Genetics, Epigenetics and Cancer
Alain L. Fymat*
International Institute of Medicine and Science, USA
Submission: March 27, 2017; Published: April 07, 2017
*Corresponding author: Alain L. Fymat, International Institute of Medicine and Science, USA, Tel: (760) 485-9149; Email: alain.iymat@fiimas.org
How to cite this article: Alain L. F. Genetics, Epigenetics and Cancer. Canc Therapy & Oncol Int J. 2017; 4(2): 555634. DOI: 10.19080/CTOIJ.2017.04.555634
Abstract
With deeper understanding of cell biology and genetics, it now appears that cancer is less an organ disease and more a disease of molecular mechanisms caused by mutations of specific genes. Cancer is fundamentally a disease of tissue growth regulation failure when the genes that regulate cell growth and differentiation are altered. Most cancers have multiple possible concurring causes, and it is not possible to prevent all such causes. However, only a small minority of cancers (5-10%) are due to inherited genetic mutations whereas the vast majority (90-95%) are non-hereditary epigenetic mutations that are caused by various agents (environmental factors, physical factors, and hormones). Thus, although there are some genetic predispositions in a small fraction of cancers, the major fraction is due to a set of new genetic mutations (called "epigenetic” mutations). After a brief primer on cancer and its genetics, this article focuses on the epigenetics of cancer. Epigenetics is the study of cellular and physiological traits inherited by daughter cells, but not caused by changes in the DNA sequence. Important examples of epigenetic mechanisms (DNA methylation, histone modification, chromatin remodeling) and agents (prions, RNA and miRNA) are discussed. Epigenetic evidence in humans and epigenetic inheritance are reviewed to include environmental exposure, genomic imprinting and trans-generational inheritance. Epigenetic carcinogens and cancer treatment are also treated.
Abbreviations: ACS: American Cancer Society; BRCA: Breast Cancer; CAR: Chimeric Antigen Receptors; DNA: DeoxyriboNucleic Acid; DNAMT: DNA methyltransferase; GF: Growth Factors; GH: Growth Hormones; GIS: Growth Inhibitory Signals; HAT: Histone Acetyl Transferase; HDAC: Histone DeACetylase; HRT: Hormone Replacement Therapy; HLMT: Histone Lysine Methyl Transferase; HRT: Hormone Replacement Therapy; IGF: Insulin Growth Factor; mi-RNA: Micro RNA; MLL: Mixed Lineage Leukemia; OS: Osteosarcoma; PAMT: Protein Arginine Methyl Transferase; RNA: RiboNucleic Acid; RP: Repressor Proteins; SM: Somatic Mutations; SR: Silencer Regions; TPG: Tumor Promoter Genes; TS: Tumor Suppressor; TSG: Tumor Suppressor Genes
Introduction
Diseases such as cancer, heart disease and stroke now account for a major percentage of deaths worldwide and, thus, pose the greatest threat to modern global public health. Regarding cancer, according to the American Cancer Society (ACS), there were approximately 1.66 million new cancer cases in 2015. But why hasn't cancer been cured despite more than a four-decade "war” against the disease and the expenditure of hundreds of billions of dollars? In short, it is essentially our lack of understanding of the basic underlying molecular mechanisms. Such mechanisms have not been elucidated by chemotherapy, as evidenced by the fact that this approach does not work universally or permanently With the more recent deeper understanding of cell biology and genetics, it now appears that cancer is less an organ disease and more a disease of molecular mechanisms caused by mutations of specific genes.
The long quest for cancer cures has recently been reviewed [1] along with developments in anti-cancer therapy [2], including the new strategy of immunotherapy. This latter strategy and its synthetic version utilizing chimeric antigen receptors (CAR) aim to harness the body's immune system to fight cancer [3,4]. Similarly, recent developments in nanomedicine research [5,6] and more specifically the application of nanotechnology in cancer research and treatment [7,8] have also been reviewed.
In this article, building upon recent advances in epigenetics, I will concentrate on the epigenetics of cancer. Brief primers on cancer (including its causes, prevention, development, malignant progression, genetics, the transformative processes from a normal cell into cancer, and how cancer cells become resistant to treatment) and on epigenetics (including mechanisms, changes, classification, agents, evidence in humans and inheritance) will serve as a helpful background.
A Brief Primer on Cancer
Causes
A small minority of cancers, some 5-10%, are due to inherited genetics being caused by an inherited genetic defect (inherited mutations in the genes such as BRCA1 and BRCA2). The vast majority of cancers, some 90-95% of cases, are non- hereditary and due to various agents (environmental factors, physical factors, and hormones).
Environmental factors include lifestyle (diet and obesity; excessive tobacco smoking; alcohol over-consumption; stress; lack of physical activity), physical factors (pollutants; viral, bacterial and parasitic infections; ionizing and non-ionizing radiations), and economic and behavioral factors. It is nearly impossible to prove what truly caused a cancer in any individual, because most cancers have multiple possible concurring causes. Also, excepting the rare transmissions that occur with pregnancies and only a marginal few organ donors, cancer is generally not a transmissible disease. Some substances cause cancer primarily through their physical, rather than chemical, effects on cells (asbestos; synthetic asbestos-like fibers; non-fibrous particulate materials). It is controversial whether chronic inflammation can directly cause mutations. It is recognized, however, that inflammation can contribute to the proliferation, survival, angiogenesis and migration of cancer cells by influencing the microenvironment around tumors. Furthermore, oncogenes are known to build up an inflammatory pro-tumorigenic microenvironment.
Some hormones play a role in the development of cancer by promoting cell proliferation. Likewise, insulin-like growth factors (IGF) and their binding proteins play a key role in cancer cell proliferation, differentiation and apoptosis, suggesting possible involvement in carcinogenesis. Hormones are also important agents in cancers of the sex organs (breasts, endometrium, ovaries, prostate, and testes), and also of thyroid and bone cancer. Other factors are also relevant: obese people have higher levels of some hormones associated with cancer and a higher rate of those cancers. Women who take hormone replacement therapy (HRT) have a higher risk of developing cancers associated with those hormones. On the other hand, people who exercise far more than average have lower levels of these hormones, and lower risk of cancer. Growth hormones (GH) may promote osteosarcoma (OS) and, by artificially reducing hormone levels, hormone-sensitive cancers may be discouraged.
Development
Individuals differ in their inherited tendency to develop cancer. The process of cancer development in the body is a combination of events. Mutations occasionally occur within cells in the body as they divide. Unless they occur in germ cells, these mutations will not be inherited by any offspring, although they can affect the behavior of cells, sometimes causing them to grow and divide more frequently. There are biological mechanisms that attempt to stop this process: signals are given to inappropriately dividing cells that should trigger cell death (apoptosis), but sometimes additional mutations occur that cause cells to ignore these messages. An internal process of natural selection occurs within the body and eventually mutations accumulate within cells to promote their own growth, creating a cancerous tumor that grows and invades various tissues of the body.
Normally, a cell divides in response to signals called Growth Factors (GF) and stops growing once in contact with surrounding cells in response to Growth Inhibitory Signals (GIS). It usually then divides a limited number of times and dies, staying within the epithelium where it is unable to migrate to other organs. To become cancerous, a cell has to accumulate mutations in a number of genes that allow it to bypass this regulation and then no longer needs GFs to divide. It continues growing when making contact with neighboring cells, ignoring inhibitory signals, and growing indefinitely to become immortal. It will then escape from the epithelium and ultimately from the primary tumor, cross the endothelium of a blood vessel, be transported by the blood stream to colonize a new organ, forming metastases. Although there are some genetic predispositions in a small fraction of cancers, the major fraction is due to a set of new genetic mutations. These new, somatic mutations (SM) originally appear and accumulate in one or a small number of cells that will divide to form the tumor but are not transmitted by the progeny. The most frequent mutations are a loss of function of the p53 protein, a tumor suppressor (TS), or in the p5 pathway, and gain of function mutations in the protein, or in other oncogenes.
Malignant Progression
Douglas Hanahan and Robert Weinberg [9] proposed the following steps, which they dubbed as the "Hallmarks of Cancer”:
1. "Evasion of apoptosis (or programmed cell death);
2. "Self-sufficiency in growth signaling;
3. "Insensitivity to anti-growth signals;
4. "Induction and sustainment of angiogenesis;
5. "Enabling of a limitless replicative potential;
6. "Activation of metastasis and invasion of tissue;
7. "Reprogramming of energy metabolism; and
8. "Evasion of immune destruction.”
The above multi-step progression from normal cells to cells that can form a discernible mass to outright cancer is known as "malignant progression”. When cancer begins, it invariably produces no symptoms. Signs and symptoms only appear as the mass continues to grow. Symptoms can be local or systemic, the resultant finding depending on the type and location of the cancer.
Metastatis is the spread of cancer from its original site to distant sites by one or more pathways: local spread, lymphatic spread to regional lymph nodes, and by blood. In the latter instance, cancer spreads all over the body. In the soil and seed hypothesis of cancer metastasis, cancer 'seeds' grow in certain selected site(s) only (‘soil’). The symptoms of metastatic cancers depend on the location of the tumor, and can include enlarged lymph nodes, enlarged liver, enlarged spleen, and neurological symptoms.
Prevention
While hereditary genetic disorders may cause cancer, as stated earlier, the vast majority of cancer cases are due to environmental risk factors. Many, but not all, of these environmental factors are controllable lifestyle choices. Thus, cancer is considered a largely preventable disease. Greater than 30% of cancer deaths could have been prevented by avoiding certain risk factors including; tobacco use (smoking and chewing), alcohol consumption, overweight, obesity, poor diet, physical inactivity, transmitted infections, and air pollution. However, not all environmental causes are controllable such as, for example, naturally occurring background electromagnetic radiation. Thus, it is not possible to prevent all causes of cancer.
Genetics of Cancer
Cancer is fundamentally a disease of tissue growth regulation failure. In order for a normal cell to transform into a cancer cell, the genes that regulate cell growth and differentiation must be altered. The affected genes are divided into two broad categories: tumor promoter genes (TPG) or oncogenes that promote cell growth and reproduction, and tumor suppressor genes (TSG) that inhibit cell division and survival. Malignant transformation can occur through the formation of new oncogenes, the inappropriate over-expression of normal oncogenes, or else by the under-expression or disabling of TSG. Typically, changes in many genes are required to transform a normal cell into a cancer cell.
Genetic changes can occur at different levels and by different mechanisms. The gain or loss of an entire chromosome can occur through errors in mitosis. More common are mutations, which are changes in the nucleotide sequence of genomic DNA. Large-scale mutations involve the deletion or gain of a portion of a chromosome. Genomic amplification occurs when a cell gains many copies (often 20 or more) of a small chromosomal locus, usually containing one or more oncogenes and adjacent genetic material. Genomic translocation occurs when two separate chromosomal regions become abnormally fused, often at a characteristic location. Small-scale mutations include point mutations, deletions, and insertions, which may occur in the promoter region of a gene and affect its expression, or may occur in the gene's coding and alter the function or stability of its protein product. Disruption of a single gene may also result from integration of genomic material from a DNA virus or retrovirus, leading to the expression of viral oncogenes in the affected cell and its descendants. Replication of the enormous amount of data contained within the DNA of living cells will probabilistically result in some errors (mutations). Complex error correction and prevention are built into the process to safeguard the cell against cancer. However, if a significant error occurs, the damaged cell can "self-destruct” through programmed cell death (apoptosis). If the error control processes fail, then the mutations will survive and be passed along to daughter cells.
Transformative Processes of a Normal Cell into Cancer
Some environments make errors more likely to arise and propagate. Such environments can include the presence of disruptive substances called carcinogens, repeated physical injury, heat, ionizing radiation, or hypoxia. The errors that cause cancer, are self-amplifying and compounding, for example:
a. A mutation in the error-correcting machinery of a cell might cause that cell and its daughters to accumulate errors more rapidly;
b. A further mutation in an oncogene might cause the cell to reproduce more rapidly and more frequently than its normal counterparts;
c. Another mutation may cause loss of a TSG, disrupting the apoptosis signaling pathway and result in the cell becoming immortal;
d. Yet another mutation in the signaling machinery of the cell might send error-causing signals to nearby cells; and
e. The transformation of normal cell into cancer is akin to a chain reaction caused by initial errors, which compound into more severe errors, each progressively allowing the cell to escape the controls that limit normal tissue growth. This rebellion-like scenario becomes an undesirable survival of the fittest, where the driving forces of evolution work against the body’s design and enforcement of order. Once cancer has begun to develop, this ongoing process (termed "clonal evolution”) drives progression towards more invasive stages. Clonal evolution leads to intra-tumor heterogeneity that complicates designing effective treatment strategies.
How Cancer Cells Become Resistant to Treatment
While many types of chemotherapy have been developed against cancer, oncologists do not know before starting treatment whether a patient might benefit from a particular drug. So being able to identify through a laboratory test whether a patient's tumor is either resistant or sensitive to a specific drug is crucial to enabling the rapidly developing field of "personalized medicine”
In a breakthrough discovery, Prof. Karni and his team of researchers at the Hebrew University Medical Center, Jerusalem, Israel, have identified a process by which cancer cells become resistant to certain drugs [10]. This process will lead to a reliable prediction as to which patients will be helped by chemotherapy and recover, and which patients will not be helped by these drugs. This finding could enable the reversal of the process and inhibit metastasis of malignant tumor cells. The researchers found that breast, lung and colon cancer cells change the structure of an enzyme called mnk-2, which is involved in the transmission of information from the environment/body into the cell. They also showed that the enzyme has two forms, a "normal" one that inhibits cancer and another form that promotes cancer development. They further showed that cancer cells change the structure of the mnk-2, so they eliminate the form that inhibits cancer and elevate the form that induces it, thus allowing the cancer cells to survive and grow faster. Still further, they found that the anti-cancer form of the enzyme activates a program of apoptosis (suicide) in normal cells under stress conditions.
To fight the resistance process, the Karni team developed molecules that can convert the cancerous form of the mnk-2 enzyme back into its normal form so they become sensitive to stress and to absorbing chemotherapeutic drugs. More importantly, the molecules that change the cancerous form of mnk-2 into the normal form will make it possible to overcome the drug resistance of cancer cells, making them instead sensitive and responsive to various anti-cancer treatments. Further laboratory work on this aspect is continuing.
In summary, the mechanism discovered by Prof. Karni et al explains how cancer cells eliminate the anti-cancer form of mnk-2 without changing their DNA and how they become resistant to anti-cancer treatments, a problem that exists for almost every cancer treatment today. The new molecules they developed to change the structure of the mnk-2 enzyme back to its normal form will enable re-sensitizing cancer cells into anticancer therapies. This research could lead to the development of a new biomarker for testing the sensitivity of a patient to specific drugs. The possibility of examining whether a patient will benefit from a specific drug treatment before the treatment starts is of primary medical interest. The Israeli scientists are now developing a diagnostic test for the marker they found.
Brief Primer On Epigenetics
Defining Epigenetics
The generally accepted definition of "epigenetics" (epi-from the Greek word sm meaning over, outside of, on top off, around + genetics) is the "study of cellular and physiological traits inherited by daughter cells, but not caused by changes in the DNA sequence". It is the study of stable, long-term alterations in the heritable transcriptional potential of a cell. Thus, unlike genetics, which is based on changes to the DNA sequence (the genotype), in epigenetics, the changes in gene expression (or cellular phenotype) have other causes [11].
The term epigenetics also refers to the changes themselves, the relevant changes to the genome that do not involve a change in the nucleotide sequence. Gene expression can be controlled through the action of repressor proteins (RP) that attach to silencer regions (SR) of the DNA. These epigenetic changes may last through cell divisions for the duration of the cell's life, and may also last for multiple generations even though they do not involve changes in the underlying DNA sequence of the organism; instead, non-genetic factors cause the organism's genes to behave (or "express themselves") differently.
Epigenetic Mechanisms
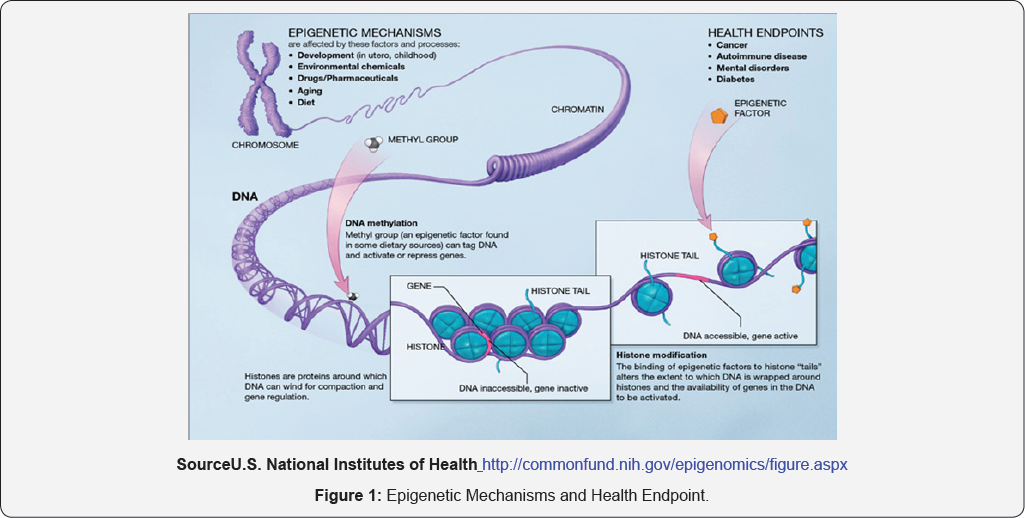
Several types of epigenetic inheritance systems may play a role in what has become known as "cell memory". (Note, however, that not all of these are universally accepted as examples of epigenetics.). Important examples of epigenetic mechanisms are DNA methylation, histone modification and chromatin remodeling, each of which alters how genes are expressed without altering the underlying DNA sequence. Epigenetic mechanisms are affected by several factors and processes including development in utero and in childhood, environmental chemicals, drugs and pharmaceuticals, aging, and diet. DNA methylation is what occurs when methyl groups, an epigenetic factor found in some dietary sources, can tag DNA and activate or repress genes. Histones are proteins around which DNA can wind for compaction and gene regulation. Histone modification occurs when the binding of epigenetic factors to histone "tails” alters the extent to which DNA is wrapped around histones and the availability of genes in the DNA to be activated. All of these factors and processes can have an effect on people's health and influence their health possibly resulting in cancer, autoimmune disease, mental disorders, or diabetes among other illnesses (Figure 1).
DNA Methylation
Much is known about the mechanism of heritability of DNA methylation state during cell division and differentiation. Heritability of methylation state depends on certain enzymes, such as the DNA methyltransferase (DNAMT), that have a higher affinity for 5-methylcytosine than for cytosine. If this enzyme reaches a "hemimethylated” portion of DNA (where 5-methylcytosine is in only one of the two DNA strands) the enzyme will methylate the other half.
DNA methylation patterns are known to be established and modified in response to environmental factors by a complex interplay of at least three independent DNA methyltransferases (DNAMT1, DNAMT3A, and DNAMT3B). DNAMT1 is often referred to as the maintenance methyltransferase. It is essential for proper embryonic development, imprinting, and X-inactivation. To emphasize the difference of this molecular mechanism of inheritance from the canonical Watson-Crick base-pairing mechanism of transmission of genetic information, the term epigenetic templating was introduced. Furthermore, in addition to the maintenance and transmission of methylated DNA states, the same principle could work in the maintenance and transmission of histone modifications.
DNA methylation is an important regulator of gene transcription and a large body of evidence has demonstrated that aberrant DNA methylation is associated with unscheduled gene silencing, and the genes with high levels of 5-methylcytosine in their promoter region are transcriptionally silent. DNA methylation is essential during embryonic development and, in somatic cells, patterns of DNA methylation are in general transmitted to daughter cells with a high fidelity. Aberrant DNA methylation patterns have been associated with a large number of human malignancies and found in two distinct forms: hypermethylation and hypomethylation compared to normal tissue. Hypermethylation is one of the major epigenetic modifications that repress transcription via promoter region of tumor suppressor genes. Hypermethylation typically occurs at CpG islands in the promoter region and is associated with gene inactivation. Global hypomethylation has also been implicated in the development and progression of cancer through different mechanisms.
Histone Modifications
Mechanisms of heritability of histone state are not well understood. Although histone modifications occur throughout the entire sequence, the unstructured N-termini of histones (called histone tails) are particularly highly modified. These modifications include:
A. Acetylation: This is the formation of an acetyl derivative (acetyl is the atom grouping CH3CO, an acetic acid molecule from which the hydroxyl group has been removed). It is the most highly studied. It has a tendency to be associated with "active" transcription and is biophysical in nature. Because it normally has a positively charged nitrogen at its end, lysine can bind the negatively charged phosphates of the DNA backbone. The acetylation event converts the positively charged amine group on the side chain into a neutral amide linkage. This removes the positive charge, thus loosening the DNA from the histone. When this occurs, complexes and other transcriptional factors can bind to the DNA and allow transcription to occur. This is the "cis" model of epigenetic function. In other words, changes to the histone tails have a direct effect on the DNA itself. Another model of epigenetic function is the "trans" model in which changes to the histone tails act indirectly on the DNA. Further, acetylation at one position is likely to function differently from acetylation at another position. The idea that multiple dynamic modifications regulate gene transcription in a systematic and reproducible way is called the histone code.
B. Methylation: This is the addition of methyl groups (methyl is the radical - CH3). It bears the idea that modifications act as docking modules for related factors. It is a chemical endogenous damage to DNA and an important regulator of gene transcription.
C. Phosphorylation: This is the addition of phosphate to an organic compound such as glucose to produce glucose monophosphate through the action of a phosphotransferase (phosphorylase) or kinase.
D. Ribosylation: This is the chemical transformation into a ribosyl (a radical formed by loss of the hemiacetal OH group from either of two cyclic forms of ribose) yielding ribofuranosyl andrybopyranosyl compounds by combination with an H of -nH- or -CH group.
Other modifications include citrillination, sumoylation and ubiquitylation.
Chromatin Remodeling
Because chromatin remodeling (and other mechanisms such as DNA methylation) play such a central role in many types of epigenetic inheritance, the word "epigenetics" is sometimes, albeit misleadingly, used as a synonym for these processes. Chromatin remodeling is not always inherited and not all epigenetic inheritance involves chromatin remodeling. Chromatin remodeling is accomplished through two main mechanisms:
a) Post-translational modification of the amino acids that make up histone proteins: Histone proteins are made up of long chains of amino acids. If the amino acids that are in the chain are changed, the shape of the histone might be modified. DNA is not completely unwound during replication. It is possible, then, that the modified histones may be carried into each new copy of the DNA. Once there, these histones may act as templates, initiating the surrounding new histones to be shaped in the new manner. By altering the shape of the histones around them, these modified histones would ensure that a lineage-specific transcription program is maintained after cell division.
b) Addition of methyl groups to the DNA, mostly at CpG sites, to convert cytosyne to 5-methylcytosine: 5-Methylcytosine performs much like a regular cytosine, pairing with a guanine in double-stranded DNA. However, some areas of the genome are methylated more heavily than others, and highly methylated areas tend to be less transcriptionally active, through a mechanism not fully understood. Methylation of cytosines can also persist from the germ line of one of the parents into the zygote, marking the chromosome as being inherited from one parent or the other (genetic imprinting).
Epigenetic Changes
As we know, epigenetic changes can modify the activation of certain genes, but not the sequence of DNA. Additionally, the chromatin proteins associated with DNA may be activated or silenced. This is why the differentiated cells in a multi-cellular organism express only the genes that are necessary for their own activity.
Epigenetic changes are preserved when cells divide. Most epigenetic changes only occur within the course of one individual organism's lifetime, but, if gene inactivation occurs in a sperm or egg cell that results in fertilization, then some epigenetic changes can be transferred to the next generation. This raises the question of whether or not epigenetic changes in an organism can alter the basic structure of its DNA (in contradistinction with the very definition of epigenetics)
Specific epigenetic processes are multiple and varied: bookmarking, carcinogenesis progress, cloning technical limitations, gene silencing, heterochromatin, histone modification, regulation, imprinting, maternal effects, paramutation, pathogenesis technical limitations, position effect, reprogramming, teratogens' effects, transvection, andX-chromosome inactivation.
DNA damages can also cause epigenetic changes. They are very frequent, occurring on average about 10,000 times a day per cell of the human body. These damages are largely repaired, but at the site of a DNA repair, epigenetic changes can remain. DNA- damaging chemicals, such as benzene, hydroquinone, styrene, carbon tetrachloride, and trichloroethylene cause considerable hypomethylation of DNA.
Foods are known to alter the epigenetics of rats on different diets. Some food components epigenetically increase the levels of DNA repair enzymes while others can reduce DNA damage, such as soy isoflavones and bilberry anthocyanins.
Epigenetics Classification
Epigenetics can be divided into predetermined and probabilistic epigenesis. Predetermined epigenesis is a unidirectional movement from structural development in DNA to the functional maturation of the protein. Predetermined here means that development is scripted and predictable. Probabilistic epigenesis, on the other hand, is a bidirectional structure-function development with experiences and external molding development.
Epigenetic Agents
Epigenetic agents are prions, RNA and micro-RNA;
Prions
In general, proteins fold into discrete units that perform distinct cellular functions, but some proteins are also capable of forming an infectious conformational state known as a prion (protein+infection). Although often viewed in the context of infectious diseases, prions are more loosely defined by their ability to catalytically convert other native state versions of the same protein to an infectious conformational state. It is in this latter sense that they can be viewed as epigenetic agents capable of inducing a phenotypic change without a modification of the genome.
Fungal prions are considered by some to be epigenetic because the infectious phenotype caused by the prion can be inherited without modification of the genome.
RNA and Micro-RNA
Sometimes a gene, after being turned on, transcribes a product that (directly or indirectly) maintains the activity of that gene. RNA signaling includes differential recruitment of a hierarchy of generic chromatin-modifying complexes and DNAMTs to specific loci by RNAs during differentiation and development. Other epigenetic changes are mediated by the production of different splice forms of RNA, or by formation of double-stranded RNA. Descendants of the cell in which the gene was turned on will inherit this activity, even if the original stimulus for gene-activation is no longer present. These genes are often turned on or off by signal transduction, although in some systems RNA may spread directly to other cells or nuclei by diffusion. A large amount of RNA and protein is contributed to the zygote by the mother during oogenesis or via nurse cells resulting in maternal effects' phenotypes. A smaller quantity of sperm RNA is transmitted from the father, but there is recent evidence that this epigenetic information can lead to visible changes in several generations of offspring.
Micro-RNAs (mi-RNAs) are members of non-coding RNAs that range in size from 17 to 25 nucleotides. They regulate a large variety of biological functions. About 2000 mi-RNAs have so far been discovered in humans. It appears that about 60% of human protein coding genes are regulated by mi-RNAs. Many mi-RNAs are epigenetically regulated. About 50% of mi-RNA genes are associated with CpG islands that may be repressed by epigenetic methylation. Other mi-RNAs are epigenetically regulated by either histone modifications or by combined DNA methylation and histone modification.
Evidence in Humans
There are at least three types of epigenetic evidence in humans [12,13]:
Environmental Exposure
Epigenetic changes have been observed to occur in response to environmental exposure. In the case of humans with different environmental exposures, monozygotic (identical) twins were epigenetically indistinguishable during their early years, while older twins had remarkable differences in the overall content and genomic distribution of 5-methylcytosine DNA and histone acetylation. The twin pairs who had spent less of their lifetime together and/or had greater differences in their medical histories were those who showed the largest such differences.
Recent studies involving both dizygotic (not identical) and monozygotic twins have produced some evidence of epigenetic influence in humans. Direct comparisons between identical twins constitute the ideal experimental model for testing environmental epigenetics, because DNA sequence differences that would be abundant in a singleton-based study do not interfere with the analysis. Research has shown that a difference in the environment can produce long-term epigenetic effects, and different developmental monozygotic twin subtypes may be different with respect to their susceptibility to be discordant from an epigenetic point of view.
One of the first high-throughput studies of epigenetic differences between monozygotic twins focused on comparing global and locus-specific changes in DNA methylation and histone modifications in a sample of 40 monozygotic twin pairs. In this case, only healthy twin pairs were studied, but a wide range of ages was represented, between 3 and 74 years.
One of the major conclusions from this study was that there is an age-dependent accumulation of epigenetic differences between the two siblings of twin pairs. This accumulation suggests the existence of epigenetic drift. A more recent study, where 114 monozygotic twins and 80 dizygotic twins were analyzed for the DNA methylation status of around 6,000 unique genomic regions, concluded that epigenetic similarity at the time of blastocyst splitting may also contribute to phenotypic similarities in monozygotic co-twins. This supports the notion that the microenvironment at early stages of embryonic development can be quite important for the establishment of epigenetic marks.
Genomic Imprinting
Some human disorders are associated with genomic imprinting, a phenomenon in mammals where the father and mother contribute different epigenetic patterns for specific genomic loci in their germ cells. The best-known cases of imprinting in human disorders are those in Angelman, Prader- Willi, and Beckwith-Wiedemann syndrome. The former two syndromes can be produced by the same genetic mutation (chromosome 15q partial deletion), and the particular syndrome that will develop depends on whether the mutation is inherited from the child’s mother or father. This is due to the presence of genomic imprinting in the region. The latter syndrome is often caused by abnormalities in maternal genomic imprinting of a region on chromosome 11.
Transgenerational Inheritance
In the Overkalix study, Marcus Pembrey and colleagues observed that the paternal (but not maternal) grandsons of Swedish men who were exposed during preadolescence to famine in the 19th century were less likely to die of cardiovascular disease. If food was plentiful, then diabetes mortality in the grandchildren increased, suggesting that this was a transgenerational epigenetic inheritance. The opposite effect was observed for females—the paternal (but not maternal) granddaughters of women who experienced famine while in the womb (and therefore while their eggs were being formed) lived shorter lives on average. Similar transgenerational effects were observed during the 1944 Dutch Famine. Such transgenerational inheritance traits were noted and studied in the case of these two famines because, especially in the cae of the Dutch Famine, accurate records were kept over long periods of time for both male and female descendants, some continuing even to this day. No doubt such observations could have been made in the previous numerous famines except for the lack of the required meticulous records.
Epigenetic Inheritance
Somatic epigenetic inheritance through epigenetic modifications, particularly through DNA methylation and chromatin remodeling, is very important in the development of multicellular eukaryotic organisms. The genome sequence is static (with some notable exceptions), but cells differentiate into many different types, which perform different functions, and respond differently to the environment and intercellular signaling. Thus, as individuals develop, morphogens activate or silence genes in an epigenetically heritable fashion, giving cells a "memory”.
In mammals, most cells terminally differentiate, with only stem cells retaining the ability to differentiate into several cell types ("totipotency” and "multipotency”). Some stem cells continue producing new differentiated cells throughout life, such as in neurogenesis, but mammals are not able to respond to loss of some tissues, for example, the inability to regenerate limbs, of which some other animals are capable. Unlike animals, plant cells do not terminally differentiate, remaining totipotent with the ability to give rise to a new individual plant. While plants do utilize many of the same epigenetic mechanisms as animals, such as chromatin remodeling, it has been hypothesized that some kinds of plant cells do not use or require "cellular memories”, resetting their gene expression patterns using positional information from the environment and surrounding cells to determine their fate.
Cancer Epigenetics
Defining Cancer Epigenetics
The accepted definition is: "Cancer epigenetics is the study of epigenetic modifications to the genome of cancer cells that do not involve a change in the nucleotide sequence”. Epigenetic alterations are as important as genetic mutations in the transformative processes of a normal cell into cancer. As will further be elaborated, such alterations include the silencing of tumor suppressor genes (TSG) and the activation of tumor promoter genes (TPG, or oncogenes) by altered CpG island methylation patterns, histone modifications, and dysregulation of DNA binding proteins [14-16].
Classically, cancer has been viewed as a set of diseases that are driven by progressive genetic abnormalities including mutations in oncogenes, TSGs, and chromosomal abnormalities. However, it has become apparent that cancer is also driven by epigenetic alterations, which refer to functionally-relevant modifications to the genome that do not involve a change in the nucleotide sequence. Examples of such modifications were given already (changes in DNA methylation: hyper- and hypo-methylation; histone modifications; and changes in chromosomal architecture caused by inappropriate expression of proteins).
Each of these epigenetic alterations serves to regulate gene expression without altering the underlying DNA sequence. These changes may remain through cell divisions, last for multiple generations, and can be considered to be "epimutations” (equivalent to mutations). Epigenetic alterations occur frequently in cancers. However, while large numbers of epigenetic alterations are found in cancers, the epigenetic alterations in DNA repair genes, causing reduced expression of DNA repair proteins, may be of particular importance. Such alterations are thought to occur early in progression to cancer and to be a likely cause of the genetic instability characteristic of cancers. Reduced expression of DNA repair genes causes deficient DNA repair.
In summary, as pointed out above under genetic alterations, cancer is caused by failure to regulate tissue growth, when the genes that regulate cell growth and differentiation are altered. It has become clear that these alterations are caused by both DNA sequence mutation in oncogenes and TSGs as well as by epigenetic alterations. The epigenetic deficiencies in expression of DNA repair genes, in particular, likely cause an increased frequency of mutations, some of which then occur in oncogenes and TSGs.
Epigenetics has the potential to explain mechanisms of aging, human development, and the origins of cancer, heart disease and mental illness as well as several other conditions. Some investigators even think that epigenetics may ultimately turn out to have a greater role in disease than genetics.
A variety of compounds are considered as epigenetic carcinogens, They result in an increased incidence of tumors, but they do not show mutagen activity. Examples include: diethylstilbestrol, arsenite, hexachlorobenzene, and nickel compounds.
Recent studies have shown that the mixed lineage leukemia (MLL) gene causes leukemia by rearranging and fusing with other genes in different chromosomes, which is a process under epigenetic control. Other investigations have concluded that alterations in histone acetylation and DNA methylation occur in various genes influencing prostate cancer. Gene expression in the prostate can be modulated by nutrition and lifestyle changes [17,18].
DNA Repair Epigenetics
Germ line (familial) mutations have been identified in 34 different DNA repair genes that cause a high risk of cancer, including, for example BRCA1 and ATM. However, cancers caused by such germ line mutations make up only a very small proportion of cancers. For instance, germ line mutations cause only 2% to 5% of colon cancer cases.
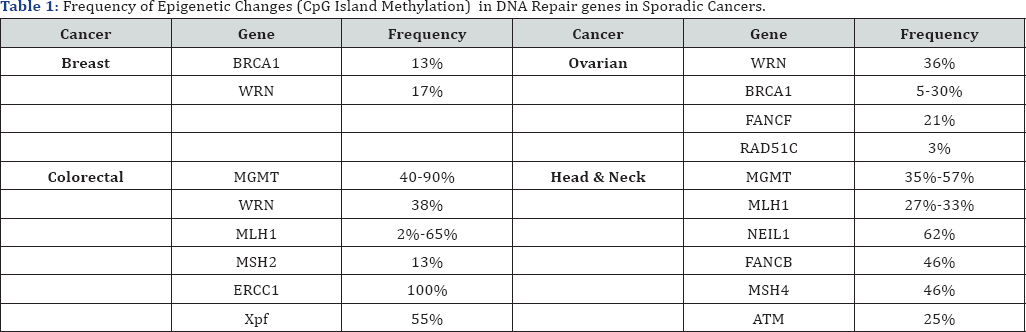
Epigenetic reductions in expression of DNA repair genes, however, are very frequent in sporadic (non-germ line) cancers, as shown among some representative cancers in Table 1, while mutations in DNA repair genes in sporadic cancer are very rare.
Deficiencies in expression of DNA repair genes cause increased mutation rates and cause genome instability, which is likely the main underlying cause of the genetic alterations leading to cancer. In fact, the first event in many sporadic neoplasias is a heritable alteration that affects genetic instability and epigenetic defects in DNA repair are somatically heritable.
Epigenetic Carcinogens
A variety of toxic compounds or pathogens (diethylstilbestrol, arsenite, hexachlorobenzene, nickel) are considered as epigenetic carcinogens in that they increase the incidence of tumors. However, they do not show mutagen activity. By epigenetic mechanisms, many teratogens (such as diethylstilbestrol) exert specific effects on the fetus and throughout the life of an affected child. However, the possibility of birth defects resulting from the child's parents and their parents' exposures have not been observed. However, a range of male-mediated abnormalities have been demonstrated [19].
Cancer Treatment
Manipulations of epigenetic mutations hold great promise for cancer prevention, detection, and therapy. Several medications which have epigenetic impact are now used in several of these diseases [20-22].
Current research has shown that epigenetic pharmaceuticals could be a replacement or adjuvant therapy for currently accepted treatment methods such as radiation and chemotherapy, or could enhance the effects of these current treatments. It has been shown that the epigenetic control of the proto-onco regions and the tumor suppressor sequences by conformational changes in histones directly affects the formation and progression of cancer. Epigenetics also has the factor of reversibility, a characteristic that other cancer treatments do not offer.
Drug development has focused mainly on Histone Acetyl Transferase (HAT) and Histone DeACetylase (HDAC), and has included the introduction to the market of the new pharmaceutical Vorinostat, an HDAC inhibitor. HDAC has been shown to play an integral role in the progression of oral squamous cancer. Current front-runner candidates for new drug targets are Histone Lysine Methyl Transferase (HLMT) and Protein Arginine Methyl Transferase (PAMT).
Applications of epigenetics to cancer sub-types (cervical, leukemia, prostate, sarcoma, etc.) will not be discussed here. More epigenetic aspects remain to be discussed such as functional epigenomics (or the engineering of the epigenome), RNA and beyond RNA-epigenetics. Such a discussion would go far beyond the scope of this article and will not likewise be presented here.
Summary and Conclusion
Epigenetics is the study of cellular and physiological traits inherited by daughter cells, but not caused by changes in the DNA sequence. Thus, unlike genetics, which is based on changes to the DNA sequence (the genotype), in epigenetics, the changes in gene expression or cellular phenotype have other causes. The term epigenetics also refers to the changes in the genome that do not involve a change in the nucleotide sequence.
The epigenetic evidence in the case of humans came from (a) twin studies, (b) genomic imprinting, and (c) transgenerational inheritance. Direct comparisons between identical twins constitute the ideal experimental model for testing environmental epigenetics. Research has shown that a difference in the environment can produce long-term epigenetic effects, and different developmental monozygotic twin subtypes may be different with respect to their susceptibility to be discordant. Some human disorders are associated with genomic imprinting, a phenomenon wherein the father and mother contribute different epigenetic patterns. Best-known cases of imprinting in human disorders are those in Angelman, Prader-Willi, and Beckwith-Wiedemann syndrome. As for transgenerational inheritance, the 1866-1868 Finland/North Sweden Famine and the 1944 Dutch Famine have provided notable evidence of the effects of famine on the descendants of those who lived through these disastrous events. No doubt such inheritance effects could have been observed in other famines had thorough records been maintained.
Epigenetic changes do take place and can modify the activation of certain genes, but not the sequence of DNA. Additionally, the chromatin proteins associated with DNA may be activated or silenced. The changes are preserved when cells divide. Whereas most epigenetic changes only occur within the course of one individual organism's lifetime, they can be transferred to the next generation if gene inactivation occurs in a sperm or egg cell. Specific epigenetic processes are multiple and varied and DNA damage can also cause epigenetic changes.
Three types of epigenetic mechanisms have been discussed (a) chromatic remodeling, (b) histone modifications, and (c) DNA methylation. Chromatin remodeling is accomplished through two main mechanisms: post-translational modification of the amino acids that make up histone protein, and addition of methyl groups to the DNA. Although histone modifications occur throughout the entire sequence, mechanisms of heritability of histone state are not well understood. These modifications include: acetylation, citrillination, methylation, phosphorylation, ribosylation, sumoylation, and ubiquitylation. Much is known about the mechanism of heritability of DNA methylation state during cell division and differentiation. Heritability of methylation state depends on certain enzymes, such as the DNA methyltransferase (DNAMT1), that have a higher affinity for 5-methylcytosine than for cytosine. If this enzyme reaches a "hemimethylated" portion of DNA (where 5-methylcytosine is in only one of the two DNA strands) the enzyme will methylate the other half. DNA methylation patterns are known to be established and modified in response to environmental factors. Its molecular mechanism of inheritance is different from the canonical Watson-Crick base-pairing mechanism.
There are various applications of epigenetics in medicine and drug development, of particular importance in cancer and cancer treatment. Epigenetic pharmaceuticals could be a replacement or adjuvant therapy for currently accepted treatment methods such as radiation and chemotherapy, or could enhance the effects of these current treatments. The epigenetic control of the proto-onco regions and the tumor suppressor sequences by conformational changes in histones directly affects the formation and progression of cancer. Epigenetics also has the factor of reversibility, a characteristic that other cancer treatments do not offer.
Drug development (e.g., Vorinostat) has focused mainly on histone acetyl transferase and histone deacetylase. Current front-runner candidates for new drug targets are histone lysine methyl transferase and protein arginine methyl transferase. The discussions of other drug developments, applications to cancer sub-types, functional epigenomics (or epigenome engineering), and RNA and beyond RNA-epigenetics remain to be fully investigated in the future.
References
- Gupta K, Miller JD, Li JZ, Russell MW, Charbonneau C (2008) Epidemiologic and socioeconomic burden of metastatic renal cell carcinoma (mRCC): a literature review. Cancer Treat Rev 34(3): 193205.
- Hollingsworth JM, Miller DC, Daignault S, Hollenbeck BK (2006) Rising incidence of small renal masses: a need to reassess treatment effect. J Natl Cancer Inst 98(18): 1331-1334.
- Hollingsworth JM, Miller DC, Daignault S, Hollenbeck BK (2007) Five- year survival after surgical treatment for kidney cancer: a population- based competing risk analysis. Cancer 109(9): 1763-1768.
- Rini BI (2009) Metastatic Renal Cell Carcinoma: Many Treatment Options, One Patient. J Clin Oncol 27(19): 3225-3234.
- Pfaffenroth EC, Linehan WM (2008) Genetic basis for kidney cancer: opportunity for disease-specific approaches to therapy. Expert Opin Biol Ther 8(6): 779-790.
- Reuter VE (2006) The pathology of renal epithelial neoplasms. Semin Oncol 33(5): 534-543.
- Linehan WM, Srinivasan R, Schmidt LS (2010) The genetic basis of kidney cancer: a metabolic disease. Nat Rev Urol 7(5): 277-285.
- Choueiri TK, Bukowski RM, Rini BI (2006) The current role of angiogenesis inhibitors in the treatment of renal cell carcinoma. Semin Oncol 33(5): 596-606.
- Motzer RJ, Bander NH, Nanus DM (1996) Renal-cell carcinoma. N Engl J Med 335(12): 865-875.
- Motzer RJ, Hutson TE, Tomczak P, Bycott PW, Baum CM, et al. (2007) Sunitinib versus interferon alfa in metastatic renal cell carcinoma. N Engl J Med 356(2): 115-124.
- Escudier B, Eisen T, Stadler WM, Hutson TE, Gore M, et al. (2007) Sorafenib in advanced clear-cell renal-cell carcinoma. N Engl J Med 356(2): 125-134.
- Hudes G, Carducci M, Tomczak P, Moore L, Motzer RJ, et al. (2007) Temsirolimus, interferon alfa, or both for advanced renal-cell carcinoma. N Engl J Med 356(22): 2271-2281.
- Cohen HT, McGovern FJ (2005) Renal-cell carcinoma. N Engl J Med 353: 2477-2490.
- Dharmawardana PG, Giubellino A, Bottaro DP (2004) Hereditary papillary renal carcinoma type I. Curr Mol Med 4(8): 855-868.
- Sudarshan S, Linehan WM (2006) Genetic basis of cancer of the kidney. Semin Oncol 33(5): 544-551.
- Launonen V, Vierimaa O, Kiuru M, Herva R, Aaltonen LA, et al. (2001) Inherited susceptibility to uterine leiomyomas and renal cell cancer. Proc Natl Acad Sci U S A 98(6): 3387-3392.
- Tomlinson IP, Alam NA, Rowan AJ, Paetau A, Salovaara R, et al. (2002) Germline mutations in FH predispose to dominantly inherited uterine fibroids, skin leiomyomata and papillary renal cell cancer. Nat Genet 30(4): 406-410.
- Merino MJ, Torres-Cabala C, Pinto P, Linehan WM (2007) The morphologic spectrum of kidney tumors in hereditary leiomyomatosis and renal cell carcinoma (HLRCC) syndrome. Am J Surg Pathol 31(10):1578-1585.
- Cheville JC, Lohse CM, Zincke H, Weaver AL, Blute ML (2003) Comparisons of outcome and prognostic features among histologic subtypes of renal cell carcinoma. Am J Surg Pathol 27(5): 612-624.
- Pavlovich CP, Walther MM, Eyler RA, et al. (2002) Renal tumors in the Birt-Hogg-Dube syndrome. Am J Surg Pathol 26(12): 1542-1552.
- Schmidt LS, Warren MB, Nickerson ML, Vera Matrosova,2 Jorge R Toro, et al. (2001) Birt-Hogg-Dube syndrome, a genodermatosis associated with spontaneous pneumothorax and kidney neoplasia, maps to chromosome 17p11.2. Am J Hum Genet 69(4): 876-882.
- Vocke CD, Yang Y, Pavlovich CP, Zbar B, Linehan WM, et al. (2005) High frequency of somatic frameshift bhd gene mutations in Birt-Hogg- Dube-associated renal tumors. J Natl Cancer Inst 97(12): 931-935.
- Motzer RJ, Bacik J, Mariani T, Russo P, Mazumdar M, Reuter V (2002) Treatment outcome and survival associated with metastatic renal cell carcinoma of non-clear-cell histology. J Clin Oncol 20(9): 2376-2381.
- Motzer RJ, Michaelson MD, Redman BG, Bello CL, Theuer CP, et al.(2006) Activity of SU11248, a multitargeted inhibitor of vascular endothelial growth factor receptor and platelet-derived growth factor receptor, in patients with metastatic renal cell carcinoma. J Clin Oncol 24(1): 16-24.
- Awada A, Hendlisz A, Gil T, Schwartz B, Piccart M, et al. (2005) Phase I safety and pharmacokinetics of BAY 43-9006 administered for 21 days on/7 days off in patients with advanced, refractory solid tumours. Br J Cancer 92(10): 1855-1861.
- Clark JW, Eder JP, Ryan D, Lathia C, Lenz HJ (2005) Safety and pharmacokinetics of the dual action Raf kinase and vascular endothelial growth factor receptor inhibitor, BAY 43-9006, in patients with advanced, refractory solid tumors. Clin Cancer Res 11(15): 54725480.
- Escudier B, Szczylik C, Hutson TE, Shah S, Bukowski RM, et al. (2009) Randomized phase II trial of first-line treatment with sorafenib versus interferon Alfa-2a in patients with metastatic renal cell carcinoma. J Clin Oncol 27(8): 1280-1289.
- Sternberg CN, Davis ID, Mardiak J, McCann L, Pandite L, et al. (2010) Pazopanib in locally advanced or metastatic renal cell carcinoma: results of a randomized phase III trial. J Clin Oncol 28(6): 1061-1068.
- Gore ME, Szczylik C, Porta C, Schoffski P, Mainwaring P, et al. (2009) Safety and efficacy of sunitinib for metastatic renal-cell carcinoma: an expanded-access trial. Lancet Oncol 10: 757-763.
- Choueiri TK, Plantade A, Elson P, Bukowski RM, Escudier B, et al. (2008) Efficacy of sunitinib and sorafenib in metastatic papillary and chromophobe renal cell carcinoma. J Clin Oncol 26: 127-131.
- Hudes G, Carducci M, Tomczak P, Robert Figlin, Anil Kapoor, et al. Temsirolimus, interferon alfa, or both for advanced renal-cell carcinoma. N Engl J Med 356: 2271-2281.
- Dutcher JP, de Souza P, McDermott D, Feingold J, Hudes G, et al. (2009) Effect of temsirolimus versus interferon-alpha on outcome of patients with advanced renal cell carcinoma of different tumor histologies. Med Oncol 26(2): 202-209.
- Nanus DM, Garino A, Milowsky MI, Larkin M, Dutcher JP (2004) Active chemotherapy for sarcomatoid and rapidly progressing renal cell carcinoma. Cancer 101:1545-1251.
- Dutcher JP, Nanus D (2010) Long-term survival of patients with sarcomatoid renal cell cancer treated with chemotherapy. Med Oncol 28(4): 1530-1533.
- Oudard S, Banu E, Vieillefond A, Medioni J, Banu A, et al. (2007) Prospective multicenter phase II study of gemcitabine plus platinum salt for metastatic collecting duct carcinoma: results of a GETUG (Groupe d'etudes des tumeurs uro-genitales) study. J Urol 177:16981702.
- Gordon MS, Hussey M, Nagle RB, Pan CX, Crawford D, et al. (2009) Phase II study of erlotinib in patients with locally advanced or metastatic papillary histology renal cell cancer SWOG S0317. J Clin Oncol 27(34): 5788-5793.






























