In Silico Analysis of Seed Storage Protein Gene Promoters Reveals Differential Occurrence of 7 Cis-Regulatory Elements in Monocot and 14 in Dicot Plants
Zaiba Hasan Khan1, Richa Patel2, Sandhya Mehrotra1 and Rajesh Mehrotra1*
1Department of Biological Sciences, BITS Pilani, KK Birla Goa campus, India
2Department of Biological Sciences, Birla Institute of Technology and Science, India
Submission: April 10,2019; Published: June 12, 2019
*Corresponding author: Rajesh Mehrotra, Department of Biological Sciences, BITS- Pilani, KK Birla Goa campus, Goa, 403726, India
How to cite this article: Zaiba Hasan Khan, Richa Patel, Sandhya Mehrotra, Rajesh Mehrotra. In Silico Analysis of Seed Storage Protein Gene Promoters Reveals Differential Occurrence of 7 Cis-Regulatory Elements in Monocot and 14 in Dicot Plants. Adv Biotech & Micro. 2019; 14(2): 555881. DOI: 10.19080/AIBM.2019.14.555881
Abstract
A major part of the human diet consists of cereals and legumes all over the world. The major dietary requirement of the developing world includes seed storage proteins of grains. However, seeds are incomplete in nutrition because of their low protein content. Synthesis of seed storage proteins is partly regulated by Cis-Regulatory Elements (CREs) and transcription factors besides post transcriptional and post-translational level. We performed an in-silico analysis to find out significant occurrences of various cis-elements/motifs in promoters of Seed Storage Proteins (SSPs) from 26 monocots and 28 dicot plant groups. In this paper, an in-silico analysis was performed on 1kb region of promoters of 26 monocots and 28 dicots plant of Seed Storage Proteins (SSPs) genes to find out the occurrences of different cis elements/ motifs. Further, a statistical method was used to test the significant occurrence of motifs among a large group of the CREs/motifs. The analysis reveals differential yet significant occurrence of 7 motifs in promoters of SSPs of monocots and 14 motifs in dicots plants among 225 motifs obtained through analysis using student’s t-test. The study will be helpful in designing promoters or chimeric promoter by modifying the architecture of promoters with respect to significant CREs to enhanced deposition of protein in seeds and to manipulate the seed storage oil contents.
Keywords: Seed storage proteins; Promoter; Cis elements; Transcription; Monocots; Dicots
Abbrevations: SSP: Seed Storage Proteins; CRE: Cis-Regulatory Elements; BP: Base Pair; LEA: Late Embryogenic Abundant; TSS: Transcription Start Site; PLACE: Plant Cis-Acting Regulatory Elements
Introduction
A major part of the human diet consists of cereals and legumes worldwide. 70% of human diet comprises of legumes and cereals. Recently, increasing seed storage proteins of any plants (rice, wheat, maize, barley) in order to improve its nutritional value has successively become one of the important goals for quality breeding [1]. Protein contents and composition are crucial to grain quality and nutritional value of these crops. Plants accumulate storage substances such as lipids, starch and proteins in certain phases of development. Seed Storage Proteins (SSPs) are proteins that accumulate significantly in the developing seed, whose main function is to act as a source of nitrogen, carbon, and sulfur skeletons for the developing embryo [2]. However, the nutritional quality is not very high and intensive research is going on to form modified seed storage proteins so as to increase its nutritional value as food for human and animal feed [3]. Seed Storage Proteins are formed during seed development and maturation phase and are of two types, mid-embryogenesis and Late Embryogenesis Abundant (LEA) proteins [4]. The various types of seed storage proteins in dicots include water soluble albumins and saline soluble globulins whereas alcohol soluble prolamins are included in monocots such as gliadin, avenin, zein and glutenin [5].
The regulation of transcription of genes encoding seed storage proteins temporally and spatially is mediated by the Cis-Regulatory Elements (CREs) residing in the promoters and corresponding sequence specific Transcription Factors (TFs). These CRES are characterized by a consensus sequence known as motifs which is a conserved or frequently occurring sequence of defined length, usually 3-25 Base Pairs (BP) of promoters [6]. Motifs can be found in non-coding part of DNA especially in promoters or enhancers upstream of a gene. Seed storage proteins within their proximal and distal part of promoters contain such multiple short conserved sequences also known as Cis-Regulatory Elements (CREs). Usually they lie upstream to the Transcription Start Site (TSS) of gene. Each motif occurring within promoter is typically associated with certain biological functions as they contain unique composition of nucleotide sequence known as Transcription Factor Binding Sites (TFBS) where Transcription Factors (TFs) binds and regulate the expression of genes encoding seed storage proteins. A wide range of CREs is required for seed specific transcription of seed storage proteins and their various subunits eventually regulate the seed storage protein synthesis.
Hence, the present analysis endeavours to find out motifs which have differential occurrence in two different group of plants: monocots and dicots. For this in-silico approaches have been employed to analyze 26 seed storage gene protein promoters from monocots and 28 from dicots were selected. In, promoter region (1000-bp) of each gene, motifs were identified using the PLACE database which is a collection of CREs (motifs) present in plant cis-acting regulatory DNA elements, taken up from previously published reports [7].
Materials and Methods
Retrieval of promoter regions and analysis of cisregulatory elements
We manually selected types and subcategories of seed storage proteins in monocots and in dicots from the NCBI (National Centre for Biotechnology Information) database. A total of 26 seed storage proteins from monocots and 28 from dicots belong to different class of SSPs were selected for the study. Categories of seed proteins chosen for monocots were glutelin, zein, avenin, globulin, vicilin, hordein etc. whereas for dicots; legumin, LEA, albumin, globulin, glycilin have been selected.
Considering the fact that motifs conferring seed-specific expression lie in the proximal part of the promoter, often within 500 bp upstream of the transcriptional start site, we have selected promoter sequence of 1000 bp upstream from Transcription Start Site (TSS) using TAIR (The Arabidopsis Information Resource) database [8]. Promoter sequences of monocots and dicots seed storage proteins were scanned to find out the occurrences of CREs, their position on plus (+) and minus (-) strand on DNA using PLACE tool.
Statistical method to test significant occurrences of motifs
To test the statistically significance of particular motifs a Student’s t-test was performed [9]. We know that the difference between the sample mean will not always be zero, even if the samples belong to the same population. It is statistically very rare for the difference in two samples means to lie on the margins of the distribution. Therefore, if the difference does lie on the margins, it is statistically significant to conclude that the samples were extracted from two different populations. An independentsamples t-test was conducted to compare frequency of occurrence of a particular motif in monocots and dicots.
Mean position of motifs
From the combined motif results mean position of all the motifs separately for monocots and dicots and for positive (+) and negative (-) strand was calculated. The code written for the same takes the unique list of motifs and creates a HashMap of the unique motif name with its details including count and list of positions of motifs in positive and negative strands. Every time the motif is encountered in the list of all motifs, it is looked up in the Hashmap in constant time. New position is added to the list and count is incremented. Once we have scanned the whole data, we calculated the average position by dividing the sum of all the positions by count (frequency of that motif).
Results
Analysis of cis-regulatory motifs in promoters of SSPs from monocots and dicots
In promoter sequences up to 1 kb upstream of the TSS of each seed storage genes of monocots and dicots were scanned using PLACE database for the recognition of CREs (motifs) as described in materials and method section. The study revealed cumulatively 225 CREs in both the monocots and dicots having various functions in plants. Individually, in 26 SSPs promoters of monocot group we got a total of 211 motifs whereas, 218 CREs have been recognized in 28 SSPs promoters from dicots plants. While there are 21 motifs unique to monocot group and 25 motifs unique to dicot group among 225 motifs.
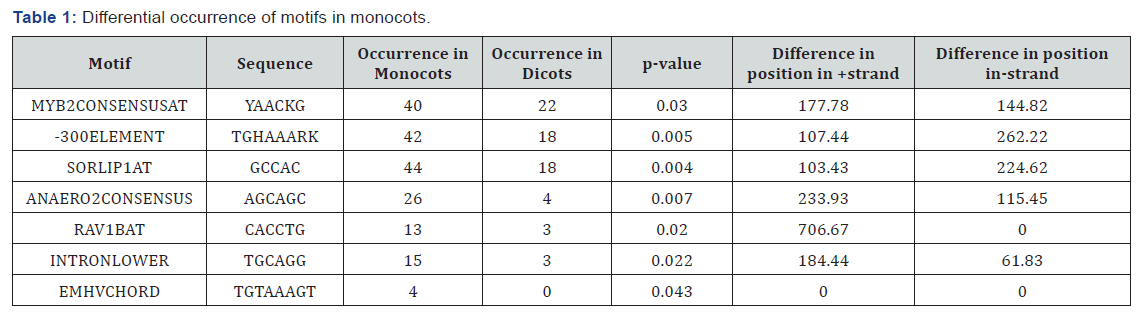
*no value indicated absence of the motif in the entire + or - strand (either in monocot or dicot)
Mean position of motifs
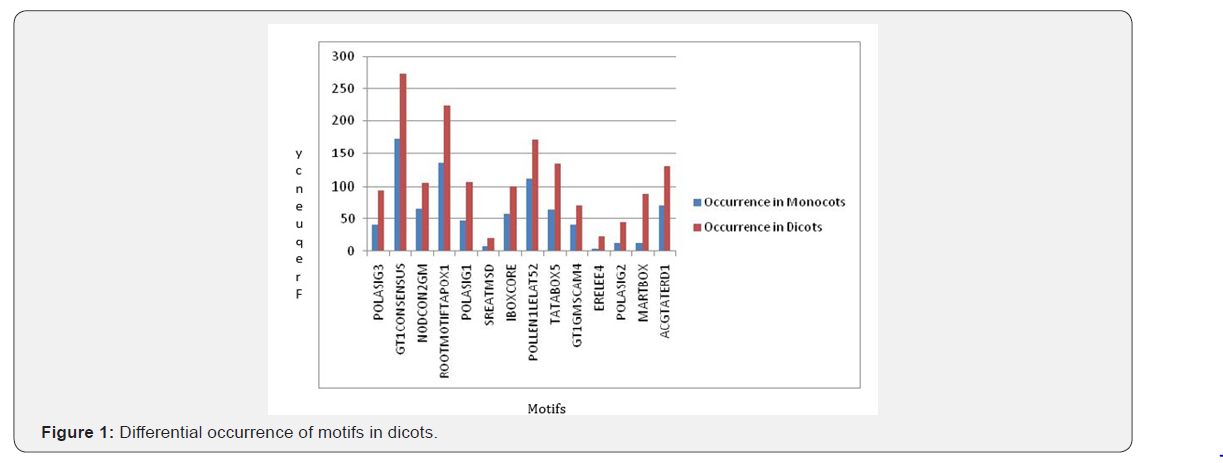
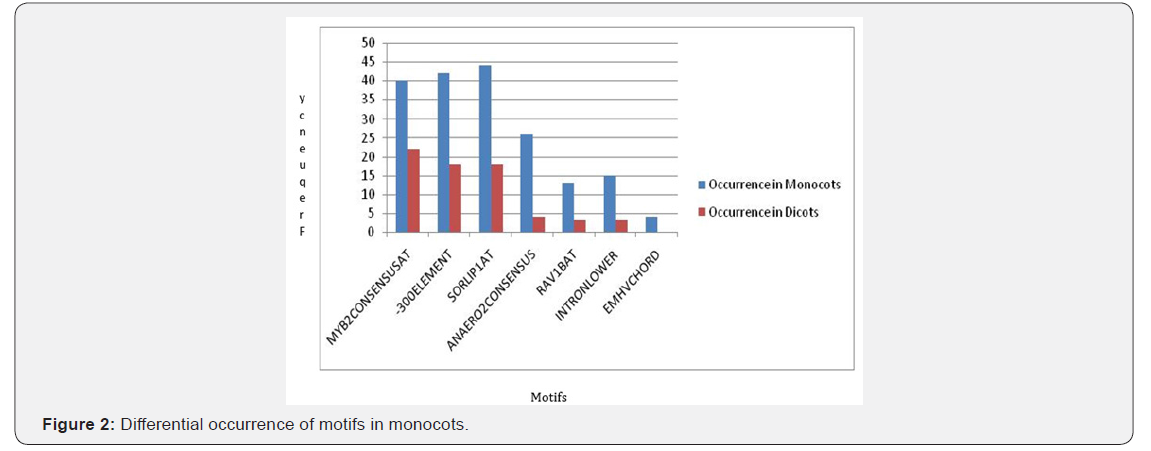
For all motifs their positions on + and - strand were determined then the average position of that particular motif on both the strands was calculated and the difference between the mean positions were calculated. By analyzing the data, we got 14 motifs in dicots whose mean positions are significantly different from the monocots (Figure1). Similarly, for monocots,7 motifs have been found out which show significant difference than dicots (Figure 2). Table 1 is showing an overview of differences in the mean positions of motifs in both monocots and dicots (last two columns). If a particular motif found either in monocot or dicot their difference could not be calculated which was indicated as 0 (Table 1).
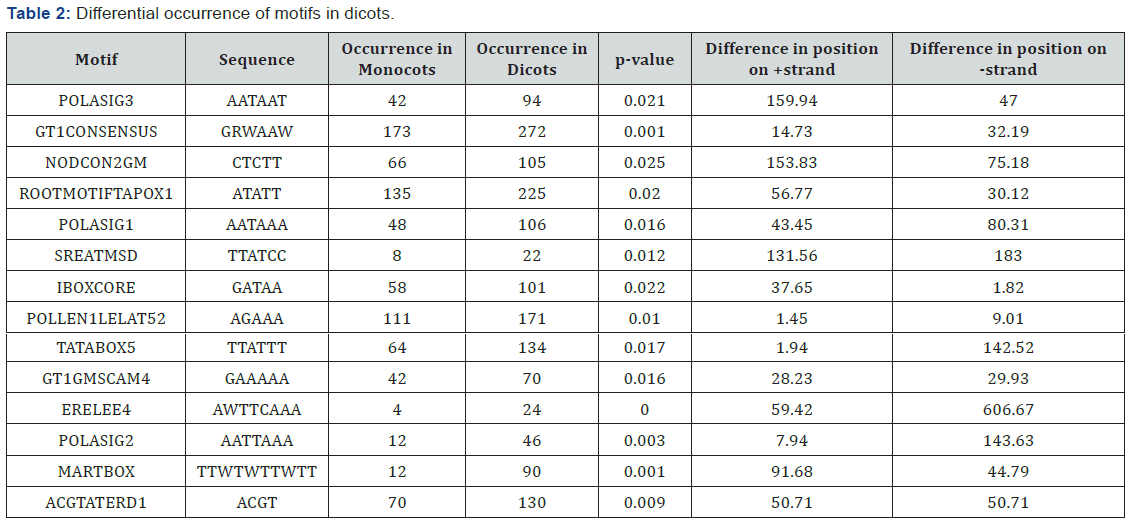
Statistical significance of occurrences of motifs
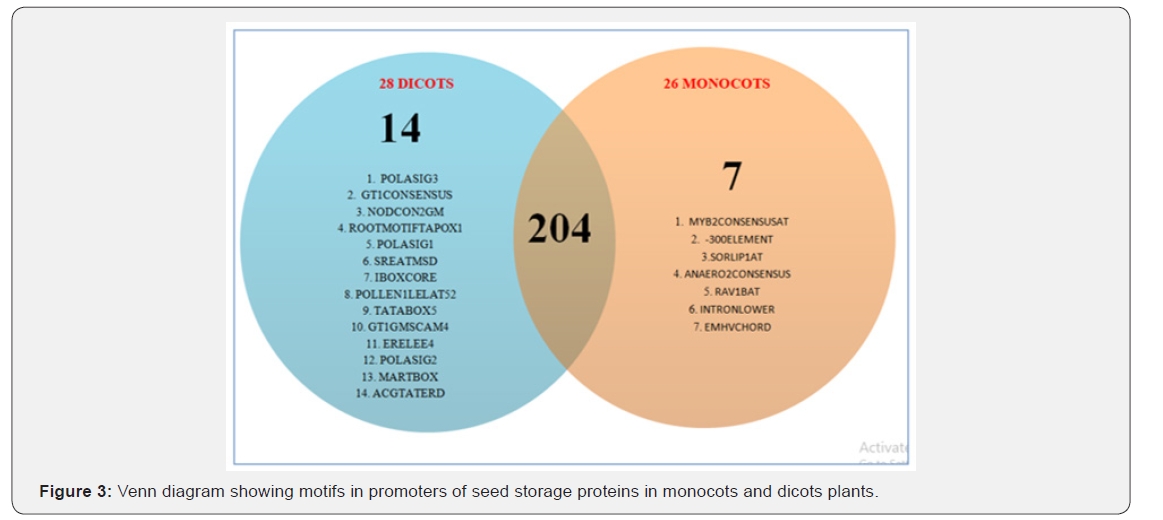
Positions of all the 225 different motifs obtained in promoters of SSPs considered in monocots and dicots were also determined on + and - strands. Among them motifs with higher levels of significance are chosen, i.e, the likelihood that the random sample chosen were not the representative of the population is less than 5% (p value <0.05). We set the level of α=0.05 so those motifs with p-value less than 0.05 were shortlisted. There are 7 motifs that are predominant in monocots as compared to dicots (Table-1). The scenario is different for dicots where there are 14 motifs showing high abundance in dicots as compared to monocots (Table-2). In total, we found out 204 motifs common to both the groups, making them the usual plant motifs. We measured how significantly different the number of occurrences of cis regulatory motifs is there in promoters of SSPs. We found that the difference was significant for motifs occurrences in dicots and monocots. This shows that 7 motifs occur significantly within the 26 promoters of SSPs genes in monocots while in dicots, 14 motifs were found to be significant in 28 promoters of SSPs genes (Figure 3). In general motifs are not exclusively present in monocots or dicots but there (as highlighted above) are motifs whose frequencies in monocots and dicots have a significant difference.

Discussion
In our study we have done cis-regulatory motifs analysis on 26 and 28 seed storage proteins of two different groups of plants: monocots and dicots respectively. Through our analysis we found out a diverse range of CREs in the promoters of SSPs, many of these upstream promoter elements are responsible for basal transcription and precision of initiation of transcription of SSPs. The fundamental problem is that many of the known regulatory regions (enhancer and promoters) do not contain very much significant motifs so this analysis definitely gave an idea about the motifs which are significant or not in promoter regions of genes. Here, we have applied statistical method to check significant occurrences of motifs in monocots and dicots plants thus the results suggest statistical preference of some of the motifs in monocots while the others are preferred in dicots only among a large group of motifs. These statistically significant motifs are generally over-representation in promoter region of SSPs.
However, studies on promoter analysis of seed storage proteins from mustard, legumes and grasses shows abundance of various motifs viz. RY-like and ACGT-like, E2Fb-like, GCN4-like motif, prolamin-box-like, Skn-1-like [10] while some negative elements like AACA were also present [6, 11]. Also, through our previous in-silico analysis on promoters of 38 SSPs we found out certain motifs like E-box and RY belongs to monocots only while GCN4 and AACA are restricted to the dicots merely. In this study we reported the statistically enriched motif in the promoters of SSPs in two different groups of plants, monocots and dicots. However, the number of common motifs present between two groups of However, studies on promoter analysis of seed storage proteins from mustard, legumes and grasses shows abundance of various motifs viz. RY-like and ACGT-like, E2Fb-like, GCN4-like motif, prolamin-box-like, Skn-1-like [10] while some negative elements like AACA were also present [6, 11]. Also, through our previous in-silico analysis on promoters of 38 SSPs we found out certain motifs like E-box and RY belongs to monocots only while GCN4 and AACA are restricted to the dicots merely. In this study we reported the statistically enriched motif in the promoters of SSPs in two different groups of plants, monocots and dicots. However, the number of common motifs present between two groups of
Analysis of cis motifs enriched in the promoter regions of SSPs genes would be helpful in better understanding of temporal and spatial regulation of their gene expression. Knowledge on ciselements individually or their combinatorial effect will allow us to effectively change the expression pattern of a gene in required way. This study would be helpful in developing transgenic plants with improved nutritional quality. Numerous approaches have been used by plant biotechnologist continuously from conventional to molecular breeding towards enhancement of nutritive content of cereals like rice, wheat, maize [13]. Uses of genetic engineering techniques like characterization studies of promoters of seed storage proteins as well as the establishment of new technologies for genetic engineering and plant transformations could be useful for improvement of grain quality in different cereals.
The study would be beneficial in better understanding of the mechanism of regulation of seed storage proteins at transcriptional level and will widen up the scope in transgenic technology where transgenic crops could be produced by manipulating the enhancers region mainly promoters [14] of seed storage proteins in order to achieve desired level of human nutrition. The observations could be used to target protein in seeds using unique motifs as well as in designing seed specific promoters (chimeric promoters) by manipulating the genetic architecture of promoters in terms of their CREs.
Acknowledgement
We would like to thank BITS-Pilani, Pilani campus and K.K. Birla Goa campus for the infrastructure and the facilities provided. RM and SM are thankful to DST-SERB for the funding. ZHK is thankful to DEPARTMENT OF SCIENCE AND TECHNOLOGY (DST) for the fellowship provided.
Authors’ contributions
ZHK wrote and edited the manuscript. RP had performed the computations and drafted the manuscript. RM conceived the original idea and SM investigate and supervised the findings of this work. All authors discussed the results and contributed to the final manuscript.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
References
- Mandal S, Mandal RK (2000) Seed storage proteins and approaches for improvement of their nutritional quality by genetic engineering. Current Sciences 79(5): 576-589.
- Krishnan HB and Coe EH (2001) Seed storage proteins. Encyclopedia of Genetics. New York: Academic Press, USA, pp.782-787.
- Shewry PR, Halford NG (2002) Cereal seed storage proteins: structures, properties and role in grain utilization. J Exp Bot 53(370): 947-958.
- Mehrotra R, Kumar S, Mehrotra S, Singh BD (2009) Seed storage protein gene regulation-A jig-saw puzzle. IJBT 8:147-158.
- Shewry PR, Napier JA, Tatham AS (1995) Seed Storage Proteins: Structures and Biosynthesis. Plant Cell 7(7): 945-956.
- Mehrotra R, Gupta G, Sethi R, Bhalothia P, Kumar N, et al. (2011) Designer promoter: an artwork of cis engineering. Plant Mol Biol 75(6): 527-536.
- Higo K, Ugawa Y, Iwamoto M, Korenaga T (1999) Plant cis-acting regulatory DNA elements (PLACE) database. Nucleic Acids Res 26(1): 297-300.
- Berardini TZ, Reiser L, Li D, Mezheritsky Y, Muller R, et al. (2015) The Arabidopsis Information Resource: Making and mining the gold standard annotated reference plant genome. Genesis 53(8): 774-785.
- Kim TK (2015) T test as a parametric statistic. Korean J Anesthesiol 68(6): 540-546.
- Fauteux F, Strömvik, MV (2009) Seed storage protein gene promoters contain conserved DNA motifs in Brassicaceae, Fabaceae and Poaceae. BMC Plant Biol 9: 126-137.
- Bhalothia P, Alok A, Mehrotra S, Mehrotra R (2013) AACA element negatively regulates expression of protein phosphatase 2C (PP2C) like promoter in Arabidopsis thaliana. Am J Plant Sci 4(3): 549-554.
- Cserhati M (2015) Motif content comparison between monocot and dicot species Genom Data 3: 128-136.
- Toenniessen GH (2002) Crop Genetic Improvement for Enhanced Human nutrition. J Nutr 132(9): 2943s-2946s.
- Mehrotra R, Sethi S, Zutshi I, Bhalothia P, Mehrotra S (2013) Patterns and evolution of ACGT repeat cis-element landscape across four plant genomes. BMC Genomics 14(1): 203.






























