Comparison of three Diagnostic Methods of Real time PCR, Reverse Dot Blot and DNA Sequencing for Analysis of KRAS Mutations in Patients with Colorectal Cancer
Fariba Jafary1, Sara Asadi2, Mansoor Salehi3,5, Farzaneh Jafary4* and Nayereh Nouri5*
1 Department of Biochemistry, Najafabad Branch, Islamic Azad University, Iran
2 Medical Genetics Laboratory, Alzahra University Hospital, Isfahan University of Medical Sciences, Iran
3 Cellular, Molecular and Genetics Research Center Isfahan University of Medical Sciences, Iran
4 Core Research Facilities (CRF), Isfahan University of Medical Science, Iran
5 Department of Genetics and Molecular Biology, Medical School, Isfahan University of Medical Sciences, Faculty of Science, Isfahan University, Iran
Submission: October 20, 2021; Published: November 08, 2021
*Corresponding Address: Dr. Nayereh Nouri, Department of Genetics and Molecular Biology, Medical School, Isfahan University of Medical Sciences, Isfahan, Iran, Faculty of Science, Isfahan University, Isfahan, Iran
How to cite this article: Nayereh N, Fariba J, Sara A, Mansoor S. Comparison of three Diagnostic Methods of Real time PCR, Reverse Dot Blot and DNA Sequencing for Analysis of KRAS Mutations in Patients with Colorectal Cancer. Canc Therapy & Oncol Int J. 2021; 20(2): 556031. DOI: 10.19080/CTOIJ.2021.20.556031
Abstract
Objective: KRAS has an important function to control cell growth, cell division, cell maturation, and cell death (apoptosis). KRAS mutant cases were reported in 30–40% of colorectal tumors. Status of KRAS mutation can be predictive of the response to drugs targeting the epidermal growth factor receptor (EGFR) in metastatic colorectal cancer patients. Consequently, due to importance of KRAS mutation status analysis for treatment of patients, a sensitive; economic and easily feasible method is required.
Methods: In this study specificity and sensitivity of three methods of direct sequencing, real-time PCR and reverse dot blot for analysis of KRAS mutant status were compared together in 23 colorectal patients.
Results: The obtained results indicated that there were significant differences between accuracy of three selected methods. The percentage error in two methods of Real-Time PCR and Reverse dot blot was 5% but in sequencing method was 34.79%.
Conclusion: So, Real-Time PCR and Reverse dot blot are suitable methods while sequencing method despite of its low cost and fast manner is not reliable method.
Keywords: Colorectal cancer; KRAS gene; Sequencing method; Real-Time PCR; Reverse dot blot
Introduction
>Colorectal cancer (CRC) is one of the most prevalent cancers among human population with high frequency of metastatic. Development of metastatic disease was occurred at approximately 50% of colorectal cancer patients [1,2]. Genetic and epigenetic variations are prevalent in CRC and can be considered as a main factor of tumorgenesis [3,4]. Chromosomal instability includes activation of proto-oncogenes such as KRAS and inactivation of at least three tumor suppression genes [5-7], microsatellite instability and CpG island methylator phenotype are three responsible pathways for genetic instability in colorectal cancer [4,5,8,9].
KRAS protein with participation of RAS-RAFMAPK pathway have a critical play in control of proliferation cell division, differentiation, apoptosis and survival of eukaryotic cells. Mutagenesis in RAS gene was reported in different types of human cancer [10-12]. The frequency of KRAS mutation in NSCLC, colorectal cancer and pancreatic carcinomas is 21-34%, 33% and 75-82.4% respectively [13-15]. KRAS is a powerful molecular marker in cancer diagnostics. Mutagenesis in KRAS gene can be divided in transversions and transitions. with exchangment of the amino acid glycine to another amino acid in the protein. Both mutation types were observed in colorectal cancer.
Anti-EGFR monoclonal antibodies like Cetuximab and Panitumumab are the major treatment for the patients with metastatic colorectal cancer (mCRC) without KRAS mutation [16,17]. Lack of response to these therapies was observed in patients with RAS activating mutations while low levels of primary resistance characterize in RAS wild type (WT) patients (only about 15%) [18]. So, identification of RAS mutation status before EGFR-targeted therapy is strongly recommended in therapy guidelines. Different methods were applied for determination of this mutation, but little comparative data is available for parameters such as analytical performance, economic merits, and workflow between these methods. therefore, introducing the most appropriate and precise method is necessary.
In the present study, we focused on comparing three diagnostic methods including Taq man Real Time PCR, PCR-RDB (Reverse Dot Blot) assay and PCR sequencing. KRAS mutational analysis was performed on codons 12 and 13 (exon 1) for 23 CRCs cases.
Material and Method
Sample selection
Genomic DNA was extracted from cancer tissue of 23 patients with pathologically confirmed CRC.
PCR Amplification
PCR-direct sequencing technique was applied for studying the KRAS-codon 12-13. The PCR amplification was carried out on the KRAS gene exon 2. The primer sequences were 5´-AAGGCCTGCTGAAAATGACTG-3´ and 5-CAAAGAATGGTCCTGCACCAG-3 (Kalikaki A, Koutsopoulos A, Trypaki M, et al). Finally, NCBI-Blast and ENSMBLE database were used to analyze the obtained results. Real- time PCR-Sequence-Specific Primers (SSP-PCR) was used to determine DNA mutant specific amplification from FFPE by Real Quality RI-KRAS MuST kit (ABANALITICA). The length of PCR products for both control and mutant gene were 150bp. For third method, Reverse Dot Blot, KRAS and BRAF somatic mutant genes were detected by PCR and hybridization reaction on DNA isolated from the tissue samples.
Two KRAS gene fragments of codon 12-13 and 61 (exon 1-2) and one BRAF codon 600 (exon 15) were parallel amplified. The Sequence-Specific-Oligonucleotide Probes (SSOPs) were used next. The following mutations can be detected via this method: KRAS G12A (GCT), G12C (TGT), G12D (GAT), G12R (CGT), G12S (AGT), G12V (GTT), G13C (TGC), G13D (GAC), Q61H (CAT), Q61H (CAC), BRAF V600E (GAA). It should be mentioned that the control zones were considered in this technique.
Result
The results are summarized in Tables 1 & 2. As presented in Table 1 between the three used methods, there is a conflict in 13 cases. The results of Real- time PCR and reverse dot blot were same and acceptable in comparison with the sequencing results that were not reliable. no mutation in codon 12 and 13 of KRAS gene was detected by sequencing in 7 cases while the mutation was identified in the cases by the other two methods. Moreover, in two other cases the peak of mutation was in sequencing graph was very weak, hardly to be detect; however, the two other techniques show this mutation clearly. In one case, the identified mutation in sequencing method was different from the other two methods [19-22] (Figure 1).

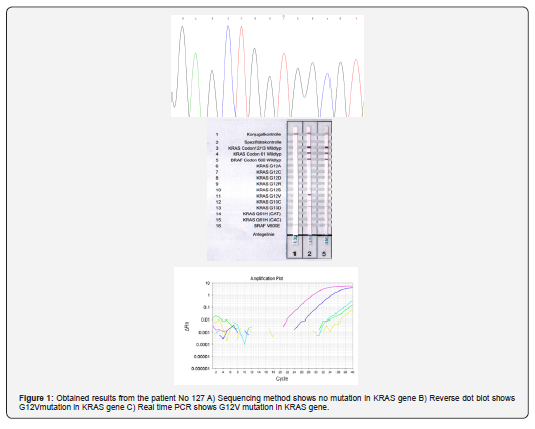

Discussion
CRC, one of the most common cancers in the word. Sporadic cases and genetic involvement were observed in 75% and 25% of patients respectively [23]. Therefore, the more effective and selective CRC therapy needs the evaluation of mutant KRAS status and finally all these aims require suitable diagnostic method [24-26]. The aim of this study was to compare the three mentioned methods (Real-Time PCR, Reverse dot blot and DNA sequencing) and introduce the most effective ones. A real-time polymerase chain reaction (Real-Time PCR) or quantitative polymerase chain reaction (qPCR), can be consider as a diagnostic test for infectious diseases, cancer and genetic abnormalities. This method is most specific, sensitive and reproducible [27-30].
Reverse dot-blot techniques reveal the most common and uncommon specific mutation. It is a non-radioactive technique with immobilize allele-specific oligonucleotide probes [31,32]. DNA sequencing is another diagnostic technique to determine DNA-nucleotide order precisely. It has wide applications range from forensic, medicine to agriculture. However, absence of complete coverage in this technique can case incorrect results. The other limit of DNA sequencing is its applications to test only for inversions or unbalanced translocations [33,34]. The obtained results revealed that DNA sequencing is not a reliable method to determine mutagenesis. In other words, if it is used alone, the irreparable damage can be resulted. Tumors with low content of KRAS mutated cells, not detectable by this method. Reverse dot-blot and Real-Time PCR have low error percentage in comparison with this method.
Despite of mutant KRAS in seven cases, the sequencing method failed to detect this mutation; furthermore, in two other cases it was hard to detect this mutation. In conclusion, regards to these mentioned issues the error percentage of the two other method was 5% and mutagenesis stages were detected correctly in all cases. Our result is in comparison with another study. By Jancik et al showed that direct sequencing failed to detect KRAS mutant in 21 patients [35]. Also, in Zinsky et al study sensitivity of direct sequencing and SnaPshot techniques was compared. The failure was reported more than 5 cases (11.1%) in DNA sequencing [36]. There is a detail discussion about determining the type of mutagenesis that was different among used techniques. It should be mentioned that incorrect probe connection may be the possible reason of this difference. Therefore, it is recommended to use two methods for mutation sensitive detection and consequently select the most effective therapy.
Conclusion
The performance of three methods of Real-Time PCR, DNA sequencing and Reverse dot-blot to detect of mutations in the KRAS gene was compared using DNA extracted from blood of 22 patients. The percentage error in direct sequencing method was about 35% whereas in two other methods were 5%. In conclusion, however sequencing method is a simple and rapid technique, it is not considered as a reliable method used alone for detection of somatic mutations especially in cancers [35,36].
Declarations
Funding
This study was funded by Isfahan University of Medical Sciences.
Conflict of Interest
The authors declare that they have no conflict of interest.
Author contributions
Fariba jafary: Doing an experiment, analysis of data
Sara Asadi: Collection of sampeles
Mansoor Salehi: Content approval
Nayereh Nouri: Doing an experiment, analysis of data
Informed consent
We are indebted to the patients who assisted us in carrying out this research. The ethical number of this research is IR.PNU.REC.1397.032.
References
- Gamelin FX, Baquet G, Berthoin S, Thevenet D, Nourry C, et al. (2009) Effect of high intensity intermittent training on heart rate variability in prepubescent children. Eur J Appl Physiol 105(5): 731-738.
- Rebecca L, Siegel Kimberly D, Miller Stacey A, Fedewa Dennis J, Ahnen Reinier GS, et al. (2016) Colorectal cancer statistics. CA: A Cancer J Clin 67(3): 177-193.
- Siegel RL, Miller KD, Jemal A (2016) Cancer statistics. CA Cancer J Clin 66: 7-30.
- Massimo P, Andrea R, Vittorio C (2012) Genetic and Epigenetic Events Generate Multiple Pathways in Colorectal Cancer Progression. Pathol Res Int 23: 348-509.
- Miew Keen C, Guy T (2012) Genetic and Epigenetic Biomarkers of Colorectal Cancer. Clin Gastroenterol Hepatol 10(1): 9–15.
- Winawer SJ, Zauber AG (2002) The advanced adenoma as the primary target of screening. Gastrointest Endosc Clin N Am 12(1): 1-9.
- Cong Tan, Xiang Du (2012) KRAS mutation testing in metastatic colorectal cancer. World J Gastroenterol 18(37): 5171–5180.
- Phipps AI, Buchanan DD, Makar KW, Win AK, Baron JA, et al. (2013) KRAS-mutation status in relation to colorectal cancer survival: the joint impact of correlated tumour markers. Br J Cancer 108(8): 1757–1764.
- Orsetti B, Selves J, Bascoul-Mollevi C, Lasorsa L, Gordien K, et al. (2014) Impact of chromosomal instability on colorectal cancer progression and outcome. BMC Cancer 14: 121-132.
- Lengauer KW, Kinzler BV, B Vogelstein (1997) Genetic instability in colorectal cancers. Nature 386(6625): 623–627.
- Hartman DJ, Davison JM, Foxwell TJ, Nikiforova MN, Chiosea SI (2013) Mutant allele-specific imbalance modulates prognostic impact of KRAS mutations in colorectal adenocarcinoma and is associated with worse overall survival. Int J Cancer 131(18): 1810–1817.
- Krasinskas AM, Moser AJ, Saka B, Adsay NV, Chiosea SI (2013) KRAS mutant allele-specific imbalance is associated with worse prognosis in pancreatic cancer and progression to undifferentiated carcinoma of the pancreas. Modern Pathol 26(10): 1346–1354.
- Payandeh M, Amirifard N, Sadeghi M, Shazad B, Farshchian N, et al. (2017) The prevalence of KRAS mutation in colorectal cancer patients in Iranian population: A systematic review and meta-analysis study. Biomed Res Ther 4(10): 1676-1692.
- Eberhard DA, Johnson BE, Amler LC, Goddard AD, Heldens SL, et al. (2005) Mutations in the epidermal growth factor receptor and in KRAS are predictive and prognostic indicators in patients with nonsmall-cell lung cancer treated with chemotherapy alone and in combination with erlotinib. J Cli Oncol 23(25): 5900–5909.
- Ogino S, Meyerhardt JA, Cantor M, Brahmandam M, Clark JW, et al. (2005) Molecular alterations in tumors and response to combination chemotherapy with gefitinib for advanced colorectal cancer. Clin Cancer Res 11(18): 6650–6656.
- Ugorcakova J, Hlavaty T, Babal P, Janega P, Bukovska G, et al. (2006) Assay design and optimization of mutant-enriched PCR based method for detection of K-ras gene mutations in pancreatic carcinoma. Neoplasma 53(5): 363–371.
- Kim HS, Heo JS, Lee J, Lee JY, Lee MY, et al. (2016) BMC Cancer. The impact of KRAS mutations on prognosis in surgically resected colorectal cancer patients with liver and lung metastases: a retrospective analysis. BMC Cancer 16: 120-128.
- Amirifard N, Sadeghi E, Farshchian N, Haghparast A, Choubsaz M (2016) Evaluation of KRAS Gene Mutations in Metastatic Colorectal Cancer Patients in Kermanshah Province. Asian Pac J Cancer Prev 17(7): 3085-3088.
- Martins M, Mansinho A, Cruz-Duarte R, Martins SL, Luis Costa L (2018) Anti-EGFR Therapy to Treat Metastatic Colorectal Cancer: Not for All. Adv Exp Med Biol 1110: 113-131.
- Karapetis CS, Khambata-Ford S, Jonker DJ, Callaghan CJ, Tu D, et al. (2008) K-ras mutations and benefit from cetuximab in advanced colorectal cancer. N Engl J Med 359(17): 1757–1765.
- Molinari M, Felicioni L, Buscarino M, Dosso S, Buttitta F, et al. (2011) Increased detection sensitivity for KRAS mutations enhances the prediction of anti-EGFR monoclonal antibody resistance in metastatic colorectal cancer. Clin Cancer Res 17(14): 4901–4914.
- Allegra CJ, Jessup JM, Somerfield MR, Hamilton SR, Hammond EH, et al. (2009) American Society of Clinical Oncology provisional clinical opinion: testing for KRAS gene mutations in patients with metastatic colorectal carcinoma to predict response to anti-epidermal growth factor receptor monoclonal antibody therapy. J Clin Oncol 27(12): 2091–2096.
- Santini D, Loupakis F, Vincenzi B, Floriani I, Stasi I, et al. (2008) High concordance of KRAS status between primary colorectal tumors and related metastatic sites: implications for clinical practice. Oncologist 13(12): 1270–1275.
- Bos JL, Fearon ER, Hamilton SR, Verlaan-de Vries M, van Boom JH, et al. (1987) Vogelstein B. Prevalence of ras gene mutations in human colorectal cancers. Nature 327(6120): 293–297.
- Perez-Ruiz E, Rueda A, Pereda T, Alcaide J, Bautista D, et al. (2012) Involvement of K-RAS mutations and amino acid substitutions in the survival of metastatic colorectal cancer patients. Tumour Biol 33: 1829–1835.
- Phipps AI, Buchanan DD, Makar KW, Win AK, Baron JA, et al. (2013) KRAS-mutation status in relation to colorectal cancer survival: the joint impact of correlated tumour markers. Br J Cancer 108(8): 1757–1764.
- Ciardiello F, Tejpar S, Normanno N, Mercadante D, Teague T, et al. (2011) Uptake of KRAS mutation testing in patients with metastatic colorectal cancer in Europe, Latin America and Asia. Target Oncol 6(3): 133–145.
- Normanno N, Pinto C, Castiglione F, Bardelli A, Gambacorta M, et al. (2011) KRAS mutations testing in colorectal carcinoma patients in Italy: from guidelines to external quality assessment. PLoS One 6(12): e29146.
- Fiore FD, Blanchard F, Charbonnier F, Pessot FL, Lamy A, et al. 007. Clinical relevance of KRAS mutation detection in metastatic colorectal cancer treated by Cetuximab plus chemotherapy. Br J Cancer 96(8): 1166–1169.
- Michael W, Vandesompele J, Kubista M (2009) Real-Time PCR: Current Technology and Applications. Caister Academic Press 978, p. 39.
- Huggett JF, Foy CA, Benes V, Emslie K, Garson JA, et al. (2013) The MIQE Guidelines: Minimum Information for Publication of Quantitative Real-Time PCR Experiments. Clin Chem 55(4): 611–622.
- Saiki RK, Walsh PS, Levenson CH, Erlich HA (1989) Genetic analysis of amplified DNA with immobilized sequence-specific oligonucleotide probes. Proc Natl Acad Sci USA 86: 6230–6234.
- Foglietta E, Bianco I, Maggio A, Giambona A (2003) Rapid detection of six common Mediterranean and three non-Mediterranean a-thalassemia point mutations by reverse dot blot. Am J Hematol 74(3): 191-195.
- Pettersson E, Lundeberg J, Ahmadian A (2009) Generations of sequencing technologies. Genomics 93: 105–111.
- Jay E, Bambara R, Padmanabhan R, Wu R (1974) DNA sequence analysis: a general, simple and rapid method for sequencing large oligodeoxyribonucleotide fragments by mapping. Nucleic Acids Research 1(3): 331–353.
- Jancik S, Drabek J, Berkovcova J, Xu YZ, Stankova M, et al. (2012) A comparison of Direct sequencing, Pyrosequencing, High resolution melting analysis,TheraScreen DxS, and the K-ras Strip Assay for detecting KRAS mutations in non-small cell lung carcinomas. J Exp Clin Cancer Res 31(1): 79.
- Zinsky R, Bolukbas S, Bartsch H, Schirren J, Fisseler-Eckhoff A (2010) Analysis of KRAS Mutations of Exon 2 Codons 12 and 13 by SNaPshot Analysis in Comparison to Common DNA Sequencing. Gastroent Res Pract 78: 93-63.






























