An Evaluation of the Performance of Two Quantification Methods for Trace DNA Casework Samples
Alketbi Salem K1,2*
1University of Central Lancashire, Preston, UK
2General Department of Forensic Science and Criminology, Dubai Police, Dubai, UAE
Submission:February 17, 2023;Published:February 22, 2023
*Corresponding author:Alketbi Salem K, University of Central Lancashire, Preston, UK
How to cite this article:Alketbi Salem K, An Evaluation of the Performance of Two Quantification Methods for Trace DNA Casework Samples. J Forensic Sci & Criminal Inves. 2023; 16(5): 555950 DOI:10.19080/JFSCI.2023.16.555950.
Abstract
Trace or Touch DNA is one of the most common types of DNA samples found at crime scenes and is often collected in vast quantities from crime scene exhibits. Quantification methods that use real-time PCR (qPCR) technology are highly sensitive compared to traditional quantification methods, therefore, this study evaluated the performance of Quantifiler™ Human and Quantifiler™ Trio quantification kits for the same set of trace DNA samples collected from handled items recovered from crime scenes at the Biology and DNA Section of the General Department of Forensic Science and Criminology of Dubai Police Force. Both methods achieved comparable results indicating that there was no significant difference between the two quantification methods to quantify the collected trace DNA from the handled items (p > 0.05).
Keywords: Forensic science; Trace DNA; Touch DNA; DNA quantification; Quantifiler™ Human DNA Quantification Kit; Quantifiler™ Trio DNA Quantification Kit; Global Filer™ PCR Amplification Kit
Introduction
Trace or Touch DNA is one of the most common types of DNA samples found at crime scenes and is often collected in vast quantities from crime scene exhibits. However, it is frequently found in minute quantities and the process of Touch DNA profiling can be impacted by many factors [1-14], which leads to low levels of DNA recovery, such as the efficiency of DNA extraction and quantification methods [4,15]. Quantification methods that use real-time PCR (qPCR) technology are highly sensitive compared to regular quantification methods such as Nano Drop™ or Gel electrophoresis, so they are the preferred technique for the quantification of trace DNA samples. This study evaluated the Quantifiler™ Trio and Quantifiler™ Human (Thermo Fisher Scientific) quantitative real-time PCR DNA kits widely used in forensic labs [1,2] on the same set of trace DNA samples collected from items recovered from crime scenes in the Biology and DNA Section of the General Department of Forensic Science and Criminology of Dubai Police Force.
Materials and Methods
Trace samples were collected with a Copan cotton swab (150C) moistened with 100 μL of molecular-grade water using a spray bottle [8] and extracted using the Prep Filer Express kit (Thermo Fisher Scientific) and the liquid handling and automation Tecan robot according to the manufacturer’s protocols. A full swab head was used with a final sample volume of 50 μL. The samples were then quantified manually using the Quantifiler™ Trio and Quantifiler™ Human, Quant Studio 5 Real-Time PCR (qPCR) and HID Real-Time PCR analysis software v1.3 (Thermo Fisher Scientific).
DNA amplification for some of the samples was performed using the Global Filer™ PCR amplification Kit on an ABI GeneAmp® 9700 PCR System (Life Technologies) for 29 cycles. The products of amplification were size-separated and detected on an ABI 3500 Genetic Analyser (Life Technologies) using 1 μl of PCR product, 9.6 μl Hi-Di™ formamide, and 0.4 μl GeneScan™ 600 LIZ® Size Standard v2.0 (Thermo Fisher Scientific). The data were statistically analysed using RStudio and factorial analysis of variance (ANOVA) and Microsoft Excel. All negative controls for DNA collection, extraction and amplification were confirmed DNA-free.
Results and Discussion
There was no significant difference between the results of the two quantification methods tested (Quantifiler™ Human DNA Quantification Kit and Quantifiler™ Trio DNA Quantification Kit) to quantify the collected trace DNA from the handled items (p > 0.05) (Figure 1). Similar quantities of DNA were recovered by both quantification methods. All samples were diluted in the same volume (10 μL from the sample with 5 μL of molecular grade water) for amplification with the Global Filer™ PCR Amplification Kit.
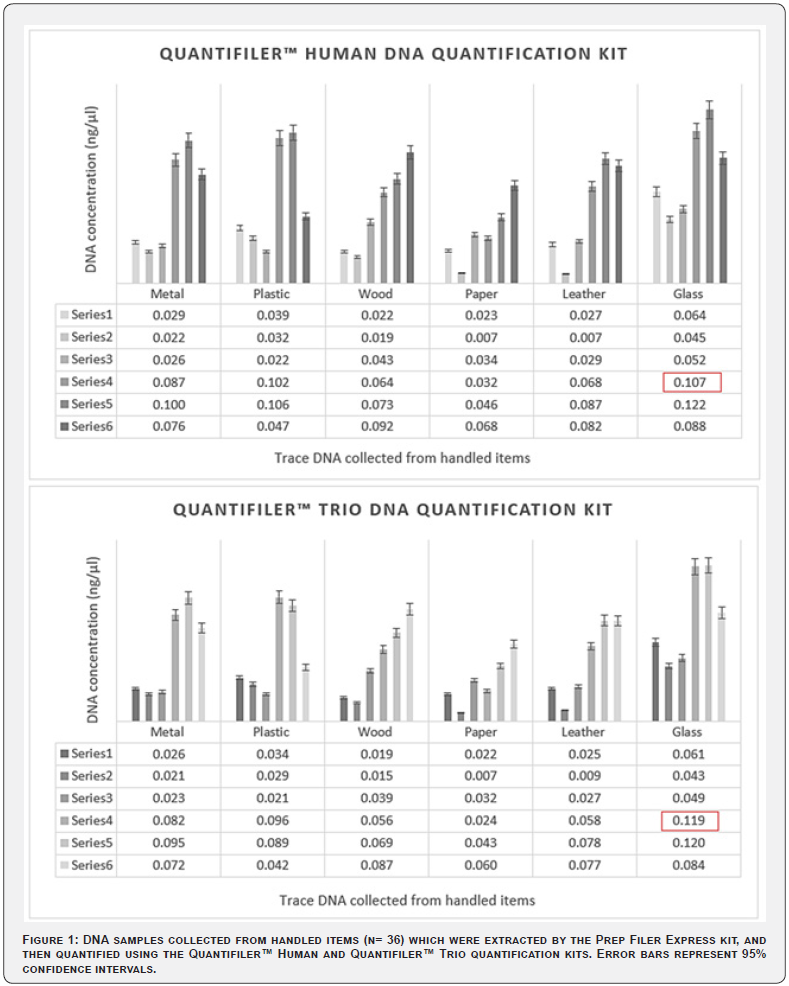

However, different volumes were required for one sample (circled in Figure 1), for which, 9.5 μL of the sample was diluted with 5.5 μL of molecular grade water for the Quantifiler™ Human kit and 8.6 μL of the sample was diluted with 6.4 μL of molecular grade water for the Quantifiler™ Trio kit. Full DNA profiles were obtained by both kits, but the peak heights (RFU) produced using the Quantifiler™ Human kit were relatively higher than the peaks obtained by the Quantifiler™ Trio kit (Figure 2). However, this small difference may be related to a human pipetting error or using an uncalibrated pipette, therefore establishing regular calibration protocols (e.g., every 3 months) or considering automated solutions to process samples for quantification can be beneficial.
Conclusion
The Quantifiler™ Human and Quantifiler™ Trio Quantification Kits are equally effective for the quantification of trace DNA but the Quantifiler™ Trio Quantification Kit provides more information about DNA degradation or human male DNA when using the large autosomal (LA) target and Y chromosome target in the same run.
Acknowledgements
This study was approved by the General Department of Forensic Science and Criminology in Dubai Police and ethical approval was granted by the University of Central Lancashire’s Research Ethics Committee (ref. no. STEMH 912).
References
- Salem Khalifa Alketbi (2018) The affecting factors of Touch DNA. Journal of Forensic Research 9(3): 424.
- Salem Khalifa Alketbi (2022) Analysis of Touch DNA [Unpublished doctoral dissertation]. University of Central Lancashire 15(5): 1-4.
- Salem Khalifa Alketbi (2020) Collection of Touch DNA from rotten banana skin. International Journal of Forensic Sciences 5(4): 000204.
- Salem Khalifa Alketbi, Goodwin W (2019) The effect of surface type, collection, and extraction methods on Touch DNA. Forensic Science International. Genetics Supplement Series 7(1): 704-706.
- Salem Khalifa Alketbi, Goodwin W (2019) The effect of time and environmental conditions on Touch DNA. Forensic Science International. Genetics Supplement Series 7(1): 701-703.
- Salem Khalifa Alketbi, Goodwin W (2019) The effect of sandy surfaces on Touch DNA. Journal of Forensic, Legal & Investigative Sciences 5: 034.
- Salem Khalifa Alketbi, Goodwin W (2021) Touch DNA collection techniques for non-porous surfaces using cotton and nylon swabs. Journal of Scientific & Technical Research 36(3): 28608-28612.
- Salem Khalifa Alketbi, Goodwin W (2019) Validating Touch DNA collection techniques using cotton swabs. Journal of Forensic Research 10(3): 445.
- Salem Khalifa Alketbi (2022) An innovative solution to collect Touch DNA for direct amplification. Journal of Forensic Sciences & Criminal Investigation 16(1): 555928.
- Salem Khalifa Alketbi, Alsoofi S (2022) Dual recovery of DNA and fingerprints using Minitapes. Journal of Forensic Sciences & Criminal Investigation 16(1): 555929.
- Salem Khalifa Alketbi, Goodwin W (2022) The impact of area size and fabric type on Touch DNA collected from fabric. Journal of Forensic Sciences & Criminal Investigation 16(1): 555926.
- Salem Khalifa Alketbi (2022) The impact of collection method on Touch DNA collected from fabric. Journal of Forensic Sciences & Criminal Investigation 15(5): 555922.
- Salem Khalifa Alketbi, Goodwin W (2022) The impact of deposition area and time on Touch DNA collected from fabric. Forensic Science International, Genetics Supplement Series 8: 45-47.
- Salem Khalifa Alketbi, Goodwin W (2023) Collection Methods for Touch DNA Direct Amplification. Journal of Forensic, Legal & Investigative Sciences 9: 072.
- Ottens Renée, Jennifer Templeton, Viviana Paradiso, Duncan Taylor, et al (2013) Application of direct PCR in forensic casework. Forensic Science International, Genetics Supplement Series 4(1): e47-e48.






























