftazidime (82% and Meropenum (81%). Sensitivity to Amikacin is moderate (62%). Bacillus spp. isolates (n=12) showed high resistance to Cefotaxime (75%), while susceptibility is high for Gentamicin, Cefotaxime, Meropenum and Chloramphenicol at 100% followed by Ciprofloxacin at 83%. Escherichia coli isolates (n= 12) were highly resistant to Cefotaxime (100%), Amikacin (91%), Trimethoprim-sulphamethoxazole (83%), Ciprofloxacin (75%) and (Gentamicin 60%). Bacillus spp. were highly sensitive to Meropenum (92%) and Chloramphenicol (75%).
Discussion
Neonatal sepsis poses great mortality risk in developing countries, regular bacterial resistance surveillances are lacking making empirical antibiotics unsatisfactory. It is a fairly neglected disease that did not benefit much from recent advances in medical care. The incidence, clinical features, risk factors [low birth weight, prematurity] and mortality rates of neonatal septicemia in Sudan were found to be similar to those previously reported in other developing nations [1,3,4,7,17]. Clinical assessment using a combination of symptoms and signs was shown to be an inadequate guide to provisional diagnosis of neonatal sepsis, a finding in agreement with previous studies. This inadequacy is probably due to lack of consensus on the clinical definition of neonatal septicemia. In addition, these definitions are inappropriate for pre-terms who constitute considerable numbers of live births. Blood culture positivity rates in our study are comparable to reports from other developing nations but are higher than those reported from others. Causative organisms’ profiles reported in this study were similar to those reported from other parts of the developing world. K. pneumoniae, P. aeruginosa and E. coli were the most resistant pathogens to routinely used antibiotics, findings concordant with previous studies [24-30].
Conclusion
In conclusion, incidence and mortality rates of neonatal septicemia were high in neonatal ICUs in Sudan. Prematurity, low birth weights and prolonged hospital stay [>7 days] are important risk factors. Causative organisms [K. pneumoniae, Pseudomonas aeruginosa, S. aureus and E. coli] are considerably sensitive to Meropenium, Vancomycin, Genamicin and Amikacin, with marked resistance to Penicilin G. There is a pressing need to conduct regular anti-microbial sensitivity patterns to better guide empirical antibiotic use and update existing guidelines.
Acknowledgements
The authors would like to thank the staff of Omdurman Maternity Hospital, Sudan for their appreciable assistance and cooperation without them this work would not have been possible.
Conflict of Interest
The authors declare no financial or any other conflict of interest.
References
- Singh M, Gray CP (2019). Neonatal Sepsis. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; [Updated 2019 Jul 15]. Available from: https://www.ncbi.nlm.nih.gov/books/NBK531478/
- Agnihotri N, Kaistha N, Gupta V (2004). Antimicrobial susceptibility of isolates from neonatal
- Jpn J Infect Dis 57: 273-275.
- Weber MW, Carlin JB, Gatchalian S, Lehmann D, Muhe L, Mulholland EK (2003). WHO Young Infants Study Group. Predictors of neonatal sepsis in developing countries. Pediatr Infect Dis J 22(8):711-7.
- Stoll BJ, Hansen N, Fanaroff AA, Wright LL, Carlo WA, Ehrenkranz RA et al., (2002). Late-onset sepsis in very low birth weight neonates: the experience of the NICHD Neonatal Research Net-work. Pediatrics 110: 285-291.
- Simonsen KA, Anderson-Berry AL, Delair SF and Dele Davies H (2014). Early-Onset Neonatal Sepsis. Clin Microbiol Rev 27(1): 21–47. doi: 10.1128/CMR.00031-13
- Kaufman D and Fairchild KD (2004). Clinical microbiology of bacterial and fungal sepsis in very-low birth weight infants. Clin Microbial Rev 17(3):638-680.
- WHO (2004). Neonatal and perinatal mortality: country, regional, and global estimates,” Geneva, Switzerland, World Health Organization.
- https://apps.who.int/iris/bitstream/handle/10665/43444/9241563206_eng.pdf;jsessionid=5D0147AFD78148712AB2E541BAB03F00?sequence=1
- Qazi SA and Stoll BJ (2009). Neonatal sepsis: a major global public health challenge. Pediatr Infect Dis J 28 (Suppl):S1-S2. 37.
- Stoll BJ (1997). The Global Impact of Neonatal infection. Clin Perinatol. 24(1):1-21
- https://doi.org/10.1016/S0095-5108(18)30181-7 9550068863
- Warren S, Tristram S and Bradbury R (2010). Maternal and neonatal sepsis caused by Haemophilus influenzae type d. J Med Micr 59:370-372.
- Nwadioha SI, Nwokedi EOP, Kashibu E, Odimayo MS, Okwori EE (2010). A review of bacterial isolates in blood cultures of children with suspected septicaemia in a Nigerian tertiary hospital. Afri J Microbiol Res 4(4):222-225.
- Franz AR, Steinbach G, Kron M, Pohlandt F (2001). Interleukin-8: a valuable tool to restrict
- antibiotic therapy in newborn infants. Acta Paediatr 90:1025-32.
- Kordek A, Giedrys-Kalemba S, Pawlus B, Podraza W, Czajka R (2003). Umbilical cord blood
- serum procalcitonin concentration in the diagnosis of early neonatal infection. J Perinatol 23:148–153.
- Chan DK, Ho LY (1997). Use fullness of C-reactive protein in the diagnosis of neonatal sepsis.
- Singapore Med J 38:252–255.
- Jordan JA (2010). Molecular diagnosis of neonatal sepsis. Clin Perinatol. 2010; 37(2):411-9. doi: 10.1016/j.clp.2010.02.001.
- https://knoema.com/atlas/Sudan/Neonatal-mortality-rate.
- Bandyopadhyay T, Kumar A, Saili A, Randhawa VS (2018). Distribution, antimicrobial resistance and predictors of mortality in neonatal sepsis. J Neonatal Perinatal Med 11(2):145-153. doi: 10.3233/NPM-1765.
- Thaver D, Ali S and Zaidi A (2009). Antimicrobial Resistance Among Neonatal Pathogens in Developing Countries. Pediatr. Infect. Dis. J.; 28-1 Suppl.: S19-S21.
- Moniri R., Mosayebi Z., Movahedian M., Mousavi GA (2006). Increasing Trend of Antimicrobial Drug Resistance in Pseudomonas aeruginosa causing septicemia. Ira. J. Pub. Heal 35(1):58-62.
- WHO (2001). WHO Global strategy for containment of antimicrobial resistance, WHO,
- Switzerland
- Chaurasia S, Sivanandan S, Agarwal R, Ellis S, Sharland M, Sankar MG (2019). Neonatal sepsis in South Asia: huge burden and spiralling antimicrobial resistance. BMJ 364:k5314 doi: https://doi.org/10.1136/bmj.k5314.
- Hermaj V, Diksha S, Avneet G (2013). A Review on Commonly used Biochemical Tests for Bacteria. IJLS 1:1-7.
- https://www.asm.org/getattachment/2594ce26-bd44-47f6-8287-0657aa9185ad/Kirby-Bauer-Disk-Diffusion-Susceptibility-Test-Protocol-pdf.pdf
- James Wynn (2016). Defining neonatal sepsis. Curr Opin Pediatr 28(2):135–140, DOI: 10.1097/MOP.0000000000000315
- Murty DS, Gyaneshwari M (2007). Blood cultures in pediatric patients: a study of clinical impact. Indian J Med Microbiol 25:220-4.
- Aftab R, Iqbal I (2006). Bacteriological agents of neonatal sepsis in NICU at Nishtar
- Hospital Multan. J Coll Physicians Surg Pak 16:216-9.
- Shrestha, S; Adhikari, N; Rai, B. K.; Shreepaili, A (2010). Antibiotic Resistance Pattern of Bacterial Isolates in Neonatal Care Unit. J Nepal Med Assoc 50 (180): 277-281.
- Agrawal A, Awasthi S, Ghanghoriya P, Singh S (2018). Study of current status of bacteriological prevalence and profile in an inborn unit of SNCU in central India. Int J Contemp Pediatr 5(3):764-769 http://www.ijpediatrics.com
- Gyawali N, Sanjana RK (2012). Bacteriological Profile and Antibiogram of Neonatal
- Indian J. Pediatr. 5:1-5.
- Upadhyay A, Aggarwal R, Kapil A, Sigh S, Paul VK, Deorari AK (2006). Profile of Neonatal Sepsis in a Tertiary Care Neonatal Unit from India: A Retrospective Study. J Neonatol 20(1):50-57 DOI: 10.1177/0973217920060109






























